

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147179.5 - phase: 0
(706 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
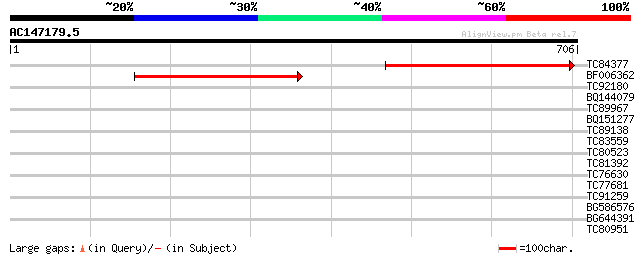
Score E
Sequences producing significant alignments: (bits) Value
TC84377 weakly similar to GP|20161282|dbj|BAB90208. hypothetical... 496 e-140
BF006362 similar to GP|21618069|gb unknown {Arabidopsis thaliana... 432 e-121
TC92180 33 0.49
BQ144079 similar to GP|6523547|emb| hydroxyproline-rich glycopro... 30 3.2
TC89967 similar to PIR|T08962|T08962 hypothetical protein F19B15... 30 3.2
BQ151277 30 4.2
TC89138 homologue to GP|15215736|gb|AAK91413.1 AT5g19690/T29J13_... 29 5.4
TC83559 similar to GP|11994399|dbj|BAB02358. contains similarity... 29 5.4
TC80523 homologue to GP|3860321|emb|CAA10128.1 beta-galactosidas... 29 5.4
TC81392 weakly similar to GP|19310589|gb|AAL85025.1 unknown prot... 29 7.1
TC76630 homologue to GP|18033654|gb|AAL57201.1 putative nodule m... 29 7.1
TC77681 similar to GP|22597156|gb|AAN03465.1 nucleolar histone d... 29 7.1
TC91259 weakly similar to GP|17065054|gb|AAL32681.1 putative pro... 28 9.2
BG586576 similar to GP|10129885|db mitochondrial elongation fact... 28 9.2
BG644391 28 9.2
TC80951 similar to PIR|D96519|D96519 mysoin-like protein 11013-... 28 9.2
>TC84377 weakly similar to GP|20161282|dbj|BAB90208. hypothetical
protein~similar to Arabidopsis thaliana chromosome 3
At3g03305, partial (14%)
Length = 899
Score = 496 bits (1276), Expect = e-140
Identities = 235/235 (100%), Positives = 235/235 (100%)
Frame = +2
Query: 469 TLTELRPFSINGHSFRLSWSWKEFFVMGCQWASLYYPLLWSALGFMFSFLLVPKALLFFQ 528
TLTELRPFSINGHSFRLSWSWKEFFVMGCQWASLYYPLLWSALGFMFSFLLVPKALLFFQ
Sbjct: 2 TLTELRPFSINGHSFRLSWSWKEFFVMGCQWASLYYPLLWSALGFMFSFLLVPKALLFFQ 181
Query: 529 MNLYTYRNFIANKGVVNGALWILQELCRVPTLWFGWIGYLFYLILFPWFMGQVFTEGKST 588
MNLYTYRNFIANKGVVNGALWILQELCRVPTLWFGWIGYLFYLILFPWFMGQVFTEGKST
Sbjct: 182 MNLYTYRNFIANKGVVNGALWILQELCRVPTLWFGWIGYLFYLILFPWFMGQVFTEGKST 361
Query: 589 VYMTYMGWAVETSNGKGKFEWVGFPDIMVLVLPHILFVVLPAILVTGALTAERAIYRERV 648
VYMTYMGWAVETSNGKGKFEWVGFPDIMVLVLPHILFVVLPAILVTGALTAERAIYRERV
Sbjct: 362 VYMTYMGWAVETSNGKGKFEWVGFPDIMVLVLPHILFVVLPAILVTGALTAERAIYRERV 541
Query: 649 LALSGKKKDDIDLNSRRPLLNGNHNSTISTLHLGKRWIRKLLCVVCLAICWKHIM 703
LALSGKKKDDIDLNSRRPLLNGNHNSTISTLHLGKRWIRKLLCVVCLAICWKHIM
Sbjct: 542 LALSGKKKDDIDLNSRRPLLNGNHNSTISTLHLGKRWIRKLLCVVCLAICWKHIM 706
>BF006362 similar to GP|21618069|gb unknown {Arabidopsis thaliana}, partial
(22%)
Length = 629
Score = 432 bits (1111), Expect = e-121
Identities = 209/209 (100%), Positives = 209/209 (100%)
Frame = +2
Query: 156 FSKYSVNGQLGRNRSVNSVTLETKERKHLFVGIDTTMSAGAGLRGPTNVFGHPTDQLLKD 215
FSKYSVNGQLGRNRSVNSVTLETKERKHLFVGIDTTMSAGAGLRGPTNVFGHPTDQLLKD
Sbjct: 2 FSKYSVNGQLGRNRSVNSVTLETKERKHLFVGIDTTMSAGAGLRGPTNVFGHPTDQLLKD 181
Query: 216 LDLELSHWDSQSEKPVTKISFGHFPLSFSAPSSSGRTLKEVFLKHSISAYLCGHLHSRFG 275
LDLELSHWDSQSEKPVTKISFGHFPLSFSAPSSSGRTLKEVFLKHSISAYLCGHLHSRFG
Sbjct: 182 LDLELSHWDSQSEKPVTKISFGHFPLSFSAPSSSGRTLKEVFLKHSISAYLCGHLHSRFG 361
Query: 276 KNLKRHHQLSNRFLSLQNFFQFNVHQNSFESTVNCSIGAPPQEFWEWEIGDWRKSRAIRI 335
KNLKRHHQLSNRFLSLQNFFQFNVHQNSFESTVNCSIGAPPQEFWEWEIGDWRKSRAIRI
Sbjct: 362 KNLKRHHQLSNRFLSLQNFFQFNVHQNSFESTVNCSIGAPPQEFWEWEIGDWRKSRAIRI 541
Query: 336 LAIDRGHVSYVDLDFKSGAKHAIILPTFP 364
LAIDRGHVSYVDLDFKSGAKHAIILPTFP
Sbjct: 542 LAIDRGHVSYVDLDFKSGAKHAIILPTFP 628
>TC92180
Length = 528
Score = 32.7 bits (73), Expect = 0.49
Identities = 12/29 (41%), Positives = 19/29 (65%)
Frame = +2
Query: 504 YPLLWSALGFMFSFLLVPKALLFFQMNLY 532
+P+LWS L F+F+F +P+ FF +Y
Sbjct: 281 FPILWSLLVFLFAFS*IPRFFSFFSFPIY 367
>BQ144079 similar to GP|6523547|emb| hydroxyproline-rich glycoprotein DZ-HRGP
{Volvox carteri f. nagariensis}, partial (12%)
Length = 1071
Score = 30.0 bits (66), Expect = 3.2
Identities = 16/61 (26%), Positives = 26/61 (42%)
Frame = -2
Query: 561 WFGWIGYLFYLILFPWFMGQVFTEGKSTVYMTYMGWAVETSNGKGKFEWVGFPDIMVLVL 620
WFGWIG+ ++ W +G + + + W V G EW+G + +V
Sbjct: 578 WFGWIGWWLWM----WRVGGLLVFVLAVFCRAVLAWPVRLLRG----EWLGLGMGLAVVC 423
Query: 621 P 621
P
Sbjct: 422 P 420
>TC89967 similar to PIR|T08962|T08962 hypothetical protein F19B15.100 -
Arabidopsis thaliana, partial (28%)
Length = 877
Score = 30.0 bits (66), Expect = 3.2
Identities = 18/44 (40%), Positives = 25/44 (55%)
Frame = +2
Query: 42 IEPKGTPDSAVWVVQLSDLHLSVHHPNRALDFTNLVGHALSFIN 85
I+PK TPDSA +VQ + SV+ P L+G LSF++
Sbjct: 173 IDPKFTPDSATHLVQKAQQESSVNDP-------KLLGWPLSFLS 283
>BQ151277
Length = 964
Score = 29.6 bits (65), Expect = 4.2
Identities = 14/37 (37%), Positives = 19/37 (50%), Gaps = 5/37 (13%)
Frame = -3
Query: 498 QWASLYYPLLWSALGFMFSFLLV-----PKALLFFQM 529
+W PL W LGF+F FL + P +LFF +
Sbjct: 137 RWPHTVVPLRWLGLGFLFLFLAIYCTFPPLLVLFFPL 27
>TC89138 homologue to GP|15215736|gb|AAK91413.1 AT5g19690/T29J13_110
{Arabidopsis thaliana}, partial (31%)
Length = 909
Score = 29.3 bits (64), Expect = 5.4
Identities = 27/99 (27%), Positives = 45/99 (45%), Gaps = 7/99 (7%)
Frame = -2
Query: 35 CCNGETVIEP------KGTPDSAVWVVQLSDLHLSVHHPNRALDFTNLVGHA-LSFINPS 87
CC G+T I+ KG P S + V + +P ++ V + F+N +
Sbjct: 416 CCQGQTWIDSPTDYTAKGIPSSVIKPVP------KIIYPILCQKLSDSVVEVRIEFMNNA 255
Query: 88 LVLITGDLTDGKSKDLLTMKQNEDEWVEYRNVMDGVIER 126
LVL G+ TD + ++ ++D+ E NV + V ER
Sbjct: 254 LVLYNGEKTDREGEN-----TDQDQNEERENVTECVPER 153
>TC83559 similar to GP|11994399|dbj|BAB02358. contains similarity to
receptor protein kinase~gene_id:MIL23.20 {Arabidopsis
thaliana}, partial (21%)
Length = 1217
Score = 29.3 bits (64), Expect = 5.4
Identities = 33/130 (25%), Positives = 52/130 (39%), Gaps = 11/130 (8%)
Frame = +2
Query: 195 GAGLRGPTNVFGHPTDQLLKDLDLELSHWDSQSEKPVTKISFGHFPLSFSAPSSSGRTLK 254
G L +N+F PT ++LK ++ + P T S + F+ SG
Sbjct: 119 GTNLTYISNLFNQPTSEILK--------YNPNIQNPDTIQSHTRLNIPFTCDCLSG---- 262
Query: 255 EVFLKHSISAYLCGHLHSRFGKNLKRHHQLSNRFLSLQNFFQFNVHQNSFES-------- 306
+FL H+ S L + G+N K ++N + S F + NS+ +
Sbjct: 263 -LFLGHTFSYKL------KEGENYKA---VANGYYSNLTTIDFLIRVNSYPATDIPAGTV 412
Query: 307 ---TVNCSIG 313
TVNCS G
Sbjct: 413 INVTVNCSCG 442
>TC80523 homologue to GP|3860321|emb|CAA10128.1 beta-galactosidase {Cicer
arietinum}, partial (41%)
Length = 1109
Score = 29.3 bits (64), Expect = 5.4
Identities = 31/129 (24%), Positives = 51/129 (39%), Gaps = 20/129 (15%)
Frame = -2
Query: 199 RGPTNVFGHPTDQLLKDLDLE--------LSHWD----SQSEKPVTKISFGHFPLSFSAP 246
R P V G Q+LK + + +W+ S++ + GH L +++P
Sbjct: 949 RDPQLVAGDSIHQILKTNHSKTQSIV*ACMVYWERNHHSKNHCKYLSLDLGHH-LPYTSP 773
Query: 247 SSSGRTLKEVFLKHSISA--------YLCGHLHSRFGKNLKRHHQLSNRFLSLQNFFQFN 298
+++ FL+ S+S +L GH HS+F K L H F
Sbjct: 772 KGHLYLIQQPFLQPSLSMHDQKLQVLFLVGHTHSQFVKELLDHLVTETAF---------- 623
Query: 299 VHQNSFEST 307
H +SFE +
Sbjct: 622 -HSSSFEQS 599
>TC81392 weakly similar to GP|19310589|gb|AAL85025.1 unknown protein
{Arabidopsis thaliana}, partial (34%)
Length = 1277
Score = 28.9 bits (63), Expect = 7.1
Identities = 18/59 (30%), Positives = 27/59 (45%)
Frame = -2
Query: 227 SEKPVTKISFGHFPLSFSAPSSSGRTLKEVFLKHSISAYLCGHLHSRFGKNLKRHHQLS 285
S + I F HFPL++ G++LK L+H I + L+ NL +H S
Sbjct: 856 SHNHINFIPFNHFPLNYGI---KGKSLKPFSLEHKIQYHYISILNHDI*NNL*LNHSHS 689
>TC76630 homologue to GP|18033654|gb|AAL57201.1 putative nodule membrane
protein {Medicago sativa}, complete
Length = 2605
Score = 28.9 bits (63), Expect = 7.1
Identities = 19/48 (39%), Positives = 26/48 (53%), Gaps = 3/48 (6%)
Frame = -2
Query: 132 SLFFDLRGN---HDSFGVPVVGGSFDFFSKYSVNGQLGRNRSVNSVTL 176
SL +GN H +F VPVVG S + S++ LGRN S + +L
Sbjct: 1368 SLSLQKKGNFRIHCTFMVPVVGHSSNVVEFTSISSHLGRN*SSQNKSL 1225
>TC77681 similar to GP|22597156|gb|AAN03465.1 nucleolar histone deacetylase
HD2-P39 {Glycine max}, partial (47%)
Length = 1233
Score = 28.9 bits (63), Expect = 7.1
Identities = 27/118 (22%), Positives = 45/118 (37%), Gaps = 16/118 (13%)
Frame = -3
Query: 461 SNDIMGRSTLTELRPFSINGHSFRLSWSWKEFFVMGCQWASLYYPL-----LWSALGFMF 515
S I+GR ++ + F W W + G QW + + L + F+
Sbjct: 538 SCSILGRCSILGISFFRFRNLQLLCFWFWFSILLCGNQWDVFIFTISFTFFLTIIIVFIV 359
Query: 516 SFL----------LVPKALLFFQMNLYTY-RNFIANKGVVNGALWILQELCRVPTLWF 562
SF+ +V F+ N+ Y RNF+ +G N L+ E+ +WF
Sbjct: 358 SFVSTEMNTSSFGIVR*LKFLFKNNIKRYLRNFVLGQGSQNKFLFTNFEVNNHRLVWF 185
>TC91259 weakly similar to GP|17065054|gb|AAL32681.1 putative protein
{Arabidopsis thaliana}, partial (6%)
Length = 723
Score = 28.5 bits (62), Expect = 9.2
Identities = 16/44 (36%), Positives = 25/44 (56%), Gaps = 4/44 (9%)
Frame = +3
Query: 667 LLNGNHNSTISTLHLGK-RWIRKLLCVV---CLAICWKHIMVMF 706
++N N+N+ TL+ RWI LLC+ C+ +C+ I MF
Sbjct: 276 IINNNNNNKC*TLNYTMFRWILLLLCLFQQQCMNLCYHFINPMF 407
>BG586576 similar to GP|10129885|db mitochondrial elongation factor G {Oryza
sativa (japonica cultivar-group)}, partial (30%)
Length = 745
Score = 28.5 bits (62), Expect = 9.2
Identities = 13/30 (43%), Positives = 18/30 (59%)
Frame = -3
Query: 444 YKAFEDTSPDRFWFQIESNDIMGRSTLTEL 473
+KAFE+ P+ F +NDI G TL E+
Sbjct: 203 FKAFENCPPESFETGCTANDITGSGTLIEV 114
>BG644391
Length = 441
Score = 28.5 bits (62), Expect = 9.2
Identities = 26/96 (27%), Positives = 43/96 (44%), Gaps = 11/96 (11%)
Frame = +2
Query: 547 ALWILQELCRVPTLWFGWIGYLFYLILFPWFMGQVF---------TEGKSTVYMTYMGWA 597
+L +L C V L GW+ YL +ILF W++ Q+ T+ +VY +
Sbjct: 38 SLLLLIAKCSVREL--GWMCYLLAIILFFWYIKQLMNATPIWC*STKCLDSVYSISIPST 211
Query: 598 VETSNGKGKFEWVGFPDI--MVLVLPHILFVVLPAI 631
+ S G F V + + +L H+L++ P I
Sbjct: 212 RQLSRLVGMFYCVVYQRLGYFLLCFVHLLYLQQPRI 319
>TC80951 similar to PIR|D96519|D96519 mysoin-like protein 11013-7318
[imported] - Arabidopsis thaliana, partial (14%)
Length = 1050
Score = 28.5 bits (62), Expect = 9.2
Identities = 13/42 (30%), Positives = 24/42 (56%), Gaps = 8/42 (19%)
Frame = -3
Query: 669 NGNHNSTISTL--------HLGKRWIRKLLCVVCLAICWKHI 702
NG+H I+ + H+ R+ R LL + CLA+C++++
Sbjct: 466 NGDHACCIACILAVYAFMVHMDGRFWRSLLLITCLALCFQNL 341
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.324 0.138 0.439
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 26,307,956
Number of Sequences: 36976
Number of extensions: 422245
Number of successful extensions: 3192
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 3151
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3192
length of query: 706
length of database: 9,014,727
effective HSP length: 103
effective length of query: 603
effective length of database: 5,206,199
effective search space: 3139337997
effective search space used: 3139337997
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 62 (28.5 bits)
Medicago: description of AC147179.5


