

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147178.10 + phase: 0
(161 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
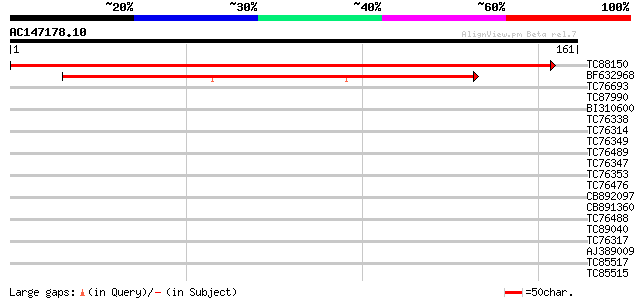
Score E
Sequences producing significant alignments: (bits) Value
TC88150 similar to GP|21537019|gb|AAM61360.1 unknown {Arabidopsi... 308 8e-85
BF632968 weakly similar to GP|9858772|gb| BAC19.4 {Lycopersicon ... 145 7e-36
TC76693 adenosylhomocysteinase [Medicago truncatula] 28 1.2
TC87990 similar to PIR|T03216|T03216 enoyl-[acyl-carrier-protein... 28 2.1
BI310600 similar to PIR|S52770|S52 subtilisin-like proteinase (E... 27 2.8
TC76338 PIR|S17435|S17435 polyubiquitin 6 - common sunflower, pa... 27 4.7
TC76314 PIR|S17435|S17435 polyubiquitin 6 - common sunflower, pa... 27 4.7
TC76349 GP|1332579|emb|CAA66667.1 polyubiquitin {Pinus sylvestri... 27 4.7
TC76489 PIR|S17435|S17435 polyubiquitin 6 - common sunflower, pa... 27 4.7
TC76347 PIR|S17435|S17435 polyubiquitin 6 - common sunflower, pa... 26 6.1
TC76353 PIR|S06921|UQPM polyubiquitin 5 - garden pea, partial (89%) 26 6.1
TC76476 PIR|S06921|UQPM polyubiquitin 5 - garden pea, partial (80%) 26 6.1
CB892097 PIR|S17435|S1 polyubiquitin 6 - common sunflower, parti... 26 6.1
CB891360 homologue to PIR|T02995|T02 unspecific monooxygenase (E... 26 6.1
TC76488 homologue to GP|14596193|gb|AAK68824.1 Unknown protein {... 26 6.1
TC89040 similar to GP|21594042|gb|AAM65960.1 unknown {Arabidopsi... 26 8.0
TC76317 PIR|S06921|UQPM polyubiquitin 5 - garden pea, complete 26 8.0
AJ389009 26 8.0
TC85517 homologue to SP|P34788|RS18_ARATH 40S ribosomal protein ... 26 8.0
TC85515 homologue to SP|P34788|RS18_ARATH 40S ribosomal protein ... 26 8.0
>TC88150 similar to GP|21537019|gb|AAM61360.1 unknown {Arabidopsis
thaliana}, partial (80%)
Length = 767
Score = 308 bits (788), Expect = 8e-85
Identities = 151/155 (97%), Positives = 152/155 (97%)
Frame = +2
Query: 1 MKDDDGLPTTTAAINKKENMDSGLFGKGRYKFWALAAILLLAFWSMFTGTVSLRWSGNLN 60
MKDDDGLPTTTAAINKKENMDSGLFGKGRYKFWALAAILLLAFWSMFTGTVSLRWSGNLN
Sbjct: 245 MKDDDGLPTTTAAINKKENMDSGLFGKGRYKFWALAAILLLAFWSMFTGTVSLRWSGNLN 424
Query: 61 TLSSDLDTPIHDDLDVLEMEEREKVVRHMWDVYTNSRRIRLPRFWQEAFEAAYEELTSDV 120
TLSSDLDTPIHDDLDVLEMEEREKVVRHMWDVYTNSRRIRLPRFWQEAFEAAYEELTSDV
Sbjct: 425 TLSSDLDTPIHDDLDVLEMEEREKVVRHMWDVYTNSRRIRLPRFWQEAFEAAYEELTSDV 604
Query: 121 PGVRDDAITEIAKMSVRSVNYDPPPIQSTVSFPFS 155
PGVRDDAITEIAKMSVRSVNYDPPPIQST + FS
Sbjct: 605 PGVRDDAITEIAKMSVRSVNYDPPPIQSTRAQEFS 709
>BF632968 weakly similar to GP|9858772|gb| BAC19.4 {Lycopersicon esculentum},
partial (56%)
Length = 644
Score = 145 bits (366), Expect = 7e-36
Identities = 77/121 (63%), Positives = 91/121 (74%), Gaps = 3/121 (2%)
Frame = +1
Query: 16 KKENMDSGLFGKGRYKFWALAAILLLAFWSMFTGTVSLRWS-GNLNTLSSDLDTPIHDDL 74
K+ M + GK YK ALAAI+L+A W MFTG+++L+ S NLN S DLD+ I DDL
Sbjct: 94 KRAIMHASAAGKRHYKLCALAAIVLIALWFMFTGSITLKLSINNLNPFSDDLDSMILDDL 273
Query: 75 DVLEMEEREKVVRHMWDVYT--NSRRIRLPRFWQEAFEAAYEELTSDVPGVRDDAITEIA 132
DVLE+EEREKVVRHMWDVYT +S I LP+FW EAFEAAYE L SDVP VRD A++EIA
Sbjct: 274 DVLEVEEREKVVRHMWDVYTRSHSNSIGLPQFWLEAFEAAYEHLVSDVPAVRDAAVSEIA 453
Query: 133 K 133
K
Sbjct: 454 K 456
>TC76693 adenosylhomocysteinase [Medicago truncatula]
Length = 2035
Score = 28.5 bits (62), Expect = 1.2
Identities = 27/106 (25%), Positives = 46/106 (42%), Gaps = 2/106 (1%)
Frame = -2
Query: 58 NLNTLSSDLDTPIHDDLDVLEMEEREKVVRHMWDVYTNSRRIR-LPRFWQEAFEA-AYEE 115
N N S L P+ L + + + H+ + + RRIR L + F + +
Sbjct: 771 NTNNSSLHLLVPLRISLQTISDNGKNNL--HLSII--SGRRIR*LTSLLKNLFSLNTFMD 604
Query: 116 LTSDVPGVRDDAITEIAKMSVRSVNYDPPPIQSTVSFPFSTHGAVS 161
SD+ + +D I A ++S PP + ++SFP GAV+
Sbjct: 603 QESDITTIINDKIWSTASTPIKSAFRAPPILLESLSFPCEHGGAVT 466
>TC87990 similar to PIR|T03216|T03216 enoyl-[acyl-carrier-protein] reductase
(NADH) (EC 1.3.1.9) precursor - common tobacco, partial
(89%)
Length = 1615
Score = 27.7 bits (60), Expect = 2.1
Identities = 17/41 (41%), Positives = 21/41 (50%)
Frame = +3
Query: 36 AAILLLAFWSMFTGTVSLRWSGNLNTLSSDLDTPIHDDLDV 76
AA L S TGTV L LN + +D+PI DLD+
Sbjct: 1197 AAFLSSPLASAITGTV-LYVDNGLNAMGVGVDSPIFSDLDI 1316
>BI310600 similar to PIR|S52770|S52 subtilisin-like proteinase (EC 3.4.21.-)
nodule-specific - Arabidopsis thaliana (fragment),
partial (19%)
Length = 648
Score = 27.3 bits (59), Expect = 2.8
Identities = 15/43 (34%), Positives = 23/43 (52%), Gaps = 3/43 (6%)
Frame = +1
Query: 62 LSSDLDTPIHDDLDVL---EMEEREKVVRHMWDVYTNSRRIRL 101
LS + T LD+L ++ + +RH W V T S+R+RL
Sbjct: 37 LSQFIQTGAQLPLDLL**QQLIRLTRTIRHYWTVLTKSQRLRL 165
>TC76338 PIR|S17435|S17435 polyubiquitin 6 - common sunflower, partial (80%)
Length = 1172
Score = 26.6 bits (57), Expect = 4.7
Identities = 23/82 (28%), Positives = 35/82 (42%), Gaps = 8/82 (9%)
Frame = -3
Query: 32 FWALAAILLLAFWSMFTGTVSLRWSGNLNTLSSDLDTPIHDDLDVLEMEEREKVVRHMWD 91
FW L + +L S F +SL WSG + +LS L L ++ +V+
Sbjct: 474 FWML*SAKVLPSSSCFPAKMSLCWSGGIPSLSCILAL----TLSMVSELSTSRVIVFPVR 307
Query: 92 VYTN--------SRRIRLPRFW 105
V+TN S + R+ FW
Sbjct: 306 VFTNICIPPRRRSTKWRVDSFW 241
Score = 25.8 bits (55), Expect = 8.0
Identities = 12/32 (37%), Positives = 18/32 (55%)
Frame = -3
Query: 32 FWALAAILLLAFWSMFTGTVSLRWSGNLNTLS 63
FW L + +L S F ++L WSG + +LS
Sbjct: 930 FWML*SAKVLPSSSCFPAKINLCWSGGMPSLS 835
>TC76314 PIR|S17435|S17435 polyubiquitin 6 - common sunflower, partial (66%)
Length = 978
Score = 26.6 bits (57), Expect = 4.7
Identities = 23/82 (28%), Positives = 35/82 (42%), Gaps = 8/82 (9%)
Frame = -3
Query: 32 FWALAAILLLAFWSMFTGTVSLRWSGNLNTLSSDLDTPIHDDLDVLEMEEREKVVRHMWD 91
FW L + +L S F +SL WSG + +LS L L ++ +V+
Sbjct: 706 FWML*SAKVLPSSSCFPAKMSLCWSGGIPSLSCILAL----TLSMVSELSTSRVIVFPVR 539
Query: 92 VYTN--------SRRIRLPRFW 105
V+TN S + R+ FW
Sbjct: 538 VFTNICIPPRRRSTKWRVDSFW 473
>TC76349 GP|1332579|emb|CAA66667.1 polyubiquitin {Pinus sylvestris}, partial
(70%)
Length = 2040
Score = 26.6 bits (57), Expect = 4.7
Identities = 23/82 (28%), Positives = 35/82 (42%), Gaps = 8/82 (9%)
Frame = -2
Query: 32 FWALAAILLLAFWSMFTGTVSLRWSGNLNTLSSDLDTPIHDDLDVLEMEEREKVVRHMWD 91
FW L + +L S F +SL WSG + +LS L L ++ +V+
Sbjct: 617 FWML*SAKVLPSSSCFPAKMSLCWSGGIPSLSCILAL----TLSMVSELSTSRVIVFPVR 450
Query: 92 VYTN--------SRRIRLPRFW 105
V+TN S + R+ FW
Sbjct: 449 VFTNICIPPRRRSTKWRVDSFW 384
Score = 25.8 bits (55), Expect = 8.0
Identities = 12/32 (37%), Positives = 18/32 (55%)
Frame = -2
Query: 32 FWALAAILLLAFWSMFTGTVSLRWSGNLNTLS 63
FW L + +L S F ++L WSG + +LS
Sbjct: 1301 FWML*SAKVLPSSSCFPAKINLCWSGGMPSLS 1206
>TC76489 PIR|S17435|S17435 polyubiquitin 6 - common sunflower, partial (54%)
Length = 805
Score = 26.6 bits (57), Expect = 4.7
Identities = 23/82 (28%), Positives = 35/82 (42%), Gaps = 8/82 (9%)
Frame = -2
Query: 32 FWALAAILLLAFWSMFTGTVSLRWSGNLNTLSSDLDTPIHDDLDVLEMEEREKVVRHMWD 91
FW L + +L S F +SL WSG + +LS L L ++ +V+
Sbjct: 408 FWML*SAKVLPSSSCFPAKMSLCWSGGIPSLSCILAL----TLSMVSELSTSRVIVFPVR 241
Query: 92 VYTN--------SRRIRLPRFW 105
V+TN S + R+ FW
Sbjct: 240 VFTNICIPPRRRSTKWRVDSFW 175
>TC76347 PIR|S17435|S17435 polyubiquitin 6 - common sunflower, partial (89%)
Length = 1421
Score = 26.2 bits (56), Expect = 6.1
Identities = 13/32 (40%), Positives = 18/32 (55%)
Frame = -3
Query: 32 FWALAAILLLAFWSMFTGTVSLRWSGNLNTLS 63
FW L + +L S F +SL WSG + +LS
Sbjct: 1065 FWML*SARVLPSSSCFPAKMSLCWSGGMPSLS 970
Score = 26.2 bits (56), Expect = 6.1
Identities = 13/32 (40%), Positives = 18/32 (55%)
Frame = -3
Query: 32 FWALAAILLLAFWSMFTGTVSLRWSGNLNTLS 63
FW L + +L S F +SL WSG + +LS
Sbjct: 837 FWML*SARVLPSSSCFPAKMSLCWSGGMPSLS 742
Score = 26.2 bits (56), Expect = 6.1
Identities = 23/82 (28%), Positives = 35/82 (42%), Gaps = 8/82 (9%)
Frame = -3
Query: 32 FWALAAILLLAFWSMFTGTVSLRWSGNLNTLSSDLDTPIHDDLDVLEMEEREKVVRHMWD 91
FW L + +L S F +SL WSG + +LS L L ++ +V+
Sbjct: 609 FWML*SANVLPSSSCFPAKMSLCWSGGIPSLSCILAL----TLSMVSELSTSRVIVFPVR 442
Query: 92 VYTN--------SRRIRLPRFW 105
V+TN S + R+ FW
Sbjct: 441 VFTNICIPPRRRSTKWRVDSFW 376
Score = 26.2 bits (56), Expect = 6.1
Identities = 13/32 (40%), Positives = 18/32 (55%)
Frame = -3
Query: 32 FWALAAILLLAFWSMFTGTVSLRWSGNLNTLS 63
FW L + +L S F +SL WSG + +LS
Sbjct: 1293 FWML*SARVLPSSSCFPAKMSLCWSGGMPSLS 1198
>TC76353 PIR|S06921|UQPM polyubiquitin 5 - garden pea, partial (89%)
Length = 1246
Score = 26.2 bits (56), Expect = 6.1
Identities = 13/32 (40%), Positives = 18/32 (55%)
Frame = -1
Query: 32 FWALAAILLLAFWSMFTGTVSLRWSGNLNTLS 63
FW L + +L S F +SL WSG + +LS
Sbjct: 298 FWML*SARVLPSSSCFPAKMSLCWSGGMPSLS 203
>TC76476 PIR|S06921|UQPM polyubiquitin 5 - garden pea, partial (80%)
Length = 1208
Score = 26.2 bits (56), Expect = 6.1
Identities = 13/32 (40%), Positives = 18/32 (55%)
Frame = -1
Query: 32 FWALAAILLLAFWSMFTGTVSLRWSGNLNTLS 63
FW L + +L S F +SL WSG + +LS
Sbjct: 527 FWML*SARVLPSSSCFPAKMSLCWSGGIPSLS 432
>CB892097 PIR|S17435|S1 polyubiquitin 6 - common sunflower, partial (29%)
Length = 648
Score = 26.2 bits (56), Expect = 6.1
Identities = 13/32 (40%), Positives = 18/32 (55%)
Frame = -2
Query: 32 FWALAAILLLAFWSMFTGTVSLRWSGNLNTLS 63
FW L + +L S F +SL WSG + +LS
Sbjct: 446 FWML*SAKVLPSSSCFPAKMSLCWSGGIPSLS 351
>CB891360 homologue to PIR|T02995|T02 unspecific monooxygenase (EC 1.14.14.1)
- common tobacco, partial (23%)
Length = 726
Score = 26.2 bits (56), Expect = 6.1
Identities = 13/32 (40%), Positives = 18/32 (55%)
Frame = -1
Query: 32 FWALAAILLLAFWSMFTGTVSLRWSGNLNTLS 63
FW L + +L S F +SL WSG + +LS
Sbjct: 651 FWML*SAKVLPSSSCFPAKMSLCWSGGIPSLS 556
>TC76488 homologue to GP|14596193|gb|AAK68824.1 Unknown protein {Arabidopsis
thaliana}, partial (55%)
Length = 493
Score = 26.2 bits (56), Expect = 6.1
Identities = 13/32 (40%), Positives = 18/32 (55%)
Frame = -2
Query: 32 FWALAAILLLAFWSMFTGTVSLRWSGNLNTLS 63
FW L + +L S F +SL WSG + +LS
Sbjct: 222 FWML*SAKVLPSSSCFPAKMSLCWSGGIPSLS 127
>TC89040 similar to GP|21594042|gb|AAM65960.1 unknown {Arabidopsis
thaliana}, partial (67%)
Length = 842
Score = 25.8 bits (55), Expect = 8.0
Identities = 10/39 (25%), Positives = 20/39 (50%)
Frame = +2
Query: 80 EEREKVVRHMWDVYTNSRRIRLPRFWQEAFEAAYEELTS 118
E+R+ + R ++D+ I + W+ E ++LTS
Sbjct: 194 EQRQTITRALFDIVKEHGPITVSNTWERVKEVGLKDLTS 310
>TC76317 PIR|S06921|UQPM polyubiquitin 5 - garden pea, complete
Length = 1375
Score = 25.8 bits (55), Expect = 8.0
Identities = 12/32 (37%), Positives = 18/32 (55%)
Frame = -3
Query: 32 FWALAAILLLAFWSMFTGTVSLRWSGNLNTLS 63
FW L + +L S F ++L WSG + +LS
Sbjct: 464 FWML*SAKVLPSSSCFPAKINLCWSGGMPSLS 369
>AJ389009
Length = 452
Score = 25.8 bits (55), Expect = 8.0
Identities = 8/27 (29%), Positives = 15/27 (54%)
Frame = +1
Query: 29 RYKFWALAAILLLAFWSMFTGTVSLRW 55
R K+W A +L++ W ++ G + W
Sbjct: 73 RRKYWPYAPLLMVLQWLLWKGKEKIEW 153
>TC85517 homologue to SP|P34788|RS18_ARATH 40S ribosomal protein S18.
[Mouse-ear cress] {Arabidopsis thaliana}, complete
Length = 736
Score = 25.8 bits (55), Expect = 8.0
Identities = 10/29 (34%), Positives = 17/29 (58%)
Frame = +1
Query: 62 LSSDLDTPIHDDLDVLEMEEREKVVRHMW 90
+S+ LD + DDL+ L+ + +RH W
Sbjct: 334 VSNQLDMKLRDDLERLKKIRNHRGLRHYW 420
>TC85515 homologue to SP|P34788|RS18_ARATH 40S ribosomal protein S18.
[Mouse-ear cress] {Arabidopsis thaliana}, complete
Length = 1018
Score = 25.8 bits (55), Expect = 8.0
Identities = 10/29 (34%), Positives = 17/29 (58%)
Frame = +1
Query: 62 LSSDLDTPIHDDLDVLEMEEREKVVRHMW 90
+S+ LD + DDL+ L+ + +RH W
Sbjct: 400 VSNQLDMKLRDDLERLKKIRNHRGLRHYW 486
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.318 0.133 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,924,054
Number of Sequences: 36976
Number of extensions: 57445
Number of successful extensions: 374
Number of sequences better than 10.0: 44
Number of HSP's better than 10.0 without gapping: 357
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 372
length of query: 161
length of database: 9,014,727
effective HSP length: 88
effective length of query: 73
effective length of database: 5,760,839
effective search space: 420541247
effective search space used: 420541247
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 54 (25.4 bits)
Medicago: description of AC147178.10


