

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147013.3 + phase: 0
(380 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
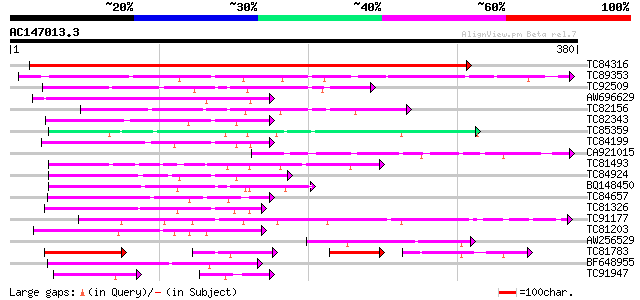
Sequences producing significant alignments: (bits) Value
TC84316 weakly similar to GP|11994612|dbj|BAB02749. gb|AAC24186.... 615 e-176
TC89353 weakly similar to GP|9279704|dbj|BAB01261.1 gb|AAF25964.... 267 6e-72
TC92509 weakly similar to PIR|T49129|T49129 hypothetical protein... 111 5e-25
AW696629 102 2e-22
TC82156 similar to GP|20197985|gb|AAM15340.1 F-box protein famil... 97 8e-21
TC82343 similar to GP|10177628|dbj|BAB10775. gb|AAF30317.1~gene_... 96 2e-20
TC85359 weakly similar to PIR|C96592|C96592 hypothetical protein... 96 3e-20
TC84199 similar to GP|9294656|dbj|BAB03005.1 gb|AAF30317.1~gene_... 95 4e-20
CA921015 homologue to SP|Q8Y7G1|DPO3 DNA polymerase III polC-typ... 93 2e-19
TC81493 weakly similar to PIR|F86249|F86249 protein F25C20.23 [i... 91 6e-19
TC84924 weakly similar to PIR|F96545|F96545 hypothetical protein... 89 2e-18
BQ148450 similar to GP|10177464|dbj gb|AAD21700.1~gene_id:MQB2.1... 89 4e-18
TC84657 weakly similar to GP|9294656|dbj|BAB03005.1 gb|AAF30317.... 87 1e-17
TC81326 weakly similar to PIR|T45651|T45651 hypothetical protein... 86 2e-17
TC91177 similar to GP|9294656|dbj|BAB03005.1 gb|AAF30317.1~gene_... 85 4e-17
TC81203 weakly similar to GP|10177464|dbj|BAB10855. gb|AAD21700.... 84 7e-17
AW256529 homologue to GP|496669|emb|C C-169 protein {Saccharomyc... 78 6e-15
TC81783 similar to GP|9294656|dbj|BAB03005.1 gb|AAF30317.1~gene_... 52 1e-14
BF648955 similar to GP|18043667|gb| Unknown (protein for IMAGE:3... 75 4e-14
TC91947 similar to GP|8096410|dbj|BAA95880.1 hypothetical protei... 49 6e-14
>TC84316 weakly similar to GP|11994612|dbj|BAB02749.
gb|AAC24186.1~gene_id:MGL6.4~strong similarity to
unknown protein {Arabidopsis thaliana}, partial (10%)
Length = 892
Score = 615 bits (1585), Expect = e-176
Identities = 294/296 (99%), Positives = 295/296 (99%)
Frame = +3
Query: 14 PVAETTANKPLPFLPEELIVIILLRLPVRSLLRFKCVCKSWKTLFSDTHFANNHFLISTV 73
PVAETTANKPLPFLPEELIVIILLRLPVRSLLRFKCVCKSWKTLFSDTHFANNHFLISTV
Sbjct: 3 PVAETTANKPLPFLPEELIVIILLRLPVRSLLRFKCVCKSWKTLFSDTHFANNHFLISTV 182
Query: 74 YPQLVACESVSAYRTWEIKTYPIESLLENSSTTVIPVSNTGHQRYTILGSCNGFLCLYDN 133
YPQLVACESVSAYRTWEIKTYPIESLLENSSTTVIPVSNTGHQRYTILGSCNGFLCLYDN
Sbjct: 183 YPQLVACESVSAYRTWEIKTYPIESLLENSSTTVIPVSNTGHQRYTILGSCNGFLCLYDN 362
Query: 134 YQRCVRLWNPSINLKSKSSPTIDRFIYYGFGYDQVNHKYKLLAVKAFSRITETMIYTFGE 193
YQRCVRLWNPSINLKSKSSPTIDRFIYYGFGYDQVNHKYKLLAVKAFSRITETMIYTFGE
Sbjct: 363 YQRCVRLWNPSINLKSKSSPTIDRFIYYGFGYDQVNHKYKLLAVKAFSRITETMIYTFGE 542
Query: 194 NSCKNVEVKDFPRYPPNRKHLGKFVSGTLNWIVDERDGRATILSFDIEKETYRQVLLPQH 253
NSCKNVEVKDFPRYPPNRKHLGKFVSGTLNWIVDERDGRATILSFDIEKETYRQVLLPQH
Sbjct: 543 NSCKNVEVKDFPRYPPNRKHLGKFVSGTLNWIVDERDGRATILSFDIEKETYRQVLLPQH 722
Query: 254 GYAVYSPGLYVLSNCICVCTSFLDTRWQLWMMKKYGVAESWTKLMSIPHENLLISN 309
GYAVYSPGLYVLSNCICVCTSFLDTRWQLWMMKKYGVAESWTKLMSIPHENL IS+
Sbjct: 723 GYAVYSPGLYVLSNCICVCTSFLDTRWQLWMMKKYGVAESWTKLMSIPHENLFISD 890
>TC89353 weakly similar to GP|9279704|dbj|BAB01261.1
gb|AAF25964.1~gene_id:MYA6.2~similar to unknown protein
{Arabidopsis thaliana}, partial (10%)
Length = 1482
Score = 267 bits (682), Expect = 6e-72
Identities = 183/393 (46%), Positives = 229/393 (57%), Gaps = 21/393 (5%)
Frame = +3
Query: 7 NNTDSTLPVAETTANKPLPFLPEELIVIILLRLPVRSLLRFKCVCKSWKTLFSDTHFANN 66
N T T V+E TA+ +PEE+IV ILLRLPVRSLL+F+CVCK WKTL SD FA
Sbjct: 60 NVTVFTQSVSEITAD-----MPEEIIVEILLRLPVRSLLQFRCVCKLWKTLISDPQFAKK 224
Query: 67 HFLISTVYPQLVACESVSAYRTWEIKTYPIESLLENSSTTVIPVSN---TGHQRYTILGS 123
H IST YPQLV+ + + +YP++ LL+N S + ++ TI+GS
Sbjct: 225 HVSISTAYPQLVSV--FVSIAKCNLVSYPLKPLLDNPSAHRVEPADFEMIHTTSMTIIGS 398
Query: 124 CNGFLCLYDNYQRCVRLWNPSINLKSKSSPTI--------DRFIYYGFGYDQVNHKYKLL 175
CNG LCL D YQ LWNPSI LKSK SPTI RF+Y GFGYDQVN +YK+L
Sbjct: 399 CNGLLCLSDFYQ--FTLWNPSIKLKSKPSPTIIAFDSFDSKRFLYRGFGYDQVNDRYKVL 572
Query: 176 AVKAFS---RITETMIYTFGENSCKNVEVKDFPRYPP----NRKHLGKFVSGTLNWIVDE 228
AV T+T+IYTFG ++ FP P R +GKFVSG LNWIV +
Sbjct: 573 AVVQNCYNLDETKTLIYTFGGKDWTTIQ--KFPCDPSRCDLGRLGVGKFVSGNLNWIVSK 746
Query: 229 RDGRATILSFDIEKETYRQVLLPQHGYAVYSPGLYVLSNCICVCTSFLD-TRWQLWMMKK 287
+ I+ FDIEKETY ++ LPQ Y + LYV SN I V + T W +WMMK+
Sbjct: 747 K----VIVFFDIEKETYGEMSLPQ-DYGDKNTVLYVSSNRIYVSFDHSNKTHWVVWMMKE 911
Query: 288 YGVAESWTKLMSIPHENLLISNISLWPCVEPLFISENGVVLLMNTSSSQLILYNLNSRG- 346
YGV ESWTKLM IP + L ++ + + LFISE+G VLLM S+L +YNLN+ G
Sbjct: 912 YGVVESWTKLMIIPQDKL--TSPGPYCLSDALFISEHG-VLLMRPQHSKLSVYNLNNDGG 1082
Query: 347 -LYFPRIPPSYLNIYYIEPKLDLHIYHESLLSP 378
Y I + LHIY+ESL+SP
Sbjct: 1083LDYCTTISGQFARY--------LHIYNESLVSP 1157
>TC92509 weakly similar to PIR|T49129|T49129 hypothetical protein F26G5.80 -
Arabidopsis thaliana, partial (8%)
Length = 858
Score = 111 bits (277), Expect = 5e-25
Identities = 83/243 (34%), Positives = 124/243 (50%), Gaps = 20/243 (8%)
Frame = +3
Query: 23 PLPFLPEELIVIILLRLPVRSLLRFKCVCKSWKTLFSDTHFANNHFLISTVYPQLVACES 82
PLP LP EL+ IL RLPV+ L + +C+CKS+ +L SD FA H L S+ P + S
Sbjct: 162 PLPTLPFELVAEILCRLPVKLLXQLRCLCKSFNSLISDPKFAKKH-LHSSTTPHHLILRS 338
Query: 83 VSAYRTWEIKTYPIESLLENSSTTVIPVSNTGHQRYTILG----------SCNGFLCLYD 132
+ + + PI+S+L ST+ +PV T T L SC+G +CL
Sbjct: 339 NNGSGRFALIVSPIQSVL---STSTVPVPQTQLTYPTCLTEEFASPYEWCSCDGIICLTT 509
Query: 133 NYQRCVRLWNPSINLKSKSSPTIDRF-------IYYGFGYDQVNHKYKLLAVKAFSRITE 185
+Y V LWNP IN K K+ P + + FGYD YK+ A+ + T
Sbjct: 510 DYSSAV-LWNPFIN-KFKTLPPLKYISLKRSPSCLFTFGYDPFADNYKVFAITFCVKRTT 683
Query: 186 TMIYTFGENSCKNVEVKDFPRYP--PNRKHLGKFVSGTLNWIVDERDG-RATILSFDIEK 242
++T G +S + +E DFP + P+ G FV+G ++W+ + G + I+S D+E
Sbjct: 684 VEVHTMGTSSWRRIE--DFPSWSFIPDS---GIFVAGYVHWLTYDGPGSQREIVSLDLED 848
Query: 243 ETY 245
E+Y
Sbjct: 849 ESY 857
>AW696629
Length = 663
Score = 102 bits (254), Expect = 2e-22
Identities = 63/174 (36%), Positives = 91/174 (52%), Gaps = 12/174 (6%)
Frame = +2
Query: 16 AETTANKPLPFLPEELIVIILLRLPVRSLLRFKCVCKSWKTLFSDTHFANNHFLISTVYP 75
+E + P P LP +LI+ IL LPV+ L+RF V K WK+L D +FA H S
Sbjct: 35 SEEATSSP-PILPSDLIMQILSWLPVKLLIRFTSVSKHWKSLILDPNFAKLHLQKSPKNT 211
Query: 76 QLVACESVSAYRTWEIKTYPIES-LLENSSTTVIPVSNTGHQRYTILGSCNGFLCL---- 130
++ TW + YP+ S LLE SS + + Y I+GS NG +CL
Sbjct: 212 HMILTALDDEDDTWVVTPYPVRSLLLEQSSFSDEECCCFDYHSYFIVGSTNGLVCLAVEK 391
Query: 131 ---YDNYQRCVRLWNPSINLKSKSSPTIDRFIY----YGFGYDQVNHKYKLLAV 177
Y+ ++ WNPS+ L+SK +P+++ +Y GFGYD +N YK +AV
Sbjct: 392 SLENRKYELFIKFWNPSLRLRSKKAPSLNIGLYGTARLGFGYDDLNDTYKAVAV 553
>TC82156 similar to GP|20197985|gb|AAM15340.1 F-box protein family AtFBX9
{Arabidopsis thaliana}, partial (4%)
Length = 1136
Score = 97.4 bits (241), Expect = 8e-21
Identities = 76/234 (32%), Positives = 116/234 (49%), Gaps = 12/234 (5%)
Frame = +3
Query: 48 KCVCKSWKTLFSDTHFANNHFLISTVYPQLVACESVSAYRTWEIKTYPIESLLENSSTTV 107
K V KSWK+L SD++F + +ST +L+ + + T + SL+ T
Sbjct: 459 KSVSKSWKSLISDSNFTKKNLRVSTTSHRLLFPKLTKGQYIFNACT--LSSLITTKGTAT 632
Query: 108 IPVSNTGHQRYT-ILGSCNGFLCLYDNYQRCVRLWNPSINLKSKSSPTID----RFIYYG 162
+++ I GSC+G LCL + +QR LWNP IN K S P ++ IY
Sbjct: 633 AMQHPLNIRKFDKIRGSCHGILCL-ELHQRFAILWNPFIN-KYASLPPLEIPWSNTIYSC 806
Query: 163 FGYDQVNHKYKLLAVKAF---SRITETMIYTFGENSCKNVEVKDFPRYPPNRKHLGKFVS 219
FGYD YK+ A + S I +T ++T G S + ++ DFP P + GKFVS
Sbjct: 807 FGYDHSTDSYKVAAFIKWMPNSEIYKTYVHTMGTTSWRMIQ--DFPCTPYLKS--GKFVS 974
Query: 220 GTLNWIVDERD---GRATILSFDIEKETYRQVLLPQHGYA-VYSPGLYVLSNCI 269
T NW+ + ++S +E E+Y ++L P +G V S L+VL +C+
Sbjct: 975 WTFNWLAYKDKYSVSSLLVVSLHLENESYGEILQPDYGSVDVLSISLWVLRDCL 1136
>TC82343 similar to GP|10177628|dbj|BAB10775.
gb|AAF30317.1~gene_id:K6M13.17~similar to unknown
protein {Arabidopsis thaliana}, partial (9%)
Length = 666
Score = 96.3 bits (238), Expect = 2e-20
Identities = 57/160 (35%), Positives = 87/160 (53%), Gaps = 7/160 (4%)
Frame = +2
Query: 25 PFLPEELIVIILLRLPVRSLLRFKCVCKSWKTLFSDTHFANNHFLISTVYPQLVACESVS 84
P+LP+ELI IL+RLPV+SL+RFK VCKSW +L SD HFAN+HF ++ L +
Sbjct: 59 PYLPDELITKILVRLPVKSLIRFKSVCKSWFSLISDNHFANSHFQVTAATHTL---RILF 229
Query: 85 AYRTWEIKTYPIESLLENSSTTVIPVS-NTGHQRY----TILGSCNGFLCLYDNYQRCVR 139
T E + ++SL + +P++ N + I SC GF+ ++ +
Sbjct: 230 LTATPEFRYIAVDSLFTDDYNEPVPLNPNFPLPEFEFDLEIKASCRGFIYVHTCSE--AY 403
Query: 140 LWNPSINLKSK--SSPTIDRFIYYGFGYDQVNHKYKLLAV 177
+WNPS + P + I+YGFGYD+ Y +++V
Sbjct: 404 IWNPSTGFLKQIPFPPNVSNLIFYGFGYDESTDDYLVVSV 523
>TC85359 weakly similar to PIR|C96592|C96592 hypothetical protein T7N22.2
[imported] - Arabidopsis thaliana, partial (7%)
Length = 1955
Score = 95.5 bits (236), Expect = 3e-20
Identities = 90/361 (24%), Positives = 147/361 (39%), Gaps = 72/361 (19%)
Frame = +2
Query: 27 LPEELIVIILLRLPVRSLLRFKCVCKSWKTLFSDTHFAN--------------------- 65
LP +L ILL+LP++SLL +CVCK W TL S+ HFA
Sbjct: 512 LPSQLTTHILLKLPIKSLLICRCVCKIWNTLISEPHFAKLQFERAPVSFVIRNLDNIGVS 691
Query: 66 ------------------NHFLISTVYPQLVACESVSAYRTWEIKTYPIESLLENSSTTV 107
NH + ++ +L C+ +S+ + K Y + + + S
Sbjct: 692 RNLYLLECEAEKFEIGSKNHVKLDPIF-ELPLCKDISSRDKNDAKFYKV--IKKKKSKIR 862
Query: 108 IPVSNTGHQRYTILGSCNGFLCLYD-NYQRCVRLWNP---SINLKSKSSPTIDRF----- 158
+ ++ I+ SCNG LCL + + + + NP + + + T D F
Sbjct: 863 YFTLTSSRDKFGIVNSCNGLLCLSETSIGSPLVICNPVTREFTILPELTTTSDWFNRARA 1042
Query: 159 -IYYGFGYDQVNHKYKLLAV----------KAFSRITETMIYTFGENSCKNVEVKDFPRY 207
+ GFG+ ++YK++ + F R+ I+T G S + VEV P+
Sbjct: 1043RVQAGFGFQPKTNEYKVIIMWNKYVRRNNRLVFERVV-LEIHTLGTPSWRKVEVD--PQI 1213
Query: 208 PPNRKHLGKFVSGTLNWIVDERDGRATILSFDIEKETYRQVLLPQHGYAVYSPG------ 261
+ V+G L+WI+ E + +IL F+ E E + P H + + G
Sbjct: 1214SFLKLLNPTCVNGALHWIIFETGQQKSILCFNFESERLQSFPSPPHVFGNHDNGFPLSMP 1393
Query: 262 --LYVLSNCICVCTSFLDTRWQLWMMKKYGVAESWTKLMSIPHENLLISNISL-----WP 314
L L + +C +W+M +YG+ ESWT + SI LL+ L W
Sbjct: 1394IRLGELKGFLYICHISSLENVTMWVMNEYGIGESWTIVYSIDTSLLLMPGTCLGYPDPWR 1573
Query: 315 C 315
C
Sbjct: 1574C 1576
>TC84199 similar to GP|9294656|dbj|BAB03005.1
gb|AAF30317.1~gene_id:F14O13.6~similar to unknown
protein {Arabidopsis thaliana}, partial (14%)
Length = 673
Score = 95.1 bits (235), Expect = 4e-20
Identities = 64/166 (38%), Positives = 91/166 (54%), Gaps = 10/166 (6%)
Frame = +1
Query: 22 KPLPFLPEELIVIILLRLPVRSLLRFKCVCKSWKTLFSDTHFANNHFLISTVYPQLVACE 81
K P+LP ELI+ ILL LPV+SLLRFKCVCKSW +L SDTHFAN+HF I+ + + V
Sbjct: 94 KTAPYLPLELIIQILLWLPVKSLLRFKCVCKSWFSLISDTHFANSHFQITAKHSRRVLFM 273
Query: 82 SVSAYRTWEIKTYPIESLLENSSTTVIP------VSNTGHQRYTILGSCNGFLCLYDNYQ 135
T + E+L +++ + IP V T SC GF+ L+++
Sbjct: 274 LNHVPTTLSL---DFEALHCDNAVSEIPNPIPNFVEPPCDSLDTNSSSCRGFIFLHNDPD 444
Query: 136 RCVRLWNPSINLKSK--SSPTIDRFIY--YGFGYDQVNHKYKLLAV 177
+ +WNPS + + SP + YGFGYDQ+ Y +++V
Sbjct: 445 --LFIWNPSTRVYKQIPLSPNDSNSFHCLYGFGYDQLRDDYLVVSV 576
>CA921015 homologue to SP|Q8Y7G1|DPO3 DNA polymerase III polC-type (EC
2.7.7.7) (PolIII). {Listeria monocytogenes}, partial
(0%)
Length = 824
Score = 92.8 bits (229), Expect = 2e-19
Identities = 72/223 (32%), Positives = 114/223 (50%), Gaps = 7/223 (3%)
Frame = -1
Query: 163 FGYDQVNHKYKLLAVKAFSRITETMIYTFGENSCKNVEVKDFPRYPPNRKHLGKFVSGTL 222
FGYD +++ V + ++ +YT G + K +E P N G FVSGT+
Sbjct: 821 FGYDHFIDNIRVMFVLPKNEVS---VYTLGTDYWKRIE-----DIPYNIFEEGVFVSGTV 666
Query: 223 NWIVDERDGRATILSFDIEKETYRQVLLPQHGYAVYSPGLYVLSNCICVCTS---FLDTR 279
NW+ + + ILS D+EKE+Y+QVLLP ++ G VL NC+C+ + FLD
Sbjct: 665 NWLASDD---SFILSLDVEKESYQQVLLPDTENDLWILG--VLRNCLCILATSNLFLD-- 507
Query: 280 WQLWMMKKYGVAESWTKLMSIPHENLLISNISLWPCVEPLFISENGVVLL----MNTSSS 335
+W+M +YG ESWTKL S+P N+ + + L+ SE+ +LL M +
Sbjct: 506 --VWIMNEYGNQESWTKLYSVP--NMQDHGLEAYTV---LYSSEDDQLLLEFNEMRSDKV 348
Query: 336 QLILYNLNSRGLYFPRIPPSYLNIYYIEPKLDLHIYHESLLSP 378
+L++Y+ + L P +Y ++ + Y ESL+SP
Sbjct: 347 KLVVYDSKTGTLNIPEFQNNY-------DQICQNFYIESLISP 240
>TC81493 weakly similar to PIR|F86249|F86249 protein F25C20.23 [imported] -
Arabidopsis thaliana, partial (9%)
Length = 728
Score = 91.3 bits (225), Expect = 6e-19
Identities = 75/249 (30%), Positives = 114/249 (45%), Gaps = 24/249 (9%)
Frame = +1
Query: 27 LPEELIVIILLRLPVRSLLRFKCVCKSWKTLFSDTHFANNHFLISTVYPQLVACESVSAY 86
LP E+ IL R+P + LLR + CK W+ L T F H +S ++ S
Sbjct: 16 LPTEVTTEILSRVPAKPLLRLRSTCKWWRNLIDSTDFIFLH--LSKSRDSVIILRQHS-- 183
Query: 87 RTWEIKTYPIESLLENSSTTVIPVSNTGHQRYTILGSCNGFLCLYDNYQRCVRLWNPS-- 144
R +E+ ++ + E ++ SN R +LGSCNG LC+ N + WNP+
Sbjct: 184 RLYELDLNSMDRVKE-LDHPLMCYSN----RIKVLGSCNGLLCIC-NIADDIAFWNPTIR 345
Query: 145 ----------INLKSKSSPTIDRFI---YYGFGYDQVNHKYKLLAVKAF------SRITE 185
I ++ + TI + YGFGYD YKL+++ F S +
Sbjct: 346 KHRIIPSEPLIRKETNENNTITTLLAAHVYGFGYDSATDDYKLVSISNFVDLHNRSYDSH 525
Query: 186 TMIYTFGENSCKNVEVKDFPRYPPNRKHLGKFVSGTLNWIVD---ERDGRATILSFDIEK 242
IYT G + + + P + +G FVSG L+W+V E D R I++FD+
Sbjct: 526 VTIYTMGSDVW--MPLPGVPYALCCARPMGVFVSGALHWVVPRALEPDSRDLIVAFDLRF 699
Query: 243 ETYRQVLLP 251
E +R+V LP
Sbjct: 700 EVFREVALP 726
>TC84924 weakly similar to PIR|F96545|F96545 hypothetical protein F8A12.9
[imported] - Arabidopsis thaliana, partial (9%)
Length = 575
Score = 89.4 bits (220), Expect = 2e-18
Identities = 59/172 (34%), Positives = 94/172 (54%), Gaps = 9/172 (5%)
Frame = +1
Query: 27 LPEELIVIILLRLPVRSLLRFKCVCKSWKTLFSDTHFANNHFLIS-TVYPQLVACESVSA 85
LP+ELI+ LLRLPV+SLL FKC+CK W ++ SD HFAN+HF ++ + + C S +
Sbjct: 61 LPQELIIQFLLRLPVKSLLVFKCICKLWFSIISDPHFANSHFQLNHAKHTRRFLCISALS 240
Query: 86 YRTWEIKTYPIESLLENSSTTVIPVSN----TGHQRYTILGSCNGFLCLYDNYQRCVRLW 141
EI++ ++ L ++ + P N + + I GSC GF+ +Y + + +W
Sbjct: 241 P---EIRSIDFDAFLNDAPAS--PNFNCSLPDSYFPFEIKGSCRGFIFMYRHPN--IYIW 399
Query: 142 NPSINLKSK---SSPTIDRFI-YYGFGYDQVNHKYKLLAVKAFSRITETMIY 189
NPS K + S+ +I YGFGYDQ Y ++ + S I +++
Sbjct: 400 NPSTGSKRQILMSAFNTKAYINLYGFGYDQSRDDYVVVFIVQ*SLIHSPLVF 555
>BQ148450 similar to GP|10177464|dbj gb|AAD21700.1~gene_id:MQB2.18~similar to
unknown protein {Arabidopsis thaliana}, partial (9%)
Length = 666
Score = 88.6 bits (218), Expect = 4e-18
Identities = 67/197 (34%), Positives = 100/197 (50%), Gaps = 18/197 (9%)
Frame = +3
Query: 27 LPEELIVIILLRLPVRSLLRFKCVCKSWKTLFSDTHFANNHFLIS-TVYPQLVACESVSA 85
LP E+ V IL RLPV+ L++F+CVCK WK+ S F H +S T + L+ +S
Sbjct: 3 LPFEIQVEILSRLPVKYLMQFQCVCKLWKSQISKPDFVKKHLRVSNTRHLFLLTFSKLSP 182
Query: 86 YRTWEIKTYPIESLLENSSTTVIPVS---NTGHQRYTILGSCNGFLCLYDNYQRCVRLWN 142
IK+YP+ S+ + T + N + +++GSC+G LC+ N V LWN
Sbjct: 183 ELV--IKSYPLSSVFTEMTPTFTQLEYPLNNRDESDSMVGSCHGILCIQCNLSFPV-LWN 353
Query: 143 PSINLKSKSSPTID----RFI--YYGFGYDQVNHKYKLLAVKAFSRI--------TETMI 188
PSI K P+ + +FI Y FGYD + YK++AV S I T +
Sbjct: 354 PSIR-KFTKLPSFEFPQNKFINPTYAFGYDHSSDTYKVVAVFCTSNIDNGVYQLKTLVNV 530
Query: 189 YTFGENSCKNVEVKDFP 205
+T G N + ++ +FP
Sbjct: 531 HTMGTNCWRRIQT-EFP 578
>TC84657 weakly similar to GP|9294656|dbj|BAB03005.1
gb|AAF30317.1~gene_id:F14O13.6~similar to unknown
protein {Arabidopsis thaliana}, partial (14%)
Length = 884
Score = 86.7 bits (213), Expect = 1e-17
Identities = 62/167 (37%), Positives = 89/167 (53%), Gaps = 15/167 (8%)
Frame = +1
Query: 26 FLPEELIVIILLRLPVRSLLRFKCVCKSWKTLFSDTHFANNHFLIST-VYPQLVACESVS 84
+LP ELI+ I+LRLPV+SL+RFKCVCKS L SD +FA +HF +ST + + S
Sbjct: 325 YLPHELIIQIMLRLPVKSLIRFKCVCKSLLALISDHNFAKSHFELSTATHTNRIVFMSTL 504
Query: 85 AYRTWEIKTYPIESLLENSSTTVIPVSNTGHQRYT---ILGSCNGFLCLYDNYQRCVRLW 141
A T I SL ++S++T + ++ + Y+ I SC GF+ L + LW
Sbjct: 505 ALETRSIDFE--ASLNDDSASTSLNLNFMPPESYSSLEIKSSCRGFIVL--TCSSNIYLW 672
Query: 142 NPSI-----------NLKSKSSPTIDRFIYYGFGYDQVNHKYKLLAV 177
NPS NL +K S + YGFGYD + Y +++V
Sbjct: 673 NPSTGHHKQIPFPASNLDAKYSCCL-----YGFGYDHLRDDYLVVSV 798
>TC81326 weakly similar to PIR|T45651|T45651 hypothetical protein F13I12.200
- Arabidopsis thaliana, partial (12%)
Length = 838
Score = 85.9 bits (211), Expect = 2e-17
Identities = 57/156 (36%), Positives = 79/156 (50%), Gaps = 7/156 (4%)
Frame = +3
Query: 24 LPFLPEELIVIILLRLPVRSLLRFKCVCKSWKTLFSDTHFANNHFLISTVYPQLVACESV 83
LP+LP ELI ILLRLPV+SL RFK V KSW +L S HFAN+HF +++ +
Sbjct: 159 LPYLPHELIFQILLRLPVKSLTRFKSVRKSWFSLISAPHFANSHFQLTSAKHAASRIMFI 338
Query: 84 SAYRTWEIKTYPIESLLENSSTTVIPVS---NTGHQRYTILGSCNGFLCLYDNYQRCVRL 140
S + E ++ ++ L + + ++ H I GSC GF+ LY + +
Sbjct: 339 STL-SHETRSIDFKAFLNDDDPASLNITFSLTRSHFPVEIRGSCRGFILLYRPPD--IYI 509
Query: 141 WNPSINLKS--KSSPTIDRFI--YYGFGYDQVNHKY 172
WNPS K SP + + GFGYDQ Y
Sbjct: 510 WNPSTGFKKHIHLSPVDSKSVAQCQGFGYDQSRDDY 617
>TC91177 similar to GP|9294656|dbj|BAB03005.1
gb|AAF30317.1~gene_id:F14O13.6~similar to unknown
protein {Arabidopsis thaliana}, partial (9%)
Length = 1298
Score = 85.1 bits (209), Expect = 4e-17
Identities = 92/369 (24%), Positives = 159/369 (42%), Gaps = 38/369 (10%)
Frame = +2
Query: 47 FKCVCKSWKTLFSDTHFANNHFLISTV-----------YPQLVACE-SVSAYRTWEIKTY 94
FKCVCK W +L S HFAN+HF ++T Y Q ++ + +S + T
Sbjct: 74 FKCVCKLWLSLISQPHFANSHFQLTTATHTNRIMLITPYLQSLSIDLELSLNDDSAVYTT 253
Query: 95 PIESLLEN----SSTTVIPVSNTGHQRYTIL---GSCNGFLCLYDNYQRCVRLWNPSINL 147
I L+++ SS++ + + + IL GSC GF+ L C+ WNPS
Sbjct: 254 DISFLIDDEDYYSSSSSDMDDLSPPKSFFILDFKGSCRGFILLNCYSSLCI--WNPSTGF 427
Query: 148 KSKSSPTI-----DRFIYYGFGYDQVNHKYKLLAVK------AFSRITETMIYTFGENSC 196
+ T D +YGFGYD+ Y ++++ + ++ I++ N
Sbjct: 428 HKRIPFTTIDSNPDANYFYGFGYDESTDDYLVISMSYEPSPSSDGMLSHLGIFSLRANVW 607
Query: 197 KNVEVKDFPRYPPNR--KHLGKFVSGTLNWIVDERD-GRATILSFDIEKETYRQVLLPQH 253
VE + Y N + +G ++W+ D I++F + + ++ LP
Sbjct: 608 TRVEGGNLLLYSQNSLLNLVESLSNGAIHWLAFRNDISMPVIVAFHLMERKLLELRLPNE 787
Query: 254 GYAVYSPG----LYVLSNCICVCTSFLD-TRWQLWMMKKYGVAESWTKLMSIPHENLLIS 308
+ P L+V C+ + D +Q+W+M+KY V SWTK + + +
Sbjct: 788 --IINGPSRAYDLWVYRGCLALWHILPDRVTFQIWVMEKYNVQSSWTKTLVLSFDGNPAH 961
Query: 309 NISLWPCVEPLFISENGVVLLMNTSSSQLILYNLNSRGLYFPRIPPSYLNIYYIEPKLDL 368
S W P + +++G ++ N + L N +G + SY + Y+ P +
Sbjct: 962 --SFW----PKYYTKSGDIVGRN---MRCALAKYNDKGQL--QEHHSYCDSQYVSPVV-- 1102
Query: 369 HIYHESLLS 377
+Y ESLLS
Sbjct: 1103-MYTESLLS 1126
Score = 33.5 bits (75), Expect = 0.14
Identities = 18/55 (32%), Positives = 30/55 (53%)
Frame = +3
Query: 26 FLPEELIVIILLRLPVRSLLRFKCVCKSWKTLFSDTHFANNHFLISTVYPQLVAC 80
+LP ELI+ ILLRLPV+SL+R L+ + + F + ++ ++ C
Sbjct: 12 YLPFELIIQILLRLPVKSLIRLNAFVSYGYLLYLNLTLQIHIFNLPPLHTRIELC 176
>TC81203 weakly similar to GP|10177464|dbj|BAB10855.
gb|AAD21700.1~gene_id:MQB2.18~similar to unknown protein
{Arabidopsis thaliana}, partial (9%)
Length = 715
Score = 84.3 bits (207), Expect = 7e-17
Identities = 62/183 (33%), Positives = 86/183 (46%), Gaps = 27/183 (14%)
Frame = +1
Query: 17 ETTANKPLPFLPEELIVIILLRLPVRSLLRFKCVCKSWKTLFS-DTHFANNHFLIS--TV 73
++ P L +ELIV IL RLPV++L++FKCVCKSWKTL S D FA H S
Sbjct: 169 QSNGGAPTSLLLDELIVDILSRLPVKTLMQFKCVCKSWKTLISHDPSFAKLHLQRSPRNT 348
Query: 74 YPQLVACESVSAYRTWEIKTYPIESLLENSSTTVIP---------VSNTGHQRYT----- 119
+ LV+ S + + +P+ L+E + T+ P V+ Y
Sbjct: 349 HLTLVSDRSSDDESNFSVVPFPVSHLIE-APLTITPFEPYHLLRNVAIPDDPYYVLGNMD 525
Query: 120 ---ILGSCNGFLCL------YDNYQR-CVRLWNPSINLKSKSSPTIDRFIYYGFGYDQVN 169
I+GSCNG CL Y+ + R WNP+ N S+ ++ F FGYD N
Sbjct: 526 CCIIIGSCNGLYCLRCYSLIYEEEEHDWFRFWNPATNTLSEELGCLNEFFRLTFGYDISN 705
Query: 170 HKY 172
Y
Sbjct: 706 DTY 714
>AW256529 homologue to GP|496669|emb|C C-169 protein {Saccharomyces
cerevisiae}, partial (7%)
Length = 624
Score = 77.8 bits (190), Expect = 6e-15
Identities = 41/120 (34%), Positives = 68/120 (56%), Gaps = 7/120 (5%)
Frame = +1
Query: 200 EVKDFPRYPPNRKHLGKFVSGTLNWI----VDERDGRATILSFDIEKETYRQVLLPQHGY 255
+++DFP G FVSGT+NW+ +D+ + TI+S D+ KE Y+++ P +G
Sbjct: 1 KIEDFPSLMIPYSQHGIFVSGTVNWLADYDLDDNNSLGTIVSLDLRKEIYQEISQPDYGD 180
Query: 256 AVYSPGLYVLSNCICVCTSFLDTRWQLWMMKKYGVAESWTKLMSIP---HENLLISNISL 312
L + C+CV S D+ +W+MK+YG ESW KL+ IP + +L+ N+++
Sbjct: 181 VTVKLSLGAMRECLCVF-SHSDSFDDVWLMKEYGNEESWIKLIRIPCISNRSLVFDNVNI 357
>TC81783 similar to GP|9294656|dbj|BAB03005.1
gb|AAF30317.1~gene_id:F14O13.6~similar to unknown
protein {Arabidopsis thaliana}, partial (6%)
Length = 1225
Score = 52.4 bits (124), Expect = 3e-07
Identities = 24/55 (43%), Positives = 37/55 (66%)
Frame = +2
Query: 24 LPFLPEELIVIILLRLPVRSLLRFKCVCKSWKTLFSDTHFANNHFLISTVYPQLV 78
LP LP +L+ IL RLPV+ L++ +C+CK + +L SD +FA HF S + +L+
Sbjct: 125 LPTLPFDLLPEILCRLPVKLLIQLRCLCKFFNSLISDPNFAKKHFQFSAKHHRLM 289
Score = 45.1 bits (105), Expect(3) = 1e-14
Identities = 31/94 (32%), Positives = 50/94 (52%), Gaps = 7/94 (7%)
Frame = +2
Query: 264 VLSNCICVCTSFLDTRWQLWMMKKYGVAESWTKLMSIP---HENLLISNISLWPCVEPLF 320
+L +C+CV + D + +W+MK+YG ESWTKL S+P H IS +
Sbjct: 875 MLRDCLCVYEAS-DLFFNVWIMKEYGNQESWTKLYSVPKMQHRRFKAYTIS--------Y 1027
Query: 321 ISENGVVLL----MNTSSSQLILYNLNSRGLYFP 350
I E+ +LL M +S+++L +Y+ + L P
Sbjct: 1028IYEDDQLLLGIVDMQSSNTKLAVYDSKTGTLNIP 1129
Score = 39.7 bits (91), Expect(3) = 1e-14
Identities = 17/37 (45%), Positives = 25/37 (66%)
Frame = +1
Query: 215 GKFVSGTLNWIVDERDGRATILSFDIEKETYRQVLLP 251
G FV GT+NW+ + ILS D+EKE+Y+++ LP
Sbjct: 715 GIFVGGTVNWMASDIADLLFILSLDLEKESYQKLFLP 825
Score = 31.6 bits (70), Expect(3) = 1e-14
Identities = 20/61 (32%), Positives = 27/61 (43%), Gaps = 4/61 (6%)
Frame = +3
Query: 123 SCNGFLCLYDNYQRCVRLWNPSIN----LKSKSSPTIDRFIYYGFGYDQVNHKYKLLAVK 178
SC+G C N C +WNPSI L +P FGYD YK++A+
Sbjct: 447 SCDGIFCGELNLGYCF-VWNPSIRKFKLLPPYKNPLEGDPFSISFGYDHFIDNYKVVAIS 623
Query: 179 A 179
+
Sbjct: 624 S 626
>BF648955 similar to GP|18043667|gb| Unknown (protein for IMAGE:3661715) {Mus
musculus}, partial (3%)
Length = 514
Score = 75.1 bits (183), Expect = 4e-14
Identities = 54/152 (35%), Positives = 70/152 (45%), Gaps = 8/152 (5%)
Frame = +3
Query: 26 FLPEELIVIILLRLPVRSLLRFKCVCKSWKTLFSDTHFANNHFLISTVYPQLVACESVSA 85
FLP ELIV I+ LPV+ L++F+CV K +KTL SD +F H S P L
Sbjct: 54 FLPSELIVEIISWLPVKYLMQFRCVSKFYKTLISDPYFVQMHLEKSARNPHLALMWQDDL 233
Query: 86 YR-TWEIKTYPIESLLENSSTTVIPVSNTGHQRYTILGSCNGFLCLYD------NYQRCV 138
R I + LL N TT P + I+GSCNG LCL D Y +
Sbjct: 234 LREDGSIIFLSVSRLLGNKYTTP-PFQSGTFYHSCIIGSCNGLLCLVDFHCPDYYYYYYL 410
Query: 139 RLWNPSINLK-SKSSPTIDRFIYYGFGYDQVN 169
WNP+ K T+ + FGYD ++
Sbjct: 411 YFWNPATRTKFXNILITLSXDFKFSFGYDTLS 506
>TC91947 similar to GP|8096410|dbj|BAA95880.1 hypothetical protein {Oryza
sativa (japonica cultivar-group)}, partial (6%)
Length = 872
Score = 49.3 bits (116), Expect(2) = 6e-14
Identities = 29/66 (43%), Positives = 36/66 (53%), Gaps = 7/66 (10%)
Frame = +1
Query: 30 ELIVIILLRLPVRSLLRFKCVCKSWKTLFSDTHFANNHFL-------ISTVYPQLVACES 82
E+IV IL RLPVRS+L++K VCKSWKTL + T F I + Q + C S
Sbjct: 16 EIIVEILSRLPVRSILQYKRVCKSWKTLLTSTAAYPRFFCSNGGRREIISYQTQSIHCSS 195
Query: 83 VSAYRT 88
Y T
Sbjct: 196TPIYTT 213
Score = 45.4 bits (106), Expect(2) = 6e-14
Identities = 25/54 (46%), Positives = 31/54 (57%), Gaps = 4/54 (7%)
Frame = +2
Query: 128 LCLYDNYQRCVRLWNP----SINLKSKSSPTIDRFIYYGFGYDQVNHKYKLLAV 177
LCLY +RLWNP I + SK D ++GFGYDQVN KYK+L +
Sbjct: 221 LCLYHKLLYNIRLWNPFY*IEIQIISK-----DCITHHGFGYDQVNDKYKVLYI 367
Score = 38.9 bits (89), Expect = 0.003
Identities = 35/111 (31%), Positives = 48/111 (42%), Gaps = 3/111 (2%)
Frame = +3
Query: 143 PSINLKSKSSPTIDRFIYYGFGYDQVNHKYKLLAVKAFSRITETMIYTFGENSCKNVEVK 202
PSI LKSK SP I V+H L+ +K I + C N +
Sbjct: 267 PSIELKSKLSPKI------------VSHIMVLVMIKLTISIRCCTL-------CSNSGIG 389
Query: 203 DFPRYPPNRKHLGKFVSGTLNWIV---DERDGRATILSFDIEKETYRQVLL 250
+ G +VSGTLNWI+ + I+S D+EKETY +V +
Sbjct: 390 *DNSFEFPV*TSGIYVSGTLNWIIA*DGVNSNQLVIISVDLEKETYGEVFM 542
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.322 0.138 0.435
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,447,857
Number of Sequences: 36976
Number of extensions: 272814
Number of successful extensions: 1822
Number of sequences better than 10.0: 142
Number of HSP's better than 10.0 without gapping: 1728
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1757
length of query: 380
length of database: 9,014,727
effective HSP length: 98
effective length of query: 282
effective length of database: 5,391,079
effective search space: 1520284278
effective search space used: 1520284278
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 59 (27.3 bits)
Medicago: description of AC147013.3


