

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147013.10 - phase: 0 /pseudo
(339 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
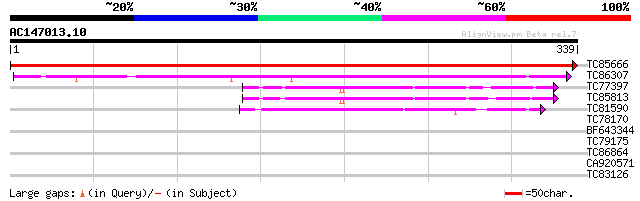
Score E
Sequences producing significant alignments: (bits) Value
TC85666 homologue to SP|P10933|FENR_PEA Ferredoxin--NADP reducta... 681 0.0
TC86307 homologue to PIR|T06773|T06773 ferredoxin--NADP+ reducta... 270 6e-73
TC77397 homologue to PIR|S37159|S37159 NADPH--ferrihemoprotein r... 73 1e-13
TC85813 homologue to GP|6503253|gb|AAC09468.2| putative NADPH-cy... 62 4e-10
TC81590 similar to GP|6041790|gb|AAF02110.1| putative NADPH-ferr... 59 3e-09
TC78170 similar to GP|17979531|gb|AAL50100.1 At1g76150/T23E18_38... 34 0.071
BF643344 33 0.16
TC79175 similar to GP|21592883|gb|AAM64833.1 cytochrome-b5 reduc... 32 0.27
TC86864 similar to PIR|F96570|F96570 unknown protein 80333-8217... 29 2.3
CA920571 similar to GP|12322018|gb unknown protein; 3519-5443 {A... 29 2.3
TC83126 similar to GP|15529155|gb|AAK97672.1 AT3g30390/T6J22_16 ... 28 5.1
>TC85666 homologue to SP|P10933|FENR_PEA Ferredoxin--NADP reductase leaf
isozyme chloroplast precursor (EC 1.18.1.2) (FNR).
[Garden pea], partial (96%)
Length = 1389
Score = 681 bits (1756), Expect = 0.0
Identities = 339/339 (100%), Positives = 339/339 (100%)
Frame = +3
Query: 1 MAAAVTAAVSFPYSNSTSLPIRTSVISPDRLVFKKVSLNNVSISGRLTVRAEVATEAPAP 60
MAAAVTAAVSFPYSNSTSLPIRTSVISPDRLVFKKVSLNNVSISGRLTVRAEVATEAPAP
Sbjct: 90 MAAAVTAAVSFPYSNSTSLPIRTSVISPDRLVFKKVSLNNVSISGRLTVRAEVATEAPAP 269
Query: 61 VKVEKISKKQEEGIVVNKFKPKTPYIGRCLLNTKITGDDAPGETWHMVFSTEGEVPYREG 120
VKVEKISKKQEEGIVVNKFKPKTPYIGRCLLNTKITGDDAPGETWHMVFSTEGEVPYREG
Sbjct: 270 VKVEKISKKQEEGIVVNKFKPKTPYIGRCLLNTKITGDDAPGETWHMVFSTEGEVPYREG 449
Query: 121 QSIGIVPDGIDKNGKPHKLRLYSIASSALGDFGDSKTVSLCVKRLVYTNDAGEVVKGVCS 180
QSIGIVPDGIDKNGKPHKLRLYSIASSALGDFGDSKTVSLCVKRLVYTNDAGEVVKGVCS
Sbjct: 450 QSIGIVPDGIDKNGKPHKLRLYSIASSALGDFGDSKTVSLCVKRLVYTNDAGEVVKGVCS 629
Query: 181 NFLCDLRPGSEVQITGPVGKEMLMPKDPNATVIMLGTGTGIAPFRSFLWKMFFEKHEDYK 240
NFLCDLRPGSEVQITGPVGKEMLMPKDPNATVIMLGTGTGIAPFRSFLWKMFFEKHEDYK
Sbjct: 630 NFLCDLRPGSEVQITGPVGKEMLMPKDPNATVIMLGTGTGIAPFRSFLWKMFFEKHEDYK 809
Query: 241 FNGLAWLFLGVPTSSSLLYKEEFEKMKEKAPENFRLDFAVSREQVNDKGEKMYIQTRMAQ 300
FNGLAWLFLGVPTSSSLLYKEEFEKMKEKAPENFRLDFAVSREQVNDKGEKMYIQTRMAQ
Sbjct: 810 FNGLAWLFLGVPTSSSLLYKEEFEKMKEKAPENFRLDFAVSREQVNDKGEKMYIQTRMAQ 989
Query: 301 YAEELWELLKKDNTFVYMCGLKGMEKGIDDIMVSLAAKD 339
YAEELWELLKKDNTFVYMCGLKGMEKGIDDIMVSLAAKD
Sbjct: 990 YAEELWELLKKDNTFVYMCGLKGMEKGIDDIMVSLAAKD 1106
>TC86307 homologue to PIR|T06773|T06773 ferredoxin--NADP+ reductase (EC
1.18.1.2) - garden pea (fragment), complete
Length = 1648
Score = 270 bits (690), Expect = 6e-73
Identities = 155/357 (43%), Positives = 211/357 (58%), Gaps = 23/357 (6%)
Frame = +3
Query: 3 AAVTAAVSFPYSNSTSLPIRTSVISPDRLVFKKVSL----------------NNVSISGR 46
+A AV+ P SN S R SV + F+ S N ++
Sbjct: 261 SASQMAVTVPVSNDFSA--RRSVFKASNINFRDKSWAPVFALDMKAKNCGWRRNQNVICM 434
Query: 47 LTVRAEVATEAPAPVKVEKISKKQEEGIVVNKFKPKTPYIGRCLLNTKITGDDAPGETWH 106
+A V A +P+++E ++ +N KPK PY + ++ G APGET H
Sbjct: 435 SVQQASVPKVAVSPLELENPAEPP-----LNLHKPKEPYTATIVSVERLVGPKAPGETCH 599
Query: 107 MVFSTEGEVPYREGQSIGIVPDGID--KNGKPHKLRLYSIASSALGDFGDSKTVSLCVKR 164
+V + +G VPY EGQS G++P G + K G PH +RLYSIAS+ GDF D KT SLCV+R
Sbjct: 600 VVINHDGNVPYWEGQSYGVIPPGENPKKPGAPHNVRLYSIASTRYGDFFDGKTASLCVRR 779
Query: 165 LVY----TNDAGEVVKGVCSNFLCDLRPGSEVQITGPVGKEMLMPKD-PNATVIMLGTGT 219
VY T GVCSNFLCD +PG +++ITGP GK ML+P+D PNAT IM+ TGT
Sbjct: 780 AVYYDPETGKEDPSKNGVCSNFLCDSKPGDKIKITGPSGKIMLLPEDDPNATHIMIATGT 959
Query: 220 GIAPFRSFLWKMFFEKHEDYKFNGLAWLFLGVPTSSSLLYKEEFEKMKEKAPENFRLDFA 279
G+AP+R +L +MF E +KF GLAWLFLGV S SLLY +EF K + PENFR D A
Sbjct: 960 GVAPYRGYLRRMFMESVPTFKFGGLAWLFLGVANSDSLLYDDEFTKYLKDYPENFRYDRA 1139
Query: 280 VSREQVNDKGEKMYIQTRMAQYAEELWELLKKDNTFVYMCGLKGMEKGIDDIMVSLA 336
+SRE+ N G KMY+Q ++ +Y++E+++LL + +Y CGLKGM GI + + +A
Sbjct: 1140LSREEKNRNGGKMYVQDKIEEYSDEIFKLL-DNGAHIYFCGLKGMMPGIQETLKRVA 1307
>TC77397 homologue to PIR|S37159|S37159 NADPH--ferrihemoprotein reductase (EC
1.6.2.4) - spring vetch, complete
Length = 2519
Score = 73.2 bits (178), Expect = 1e-13
Identities = 62/196 (31%), Positives = 92/196 (46%), Gaps = 7/196 (3%)
Frame = +2
Query: 140 RLYSIASSALGDFGDSKTVSLCVKRLVYTNDAGEVVKGVCSNFLCDLRPGSEVQITG--P 197
R YSI+SS F + C + G + KGVCS ++ + P E + P
Sbjct: 1490 RYYSISSSPR--FAPQRVHVTCAL-VEGPTPTGRIHKGVCSTWMKNAIPSEESRDCSWAP 1660
Query: 198 V---GKEMLMPKDPNATVIMLGTGTGIAPFRSFLWKMFFEKHEDYKFNGLAWLFLGVPT- 253
+ +P DP+ +IM+G GTG+APFR FL + F K + + G A LF G
Sbjct: 1661 IFIRPSNFKLPADPSIPIIMVGPGTGLAPFRGFLQERFALKEDGVQL-GPALLFFGCRNR 1837
Query: 254 SSSLLYKEEFEKMKEKAPENFRLDFAVSREQVNDKGEKMYIQTRMAQYAEELWELLKKDN 313
+Y+EE E+ + L A SRE EK Y+Q +M A W L+ +
Sbjct: 1838 QMDFIYEEELNNFVEQGSLS-ELIVAFSRE----GPEKEYVQHKMMDKASYFWSLISQGG 2002
Query: 314 TFVYMCG-LKGMEKGI 328
++Y+CG KGM + +
Sbjct: 2003 -YLYVCGDAKGMARDV 2047
>TC85813 homologue to GP|6503253|gb|AAC09468.2| putative NADPH-cytochrome P450
reductase {Pisum sativum}, complete
Length = 2581
Score = 61.6 bits (148), Expect = 4e-10
Identities = 58/197 (29%), Positives = 91/197 (45%), Gaps = 8/197 (4%)
Frame = +3
Query: 140 RLYSIASSALGDFGDSKTVSLCVKRLVYTN-DAGEVVKGVCSNFLCDLRPGSEVQITG-- 196
R YSI+SS S+ C LV+ G + +GVCS ++ + P + Q
Sbjct: 1536 RYYSISSSPR--VAPSRIHVTCA--LVHDKMPTGRIHQGVCSTWMKNSAPLEKSQDCSWA 1703
Query: 197 PV---GKEMLMPKDPNATVIMLGTGTGIAPFRSFLWKMFFEKHEDYKFNGLAWLFLGVPT 253
P+ +P D +IM+G GTG+APFR FL + K E + G + LF G
Sbjct: 1704 PIFVRQSNFRLPADNKVPIIMIGPGTGLAPFRGFLQERLALKEEGAEL-GPSVLFFGCRN 1880
Query: 254 SS-SLLYKEEFEKMKEKAPENFRLDFAVSREQVNDKGEKMYIQTRMAQYAEELWELLKKD 312
+Y++E + L A SRE K Y+Q +M + A ++W ++ +
Sbjct: 1881 RQVDYIYEDELNHFVHGGALS-ELIVAFSREGPT----KEYVQHKMIEKASDIWNMISQ- 2042
Query: 313 NTFVYMCG-LKGMEKGI 328
++Y+CG KGM K +
Sbjct: 2043 GAYIYVCGDAKGMAKDV 2093
>TC81590 similar to GP|6041790|gb|AAF02110.1| putative
NADPH-ferrihemoprotein reductase {Arabidopsis thaliana},
partial (33%)
Length = 1095
Score = 58.5 bits (140), Expect = 3e-09
Identities = 52/190 (27%), Positives = 84/190 (43%), Gaps = 7/190 (3%)
Frame = +3
Query: 138 KLRLYSIASSALGDFGDSKTVSLCVKRLVYTNDAGEVVKGVCSNFLCDLRPGSEVQITGP 197
K R +SI+SS V L V + +T KG+CS++L L P + V I
Sbjct: 27 KTRAFSISSS---QSVHPNQVHLTVSVVSWTTPYKRKKKGLCSSWLATLDPRNAVGIPAW 197
Query: 198 VGKEMLMPKDPNATVIMLGTGTGIAPFRSFLWKMFFEKHEDYKFNGLAWLFLGVPTSSSL 257
K L P+ +I++G GTG APFR F+ + + + + + F
Sbjct: 198 FQKGSLPTASPSLPLILVGPGTGCAPFRGFIEERALQS-KTISTAPIMFFFGCWNEDGDF 374
Query: 258 LYKEEFEK-------MKEKAPENFRLDFAVSREQVNDKGEKMYIQTRMAQYAEELWELLK 310
LYK+ + + E F + F+ D+ EK+Y+Q +M +++ +W LL
Sbjct: 375 LYKDLWLNHSQNNGVLSEANGGGFYVAFS------RDQPEKVYVQHKMKEHSGRVWNLL- 533
Query: 311 KDNTFVYMCG 320
D VY+ G
Sbjct: 534 ADGAAVYIAG 563
>TC78170 similar to GP|17979531|gb|AAL50100.1 At1g76150/T23E18_38
{Arabidopsis thaliana}, partial (90%)
Length = 955
Score = 34.3 bits (77), Expect = 0.071
Identities = 18/70 (25%), Positives = 36/70 (50%)
Frame = +1
Query: 257 LLYKEEFEKMKEKAPENFRLDFAVSREQVNDKGEKMYIQTRMAQYAEELWELLKKDNTFV 316
LL+ +++ ++ + P + + VS ++DKG+ ++ Y +E +LL + T V
Sbjct: 304 LLHGQQYIELCKPFPSSCHIQNKVSLAGLHDKGKAAILEIETKSYEKESGDLLCMNRTTV 483
Query: 317 YMCGLKGMEK 326
Y+ G G K
Sbjct: 484 YLRGAGGFSK 513
>BF643344
Length = 649
Score = 33.1 bits (74), Expect = 0.16
Identities = 16/31 (51%), Positives = 21/31 (67%)
Frame = +3
Query: 5 VTAAVSFPYSNSTSLPIRTSVISPDRLVFKK 35
V AV PY NSTSL IR V+ ++++FKK
Sbjct: 249 VKFAVFIPYINSTSLLIRAFVVGSEKVLFKK 341
Score = 30.4 bits (67), Expect = 1.0
Identities = 15/32 (46%), Positives = 20/32 (61%)
Frame = +1
Query: 7 AAVSFPYSNSTSLPIRTSVISPDRLVFKKVSL 38
+AV PY N TSLPIR + ++ FKK +L
Sbjct: 163 SAVFIPYINPTSLPIRVFAVGS*KVPFKK*NL 258
>TC79175 similar to GP|21592883|gb|AAM64833.1 cytochrome-b5 reductase-like
protein {Arabidopsis thaliana}, partial (84%)
Length = 1251
Score = 32.3 bits (72), Expect = 0.27
Identities = 28/123 (22%), Positives = 55/123 (43%)
Frame = +1
Query: 176 KGVCSNFLCDLRPGSEVQITGPVGKEMLMPKDPNATVIMLGTGTGIAPFRSFLWKMFFEK 235
+G S L+PG V++ GP+ K P + + M+ G+GI P + + K
Sbjct: 481 EGKMSQHFASLKPGDMVEVKGPIEKFKYTP-NMKKNIGMIAGGSGITPMLQVIEAIL--K 651
Query: 236 HEDYKFNGLAWLFLGVPTSSSLLYKEEFEKMKEKAPENFRLDFAVSREQVNDKGEKMYIQ 295
+ D K ++ ++ V + +L +++ + + + P N ++ + V KG YI
Sbjct: 652 NSDDK-TQISLIYANV-SPDDILLQQKLDILAKSHP-NLKVFYTVDNPTKTWKGGAGYIS 822
Query: 296 TRM 298
M
Sbjct: 823 KDM 831
>TC86864 similar to PIR|F96570|F96570 unknown protein 80333-82175
[imported] - Arabidopsis thaliana, partial (89%)
Length = 1226
Score = 29.3 bits (64), Expect = 2.3
Identities = 16/56 (28%), Positives = 28/56 (49%)
Frame = -2
Query: 193 QITGPVGKEMLMPKDPNATVIMLGTGTGIAPFRSFLWKMFFEKHEDYKFNGLAWLF 248
Q GP +++ + P + GT I FR+ +W EK + + + GLA+L+
Sbjct: 844 QNRGPTSINVILSRLPRGNLSDSL*GTEIW*FRNHIWSSTIEKTKTWSWKGLAFLW 677
>CA920571 similar to GP|12322018|gb unknown protein; 3519-5443 {Arabidopsis
thaliana}, partial (48%)
Length = 827
Score = 29.3 bits (64), Expect = 2.3
Identities = 19/62 (30%), Positives = 31/62 (49%)
Frame = -2
Query: 204 MPKDPNATVIMLGTGTGIAPFRSFLWKMFFEKHEDYKFNGLAWLFLGVPTSSSLLYKEEF 263
+PK+PNA +++ T G P S L E + + + + W+ G SS LL+ EF
Sbjct: 457 IPKNPNAVILVAATDDGYIPKHSVL-----ELQKAWPGSEVRWV-TGGHVSSFLLHNGEF 296
Query: 264 EK 265
+
Sbjct: 295 RR 290
>TC83126 similar to GP|15529155|gb|AAK97672.1 AT3g30390/T6J22_16
{Arabidopsis thaliana}, partial (9%)
Length = 452
Score = 28.1 bits (61), Expect = 5.1
Identities = 14/62 (22%), Positives = 33/62 (52%)
Frame = -1
Query: 23 TSVISPDRLVFKKVSLNNVSISGRLTVRAEVATEAPAPVKVEKISKKQEEGIVVNKFKPK 82
T V+ +R + K+ + + SIS +T+R ++ +++ KQ++GI + +++
Sbjct: 446 TKVVKKNRNLEKQSTKTSKSISHDITIRLSLSNSTKPHENIQRGMSKQKQGI*LKQYQLL 267
Query: 83 TP 84
P
Sbjct: 266 VP 261
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.317 0.135 0.393
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,613,977
Number of Sequences: 36976
Number of extensions: 124395
Number of successful extensions: 523
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 513
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 515
length of query: 339
length of database: 9,014,727
effective HSP length: 97
effective length of query: 242
effective length of database: 5,428,055
effective search space: 1313589310
effective search space used: 1313589310
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 59 (27.3 bits)
Medicago: description of AC147013.10


