

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
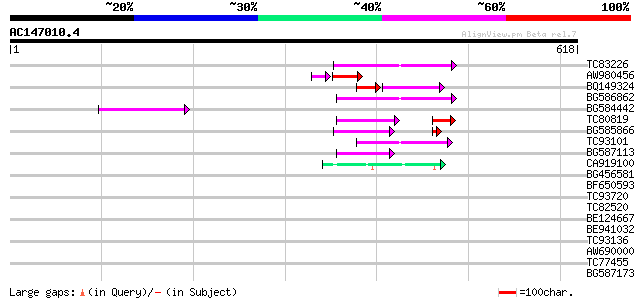
Query= AC147010.4 - phase: 0 /pseudo
(618 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC83226 weakly similar to PIR|G86419|G86419 probable reverse tra... 68 1e-11
AW980456 62 7e-11
BQ149324 44 6e-09
BG586862 59 7e-09
BG584442 52 9e-07
TC80819 weakly similar to GP|10140689|gb|AAG13524.1 putative non... 42 2e-06
BG585866 50 2e-06
TC93101 49 4e-06
BG587113 weakly similar to PIR|A84888|A8 hypothetical protein At... 47 2e-05
CA919100 homologue to PIR|G90291|G902 endoglucanase precursor [i... 42 5e-04
BG456581 41 0.001
BF650593 41 0.002
TC93720 similar to GP|21743320|dbj|BAC03314. putative nematode r... 38 0.010
TC82520 30 0.022
BE124667 36 0.050
BE941032 35 0.065
TC93136 35 0.11
AW690000 33 0.25
TC77455 similar to GP|22335695|dbj|BAC10549. nine-cis-epoxycarot... 32 0.94
BG587173 weakly similar to GP|4512630|gb| Contains reverse trans... 31 1.2
>TC83226 weakly similar to PIR|G86419|G86419 probable reverse transcriptase
100033-105622 [imported] - Arabidopsis thaliana, partial
(2%)
Length = 885
Score = 67.8 bits (164), Expect = 1e-11
Identities = 39/134 (29%), Positives = 64/134 (47%)
Frame = +3
Query: 354 WSGIWKLKVPPKFKNLVWCMCRGFQPTRVRLLDKGIQCPTNCVSCPSNHEDMSHIFFECP 413
W IW L P+ K L+W + P R L +GIQC C C S E ++H+F CP
Sbjct: 120 WKKIWSLHTIPRHKVLLWRILNDSLPVRSSLRKRGIQCYPLCPRCHSKTETITHLFMSCP 299
Query: 414 FAIHVW*MTSLWGTIQHALSSTTSAINVIFMLLETFSAELNQRLTSLFWSLWKHRNLRAW 473
+ VW ++L I + IN ++ + + + ++ ++LW RNL
Sbjct: 300 LSKRVWFGSNL--CINFDNLPNPNFIN*LYEAIL*KDECITI*IAAIIYNLWHARNLSVL 473
Query: 474 EDVTEVSVVVVERA 487
ED T + + +++RA
Sbjct: 474 EDQTILEMDIIQRA 515
>AW980456
Length = 779
Score = 61.6 bits (148), Expect(2) = 7e-11
Identities = 26/33 (78%), Positives = 28/33 (84%)
Frame = -2
Query: 352 GFWSGIWKLKVPPKFKNLVWCMCRGFQPTRVRL 384
G WSGIWKLKVPPK +NLVW MCRG PTR+RL
Sbjct: 103 GNWSGIWKLKVPPKVQNLVWRMCRGCLPTRIRL 5
Score = 23.5 bits (49), Expect(2) = 7e-11
Identities = 9/20 (45%), Positives = 11/20 (55%)
Frame = -1
Query: 330 VLCS*CLLVMCHGTR*LFSP 349
+LC CL +C GT F P
Sbjct: 173 ILCKECLSFLCRGTLRFFLP 114
>BQ149324
Length = 374
Score = 43.9 bits (102), Expect(2) = 6e-09
Identities = 21/68 (30%), Positives = 32/68 (46%)
Frame = +1
Query: 407 HIFFECPFAIHVW*MTSLWGTIQHALSSTTSAINVIFMLLETFSAELNQRLTSLFWSLWK 466
H+ F+C +++VW M TI L + N+IF L S E + WS+ K
Sbjct: 91 HLIFQCSSSLNVWSMLPFLSTISILLQQDMDSKNIIFKALHDLSNEDAALFCCVLWSI*K 270
Query: 467 HRNLRAWE 474
RN + W+
Sbjct: 271 QRNNKVWK 294
Score = 34.7 bits (78), Expect(2) = 6e-09
Identities = 14/26 (53%), Positives = 16/26 (60%)
Frame = +3
Query: 379 PTRVRLLDKGIQCPTNCVSCPSNHED 404
PTRVRL DK + CP +C C ED
Sbjct: 6 PTRVRLKDKRVTCPMDCTLCTVGSED 83
>BG586862
Length = 804
Score = 58.5 bits (140), Expect = 7e-09
Identities = 35/131 (26%), Positives = 58/131 (43%)
Frame = -1
Query: 357 IWKLKVPPKFKNLVWCMCRGFQPTRVRLLDKGIQCPTNCVSCPSNHEDMSHIFFECPFAI 416
+W +K P+ K+ +W + P + L +GI+C C C S E + H+F C
Sbjct: 654 VWGIKTIPRHKSFLWRLLHNALPVKDELHKRGIRCSLLCPRCESKIETVQHLFLNCEVTQ 475
Query: 417 HVW*MTSLWGTIQHALSSTTSAINVIFMLLETFSAELNQRLTSLFWSLWKHRNLRAWEDV 476
W + L I S + I + E LT+L +S+W RN + +E++
Sbjct: 474 KEWFGSQL--GINFHSSGVLHFHDWITNFILKNDEETIIALTALLYSIWHARNQKVFENI 301
Query: 477 TEVSVVVVERA 487
VV++RA
Sbjct: 300 DVPGDVVIQRA 268
>BG584442
Length = 775
Score = 51.6 bits (122), Expect = 9e-07
Identities = 36/99 (36%), Positives = 51/99 (51%)
Frame = +3
Query: 98 ENKFLE**MFI*GRARSYDQIRSTGYSVVCHEYFSTVVHSY*HY*KDDEFFLVGSW*NYS 157
+N FLE MF+ R D + + + +C EYFS S *KD E+F +GS S
Sbjct: 381 KN*FLEKQMFVKSYVRGND*VCTPKHIFLCDEYFSPSKFSSR*N*KDYEYFFMGSCWRKS 560
Query: 158 ARY*LDEVGEVICS*GPRWYGFQESLRFQFGYAW*ARVK 196
R LD +GE++C+* +GF Q+ AW* +K
Sbjct: 561 QRNALDVLGEIVCT*KLWRHGFYRLHDVQYSNAW*TGLK 677
>TC80819 weakly similar to GP|10140689|gb|AAG13524.1 putative non-LTR
retroelement reverse transcriptase {Oryza sativa
(japonica cultivar-group)}, partial (2%)
Length = 1262
Score = 42.0 bits (97), Expect(2) = 2e-06
Identities = 20/71 (28%), Positives = 32/71 (44%), Gaps = 2/71 (2%)
Frame = +2
Query: 357 IWKLKVPPKFKNLVWCMCRGFQPTRVRLLDKGIQCPTN--CVSCPSNHEDMSHIFFECPF 414
+W +P K VW + R PT+ L+ +G+ TN CV + E +H+F C
Sbjct: 626 VWHKHIPSKVSLFVWRLLRNRLPTKDNLVHRGVLLATNAACVCGCVDSESTTHLFLHCNV 805
Query: 415 AIHVW*MTSLW 425
+W + W
Sbjct: 806 FCSLWSLVRNW 838
Score = 28.1 bits (61), Expect(2) = 2e-06
Identities = 9/26 (34%), Positives = 16/26 (60%)
Frame = +1
Query: 461 FWSLWKHRNLRAWEDVTEVSVVVVER 486
FW +WK RN R +++ V+++R
Sbjct: 943 FWVIWKERNNRLFQNTVTAPFVLIDR 1020
>BG585866
Length = 828
Score = 49.7 bits (117), Expect(2) = 2e-06
Identities = 21/66 (31%), Positives = 31/66 (46%)
Frame = +3
Query: 354 WSGIWKLKVPPKFKNLVWCMCRGFQPTRVRLLDKGIQCPTNCVSCPSNHEDMSHIFFECP 413
WS IW+LK+P K+K +W C PT L + + C C + E H +C
Sbjct: 453 WSWIWRLKIPEKYKFFLWLACHNAVPTLSLLNHRNMVNSAICSRCGEHEESFFHCVRDCR 632
Query: 414 FAIHVW 419
F+ +W
Sbjct: 633 FSKIIW 650
Score = 20.0 bits (40), Expect(2) = 2e-06
Identities = 5/9 (55%), Positives = 7/9 (77%)
Frame = +2
Query: 462 WSLWKHRNL 470
W +W+HR L
Sbjct: 758 WWIWRHRTL 784
>TC93101
Length = 675
Score = 49.3 bits (116), Expect = 4e-06
Identities = 29/106 (27%), Positives = 55/106 (51%), Gaps = 2/106 (1%)
Frame = -1
Query: 379 PTRVRLLDKGIQCPTNCVSCPSNHEDMSHIFFECPFAIHVW*MTSLWGTIQHALSSTTSA 438
P R L +G+ CP C C N E +HIF C VW + L +I+ +ST +
Sbjct: 660 PVRXELNKRGVNCPPLCPRCYFNLETTNHIFMSCERTQRVWFGSQL--SIRFPDNSTINF 487
Query: 439 INVIFMLLETFSAELNQRLTSLFWSLWKHRNLRAWED--VTEVSVV 482
+ +F + + E+ +++++ +S+W RN +E+ V+E +++
Sbjct: 486 SDWLFDAISNQTEEIIIKISAITYSIWHARNKAIFENQFVSEDTII 349
>BG587113 weakly similar to PIR|A84888|A8 hypothetical protein At2g45230
[imported] - Arabidopsis thaliana, partial (10%)
Length = 767
Score = 47.4 bits (111), Expect = 2e-05
Identities = 18/63 (28%), Positives = 30/63 (47%)
Frame = -3
Query: 357 IWKLKVPPKFKNLVWCMCRGFQPTRVRLLDKGIQCPTNCVSCPSNHEDMSHIFFECPFAI 416
+WK PK ++ +W PT + + I +C C E ++HI F+CP+A
Sbjct: 288 VWKYNTSPKVRHFLWRCISNSLPTAANMRSRHISKDGSCSRCGMESETVNHILFQCPYAR 109
Query: 417 HVW 419
+W
Sbjct: 108 LIW 100
>CA919100 homologue to PIR|G90291|G902 endoglucanase precursor [imported] -
Sulfolobus solfataricus, partial (3%)
Length = 789
Score = 42.4 bits (98), Expect = 5e-04
Identities = 36/143 (25%), Positives = 57/143 (39%), Gaps = 9/143 (6%)
Frame = -2
Query: 342 GTR*LFSPYVGFWSGIWKLKVPPKFKNLVWCMCRGFQPTRVRLLDKGIQCPTN---CVSC 398
G+ L PY IW +VP K W + R PT+ L+ +G+ PT CVS
Sbjct: 620 GSLSLILPYAEM---IWHRQVPLKVSVFAWRLLRDRLPTKSNLIYRGV-IPTEAGLCVSG 453
Query: 399 PSNHEDMSHIFFECPFAIHVW*MTSLW-GTIQHALSSTTSAINVIFMLLETFSAELNQRL 457
E H+F C + +W + W G + + + + + + + T + +Q
Sbjct: 452 CGALESAQHLFLSCSYFASLWSLVRDWIGFV--GVDTNVLSDHFVQFVHSTGGNKASQSF 279
Query: 458 TSLF-----WSLWKHRNLRAWED 475
L W LW RN + D
Sbjct: 278 LQLIWLLCAWVLWTERNNMCFND 210
>BG456581
Length = 683
Score = 41.2 bits (95), Expect = 0.001
Identities = 18/65 (27%), Positives = 26/65 (39%)
Frame = +2
Query: 355 SGIWKLKVPPKFKNLVWCMCRGFQPTRVRLLDKGIQCPTNCVSCPSNHEDMSHIFFECPF 414
S IW +PP + W + PT L +G + C C E H+F C F
Sbjct: 74 STIWNSCIPPSHSFICWRLAHDRLPTDDNLSSRGCALVSMCSFCLEQVETSDHLFLRCKF 253
Query: 415 AIHVW 419
+ +W
Sbjct: 254 VVTLW 268
>BF650593
Length = 486
Score = 40.8 bits (94), Expect = 0.002
Identities = 24/60 (40%), Positives = 30/60 (50%), Gaps = 2/60 (3%)
Frame = +3
Query: 362 VPPKFKNLVWCMCRGFQPTRVRLLDKGI--QCPTNCVSCPSNHEDMSHIFFECPFAIHVW 419
VP K LVW + + TR L +G+ Q CV E +SH FFECPF+ VW
Sbjct: 306 VPLKVSCLVWRLFQNXLATRDNLSKRGVLDQNSIXCVXDCGREESVSHFFFECPFS-XVW 482
>TC93720 similar to GP|21743320|dbj|BAC03314. putative nematode
resistance-like protein {Oryza sativa (japonica
cultivar-group)}, partial (3%)
Length = 457
Score = 38.1 bits (87), Expect = 0.010
Identities = 16/47 (34%), Positives = 28/47 (59%)
Frame = -2
Query: 441 VIFMLLETFSAELNQRLTSLFWSLWKHRNLRAWEDVTEVSVVVVERA 487
+IF LL+ FS++ + + ++ WSLWK ++ W+ E + VV A
Sbjct: 441 LIFTLLQQFSSDQCELMATVMWSLWKSCRMKLWQQNNESNSQVVHHA 301
>TC82520
Length = 833
Score = 30.0 bits (66), Expect(2) = 0.022
Identities = 13/43 (30%), Positives = 22/43 (50%), Gaps = 2/43 (4%)
Frame = +3
Query: 357 IWKLKVPPKFKNLVWCMCRGFQPTRVRLLDKGI--QCPTNCVS 397
+W+ +P K VW + PT+V L+ + + Q T C+S
Sbjct: 186 VWQKNIPSKVSMFVWRLFHNRLPTKVNLMQRHVLQQTHTACIS 314
Score = 25.8 bits (55), Expect(2) = 0.022
Identities = 20/82 (24%), Positives = 31/82 (37%), Gaps = 9/82 (10%)
Frame = +2
Query: 403 EDMSHIFFECPFAIHVW*MTSLWGTIQHALSSTTSAINVIFMLLETFSAELNQRLTSLF- 461
E +H+F C +W W H L + I F+ + + + R T F
Sbjct: 329 ETATHLFLHCDIFGSLWSHVLRW---LHLLLVLPADIRQFFIQFTSMAG--SPRFTHSFL 493
Query: 462 --------WSLWKHRNLRAWED 475
W LWK RN R +++
Sbjct: 494 QIMWFASVWVLWKKRNNRVFQN 559
>BE124667
Length = 540
Score = 35.8 bits (81), Expect = 0.050
Identities = 13/28 (46%), Positives = 18/28 (63%)
Frame = -1
Query: 392 PTNCVSCPSNHEDMSHIFFECPFAIHVW 419
P+ C SC ++ E H+FFEC FA +W
Sbjct: 393 PSMCSSCQASSETTFHLFFECNFATKMW 310
>BE941032
Length = 435
Score = 35.4 bits (80), Expect = 0.065
Identities = 18/58 (31%), Positives = 27/58 (46%), Gaps = 2/58 (3%)
Frame = +2
Query: 370 VWCMCRGFQPTRVRLLDKGIQCPT--NCVSCPSNHEDMSHIFFECPFAIHVW*MTSLW 425
+WC+ PT+ LL +G+ +CV N D +H+F C F +W S W
Sbjct: 155 LWCVLLNRFPTKDNLLKRGVISAIYQSCVGECGNLYDATHLFLHCNFFRQIWINVSDW 328
>TC93136
Length = 722
Score = 34.7 bits (78), Expect = 0.11
Identities = 24/86 (27%), Positives = 40/86 (45%), Gaps = 2/86 (2%)
Frame = +1
Query: 402 HEDMSHIFFECPFAIHVW*MTSLWGTIQHALSSTTSAINVIFMLLETFSAELNQRLT--S 459
+E SH+F CPF VW + + G + + S +F LLE S L +
Sbjct: 259 NETSSHLFLHCPFLSSVW--SKILGWLYY------SKFVCVFRLLERRSRTKGVWLIWHA 414
Query: 460 LFWSLWKHRNLRAWEDVTEVSVVVVE 485
W +WK N R ++++++ +VE
Sbjct: 415 TIWVIWKGINNRIFKNISKAIDEIVE 492
>AW690000
Length = 652
Score = 33.5 bits (75), Expect = 0.25
Identities = 11/28 (39%), Positives = 17/28 (60%)
Frame = +2
Query: 392 PTNCVSCPSNHEDMSHIFFECPFAIHVW 419
P+ C C E H+FFEC +A+++W
Sbjct: 5 PSMCSLCCKQAESSLHLFFECSYAVNLW 88
>TC77455 similar to GP|22335695|dbj|BAC10549. nine-cis-epoxycarotenoid
dioxygenase1 {Pisum sativum}, partial (43%)
Length = 1865
Score = 31.6 bits (70), Expect = 0.94
Identities = 17/59 (28%), Positives = 24/59 (39%), Gaps = 3/59 (5%)
Frame = -2
Query: 357 IWKLKVPPKFKNLVWCMCRGFQPTRVRLLDKG---IQCPTNCVSCPSNHEDMSHIFFEC 412
+WK K P K W + PT V L + ++ CV C E + H+F C
Sbjct: 322 LWKSKAPAKVLAFSWTLFLDRIPTMVNLGKRRLLRVEDSKRCVFCGCQDETVVHLFLHC 146
>BG587173 weakly similar to GP|4512630|gb| Contains reverse transcriptase
domain (rvt) PF|00078. {Arabidopsis thaliana}, partial
(5%)
Length = 292
Score = 31.2 bits (69), Expect = 1.2
Identities = 18/68 (26%), Positives = 25/68 (36%)
Frame = +1
Query: 351 VGFWSGIWKLKVPPKFKNLVWCMCRGFQPTRVRLLDKGIQCPTNCVSCPSNHEDMSHIFF 410
V + S +W PK +W PTR R C+ C E H+F
Sbjct: 82 VDWESVVWFSGHIPKHAFNMWVANYDRLPTRCRFAAWDTSIVKTCLLCGHYDESRDHLFL 261
Query: 411 ECPFAIHV 418
C F+ H+
Sbjct: 262 RCSFSEHI 285
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.364 0.161 0.683
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 25,355,111
Number of Sequences: 36976
Number of extensions: 448237
Number of successful extensions: 7001
Number of sequences better than 10.0: 47
Number of HSP's better than 10.0 without gapping: 2073
Number of HSP's successfully gapped in prelim test: 287
Number of HSP's that attempted gapping in prelim test: 4778
Number of HSP's gapped (non-prelim): 2665
length of query: 618
length of database: 9,014,727
effective HSP length: 102
effective length of query: 516
effective length of database: 5,243,175
effective search space: 2705478300
effective search space used: 2705478300
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.5 bits)
S2: 61 (28.1 bits)
Medicago: description of AC147010.4


