

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147006.6 + phase: 0 /pseudo
(797 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
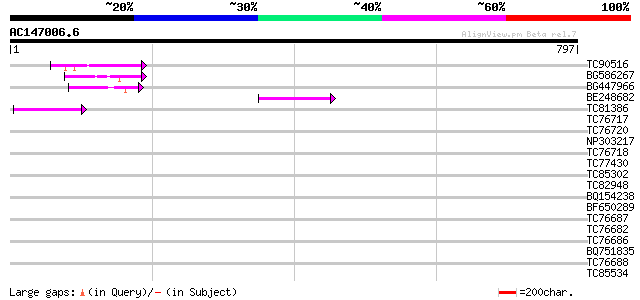
Score E
Sequences producing significant alignments: (bits) Value
TC90516 70 4e-12
BG586267 weakly similar to PIR|A84888|A8 hypothetical protein At... 59 1e-08
BG447966 weakly similar to PIR|G96509|G96 protein F27F5.21 [impo... 54 3e-07
BE248682 similar to GP|18568269|gb putative gag-pol polyprotein ... 53 5e-07
TC81386 similar to GP|10140689|gb|AAG13524.1 putative non-LTR re... 43 5e-04
TC76717 similar to PIR|T10863|T10863 extensin precursor - kidney... 35 0.15
TC76720 similar to GP|11121502|emb|CAC14888. putative extensin {... 35 0.15
NP303217 NP303217|AJ297721.1|CAC20726.1 putative repetitive prol... 34 0.25
TC76718 homologue to GP|8132441|gb|AAF73291.1| extensin {Pisum s... 32 0.95
TC77430 similar to GP|13676413|dbj|BAB41197. hypothetical protei... 32 0.95
TC85302 similar to GP|437310|gb|AAA62850.1|| nodulin {Medicago t... 32 1.2
TC82948 30 2.8
BQ154238 similar to PIR|T10863|T10 extensin precursor - kidney b... 30 2.8
BF650289 weakly similar to GP|9049283|dbj| orf129a {Beta vulgari... 30 3.6
TC76687 SP|Q40375|PRP2_MEDTR Repetitive proline-rich cell wall p... 30 3.6
TC76682 cell wall proline-rich protein 30 3.6
TC76686 homologue to SP|Q40375|PRP2_MEDTR Repetitive proline-ric... 30 3.6
BQ751835 30 3.6
TC76688 homologue to SP|Q40358|PRP_MEDSA Repetitive proline-rich... 30 4.7
TC85534 adenosylhomocysteinase [Medicago truncatula] 30 4.7
>TC90516
Length = 983
Score = 69.7 bits (169), Expect = 4e-12
Identities = 47/148 (31%), Positives = 79/148 (52%), Gaps = 13/148 (8%)
Frame = -2
Query: 58 KGKLVELWR-LLKAKDTLITQ---RSRSRWLKKGDA---------NSKFFHKCIKLRKSI 104
+G ++ LWR L++ I + RS++R G N+ +F C+K R
Sbjct: 877 RGGILFLWR*LIRTSVWPICKLFCRSKTRRCSNGQEQSG*EREGHNTTYFLACVKNRGMR 698
Query: 105 NSIKALEENEGWVVSPFEVCRKVVNYFTNHFAEDRWDRPRLDRVDFESLTEVENGLLVAP 164
NSI A E W+ + + +VN+FT+HF++ RP + +DF ++ ++N LL A
Sbjct: 697 NSISA-PRVERWMDDVAKNKQAIVNFFTHHFSDP*TYRPTMGDIDFFHISNLDNVLLSAQ 521
Query: 165 FSLLEIEEVVRDSDGGKSPGPNGYNFSF 192
F + ++E VV DG +SP P+G+N +F
Sbjct: 520 FLVSKMELVVSSLDGNESPRPDGFNLNF 437
>BG586267 weakly similar to PIR|A84888|A8 hypothetical protein At2g45230
[imported] - Arabidopsis thaliana, partial (16%)
Length = 794
Score = 58.5 bits (140), Expect = 1e-08
Identities = 40/122 (32%), Positives = 65/122 (52%), Gaps = 6/122 (4%)
Frame = +3
Query: 77 QRSRSRWLKKGDANSKFFHKCIKLRKSINSIKAL--EENEGWVVSPFEVCRKVVNYFTNH 134
Q+SR WL+ GD N+KFFH K R++ N I +L ++++ W V ++ R ++F
Sbjct: 141 QKSRLNWLRSGDRNTKFFHAVTKNRRAQNRILSLIDDDDKEWFVEE-DLGRLADSHFKLL 317
Query: 135 FAEDRWDRPRLDRVDFESL----TEVENGLLVAPFSLLEIEEVVRDSDGGKSPGPNGYNF 190
++ + + D+ S+ TE +N L+A S E+ E V D + K PGP+G N
Sbjct: 318 YSS---EDVGITLEDWNSIPAIVTEEQNAQLMAQISREEVREAVFDINPHKCPGPDGMNV 488
Query: 191 SF 192
F
Sbjct: 489 FF 494
>BG447966 weakly similar to PIR|G96509|G96 protein F27F5.21 [imported] -
Arabidopsis thaliana, partial (7%)
Length = 687
Score = 53.5 bits (127), Expect = 3e-07
Identities = 36/114 (31%), Positives = 49/114 (42%), Gaps = 9/114 (7%)
Frame = +2
Query: 83 WLKKGDANSKFFHKCIKLRKSINSIKALEENEG-WVVSPFEVCRKVVNYFTNHFAEDRWD 141
WLK GD N+KFFH R+ +N IK L++ G W V R ++ YF N F
Sbjct: 5 WLKDGDKNTKFFHSKASQRRKVNEIKKLKDETGNWCKGEENVERLLITYFNNLFTSS--- 175
Query: 142 RPRLDRVDFESLTEVENGLL--------VAPFSLLEIEEVVRDSDGGKSPGPNG 187
+ E EV G L F+ E+ E + K+PGP+G
Sbjct: 176 ----NPTAIEETCEVVKGKLSHEHIVWCEKEFTEEEVLEAINQMHPVKAPGPDG 325
>BE248682 similar to GP|18568269|gb putative gag-pol polyprotein {Zea mays},
partial (1%)
Length = 441
Score = 52.8 bits (125), Expect = 5e-07
Identities = 44/109 (40%), Positives = 56/109 (51%)
Frame = +1
Query: 350 RRRFAFIEALNKGIPLPLFCFSWRRKDLAA**GMRWTVVCLRALRLVIVVCWFLTCNMLM 409
RRR NK I LFCF RK+L ** MR V+C + L L NM M
Sbjct: 4 RRRLVSKGV*NKEIR*HLFCFFLWRKELVV**RMR*IVICSKVLMSREEERVSLIFNMPM 183
Query: 410 MLCVLVNQRWIIFGLLNLSLEALRWRWGSRLTSQRAR**E*MWIMSLWK 458
+L V +R IFGLL L + L+W +LTS RA ** M++ + W+
Sbjct: 184 ILFV*GCRRLTIFGLLKLYCKVLKWLPD*KLTSIRAL**VSMFLGTSWR 330
>TC81386 similar to GP|10140689|gb|AAG13524.1 putative non-LTR retroelement
reverse transcriptase {Oryza sativa (japonica
cultivar-group)}, partial (1%)
Length = 798
Score = 42.7 bits (99), Expect = 5e-04
Identities = 24/102 (23%), Positives = 52/102 (50%)
Frame = -1
Query: 6 KKLKGLKVILKE*IKKEYGGMEDRVELLVEDIKMLDEKGEEGTLTE*EVGLHKGKLVELW 65
+KLK LK +LK K +G + +V+ +++ + K + ++ + K + L
Sbjct: 309 QKLKNLKEVLKVWNKNTFGNVHSQVDNAYKELDDIQVKIDSIGYSDVLMDQEKAAQLNLE 130
Query: 66 RLLKAKDTLITQRSRSRWLKKGDANSKFFHKCIKLRKSINSI 107
L ++ ++S+ W +GD N+ FFH+ K++++ + I
Sbjct: 129 SALNIEEVFWHEKSKVNWHCEGDRNTAFFHRVAKIKRTSSLI 4
>TC76717 similar to PIR|T10863|T10863 extensin precursor - kidney bean,
partial (56%)
Length = 928
Score = 34.7 bits (78), Expect = 0.15
Identities = 29/105 (27%), Positives = 39/105 (36%), Gaps = 3/105 (2%)
Frame = -1
Query: 394 RLVIVVCW---FLTCNMLMMLCVLVNQRWIIFGLLNLSLEALRWRWGSRLTSQRAR**E* 450
RLV+V W F+ N VLV++ W F L+N WRW
Sbjct: 580 RLVLVHWWRRGFILVNWWWRRFVLVHRWWR*FVLVN-------WRW-------------- 464
Query: 451 MWIMSLWKWLSII*IVVCGVPVLNIWVFRWVLTRIVCLHGSRWWR 495
W+W+ I W+ RW R+V +H WWR
Sbjct: 463 ------WRWVFI-------------WLLRWWWRRLVLVH---WWR 395
Score = 32.0 bits (71), Expect = 0.95
Identities = 33/125 (26%), Positives = 45/125 (35%), Gaps = 9/125 (7%)
Frame = -1
Query: 384 RWTVVCLRAL---RLVIVVCW---FLTCNMLMMLCVLVNQRWIIFGLLNLS---LEALRW 434
RW + L RLV+V W F+ + VLVN W F L+N L + W
Sbjct: 460 RWVFIWLLRWWWRRLVLVHWWRRGFILVHRWRR*FVLVNWWWRRFILVNWWRR*LVLVNW 281
Query: 435 RWGSRLTSQRAR**E*MWIMSLWKWLSII*IVVCGVPVLNIWVFRWVLTRIVCLHGSRWW 494
W W+W+ I W+ RW R++ +H WW
Sbjct: 280 WW--------------------WRWIFI-------------WLLRWWRRRLILIH---WW 209
Query: 495 RT*VV 499
R * V
Sbjct: 208 RR*FV 194
>TC76720 similar to GP|11121502|emb|CAC14888. putative extensin {Nicotiana
sylvestris}, partial (51%)
Length = 1290
Score = 34.7 bits (78), Expect = 0.15
Identities = 29/105 (27%), Positives = 39/105 (36%), Gaps = 3/105 (2%)
Frame = -3
Query: 394 RLVIVVCW---FLTCNMLMMLCVLVNQRWIIFGLLNLSLEALRWRWGSRLTSQRAR**E* 450
RLV+V W F+ N VLV++ W F L+N WRW
Sbjct: 193 RLVLVHWWRRGFILVNWWWRRFVLVHRWWR*FVLVN-------WRW-------------- 77
Query: 451 MWIMSLWKWLSII*IVVCGVPVLNIWVFRWVLTRIVCLHGSRWWR 495
W+W+ I W+ RW R+V +H WWR
Sbjct: 76 ------WRWVFI-------------WLLRWWWRRLVLVH---WWR 8
>NP303217 NP303217|AJ297721.1|CAC20726.1 putative repetitive proline-rich
protein
Length = 525
Score = 33.9 bits (76), Expect = 0.25
Identities = 24/83 (28%), Positives = 31/83 (36%), Gaps = 28/83 (33%)
Frame = -1
Query: 275 WWTGSWW*MSWW----------IM*RRKSK--------------NVLFLRW--TSRRRMI 308
WW WW + WW + RR K N LF W SRRR++
Sbjct: 351 WWL*QWWFLVWWCINWSLTRWRFLVRRFRKWWL*Q*WFLVW*FINRLFFNWRLVSRRRLV 172
Query: 309 RLIGVSWSI**L--GWECVPSGW 329
V+WS+ + W CV W
Sbjct: 171 NWRIVNWSVSWIFVNWSCVNWIW 103
>TC76718 homologue to GP|8132441|gb|AAF73291.1| extensin {Pisum sativum},
partial (63%)
Length = 712
Score = 32.0 bits (71), Expect = 0.95
Identities = 33/125 (26%), Positives = 45/125 (35%), Gaps = 9/125 (7%)
Frame = -2
Query: 384 RWTVVCLRAL---RLVIVVCW---FLTCNMLMMLCVLVNQRWIIFGLLNLS---LEALRW 434
RW + L RLV+V W F+ + VLVN W F L+N L + W
Sbjct: 543 RWVFIWLLRWWWRRLVLVHWWRRGFILVHRWRR*FVLVNWWWRRFILVNWWRR*LVLVNW 364
Query: 435 RWGSRLTSQRAR**E*MWIMSLWKWLSII*IVVCGVPVLNIWVFRWVLTRIVCLHGSRWW 494
W W+W+ I W+ RW R++ +H WW
Sbjct: 363 WW--------------------WRWIFI-------------WLLRWWRRRLILIH---WW 292
Query: 495 RT*VV 499
R * V
Sbjct: 291 RR*FV 277
>TC77430 similar to GP|13676413|dbj|BAB41197. hypothetical protein {Glycine
max}, partial (71%)
Length = 2529
Score = 32.0 bits (71), Expect = 0.95
Identities = 11/30 (36%), Positives = 15/30 (49%), Gaps = 3/30 (10%)
Frame = +3
Query: 275 WWTGSWW*M---SWWIM*RRKSKNVLFLRW 301
WWT WW + WW+ +RK + RW
Sbjct: 1947 WWTCPWWLLWKGVWWLWRKRKRRGTWHARW 2036
>TC85302 similar to GP|437310|gb|AAA62850.1|| nodulin {Medicago truncatula},
partial (72%)
Length = 907
Score = 31.6 bits (70), Expect = 1.2
Identities = 16/37 (43%), Positives = 19/37 (51%)
Frame = -3
Query: 265 LNRLFLNVATWWTGSWW*MSWWIM*RRKSKNVLFLRW 301
+NR FLN W WW + WW + RR LF RW
Sbjct: 611 INRGFLN---GWFVHWWFLYWWFVYRR-----LFYRW 525
Score = 29.3 bits (64), Expect = 6.2
Identities = 15/42 (35%), Positives = 20/42 (46%), Gaps = 5/42 (11%)
Frame = -3
Query: 265 LNRLFLNVATWWTGSWW*M-----SWWIM*RRKSKNVLFLRW 301
+NR FLN WW WW + +WW++ R L RW
Sbjct: 806 VNRWFLN---WWLVHWWFLYRWLLNWWLVSWRLLNWWLVYRW 690
>TC82948
Length = 705
Score = 30.4 bits (67), Expect = 2.8
Identities = 13/25 (52%), Positives = 17/25 (68%)
Frame = +3
Query: 560 KISWVKWELVCQNKRNGSLGVNSLS 584
K+ V WE VC+ + GSLG+ SLS
Sbjct: 255 KVVKVSWEKVCRPIKEGSLGIRSLS 329
>BQ154238 similar to PIR|T10863|T10 extensin precursor - kidney bean, partial
(22%)
Length = 574
Score = 30.4 bits (67), Expect = 2.8
Identities = 24/94 (25%), Positives = 33/94 (34%)
Frame = -1
Query: 402 FLTCNMLMMLCVLVNQRWIIFGLLNLSLEALRWRWGSRLTSQRAR**E*MWIMSLWKWLS 461
F+ N VLV++ W F L+N WRW W+W+
Sbjct: 196 FILVNWWWRRFVLVHRWWR*FVLVN-------WRW--------------------WRWVF 98
Query: 462 II*IVVCGVPVLNIWVFRWVLTRIVCLHGSRWWR 495
I W+ RW R+V +H WWR
Sbjct: 97 I-------------WLLRWWWRRLVLVH---WWR 44
>BF650289 weakly similar to GP|9049283|dbj| orf129a {Beta vulgaris}, partial
(69%)
Length = 616
Score = 30.0 bits (66), Expect = 3.6
Identities = 12/33 (36%), Positives = 20/33 (60%)
Frame = +3
Query: 160 LLVAPFSLLEIEEVVRDSDGGKSPGPNGYNFSF 192
LL + F+ +E++ + D K+PG +GYN F
Sbjct: 9 LLCSEFTAVEVKNALFSMDSSKAPGIDGYNVHF 107
>TC76687 SP|Q40375|PRP2_MEDTR Repetitive proline-rich cell wall protein 2
precursor. [Barrel medic] {Medicago truncatula}, partial
(63%)
Length = 800
Score = 30.0 bits (66), Expect = 3.6
Identities = 8/17 (47%), Positives = 11/17 (64%)
Frame = -2
Query: 275 WWTGSWW*MSWWIM*RR 291
WW WW + WW++ RR
Sbjct: 274 WWLVHWWFLYWWLVNRR 224
>TC76682 cell wall proline-rich protein
Length = 1779
Score = 30.0 bits (66), Expect = 3.6
Identities = 12/37 (32%), Positives = 17/37 (45%)
Frame = -3
Query: 279 SWW*MSWWIM*RRKSKNVLFLRWTSRRRMIRLIGVSW 315
+WW + WW + R L RW RR++ V W
Sbjct: 532 NWWLVHWWFLYRWLINRWLLYRWFVDRRLLNWWLVHW 422
Score = 30.0 bits (66), Expect = 3.6
Identities = 8/17 (47%), Positives = 11/17 (64%)
Frame = -3
Query: 275 WWTGSWW*MSWWIM*RR 291
WW WW + WW++ RR
Sbjct: 349 WWLVHWWFLYWWLVNRR 299
Score = 29.3 bits (64), Expect = 6.2
Identities = 12/37 (32%), Positives = 16/37 (42%)
Frame = -3
Query: 279 SWW*MSWWIM*RRKSKNVLFLRWTSRRRMIRLIGVSW 315
+WW + WW + R L RW RR+ V W
Sbjct: 442 NWWLVHWWFLYRWLVNRWLLYRWFVDRRLFNWWLVHW 332
>TC76686 homologue to SP|Q40375|PRP2_MEDTR Repetitive proline-rich cell wall
protein 2 precursor. [Barrel medic] {Medicago
truncatula}, partial (73%)
Length = 825
Score = 30.0 bits (66), Expect = 3.6
Identities = 12/37 (32%), Positives = 17/37 (45%)
Frame = -3
Query: 279 SWW*MSWWIM*RRKSKNVLFLRWTSRRRMIRLIGVSW 315
+WW + WW + R L RW RR++ V W
Sbjct: 277 NWWLVHWWFLYRWLINRWLLYRWFVDRRLLNWWLVHW 167
Score = 30.0 bits (66), Expect = 3.6
Identities = 8/17 (47%), Positives = 11/17 (64%)
Frame = -3
Query: 275 WWTGSWW*MSWWIM*RR 291
WW WW + WW++ RR
Sbjct: 94 WWLVHWWFLYWWLVNRR 44
Score = 29.3 bits (64), Expect = 6.2
Identities = 12/37 (32%), Positives = 16/37 (42%)
Frame = -3
Query: 279 SWW*MSWWIM*RRKSKNVLFLRWTSRRRMIRLIGVSW 315
+WW + WW + R L RW RR+ V W
Sbjct: 187 NWWLVHWWFLYRWLVNRWLLYRWFVDRRLFNWWLVHW 77
>BQ751835
Length = 570
Score = 30.0 bits (66), Expect = 3.6
Identities = 20/44 (45%), Positives = 23/44 (51%)
Frame = -1
Query: 693 RMGCLR*NRNL*NWRALFLVRFCGGRRKNELLRSCGRVWRLRRW 736
RM R R L +W A+ LVR G RR + LL R WRL W
Sbjct: 414 RMPQPRDQRGLSSW-AVVLVRLLGTRRCSRLLGRGFRGWRLSSW 286
>TC76688 homologue to SP|Q40358|PRP_MEDSA Repetitive proline-rich cell wall
protein precursor (MSPRP). [Alfalfa] {Medicago sativa},
partial (91%)
Length = 667
Score = 29.6 bits (65), Expect = 4.7
Identities = 14/39 (35%), Positives = 20/39 (50%), Gaps = 5/39 (12%)
Frame = -2
Query: 275 WWTGSWW*MSWWIM*R----RKSKNVLFL-RWTSRRRMI 308
WW WW + WW++ R R + FL RW RR++
Sbjct: 198 WWLVHWWFLYWWLINRWLLYRGFVDRGFLNRWLVNRRLV 82
>TC85534 adenosylhomocysteinase [Medicago truncatula]
Length = 1874
Score = 29.6 bits (65), Expect = 4.7
Identities = 15/44 (34%), Positives = 17/44 (38%)
Frame = +1
Query: 653 FSLGEGIYSCGKRRCF*ALGRIWKG*VCLTMLMCGSGNWMRMGC 696
F G G Y C K C + R WKG C C +W C
Sbjct: 823 FDEGYGCYDCWKGGCCRWIWRCWKGLCC-----CPEASWSSCDC 939
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.356 0.160 0.625
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 33,876,595
Number of Sequences: 36976
Number of extensions: 644140
Number of successful extensions: 8379
Number of sequences better than 10.0: 47
Number of HSP's better than 10.0 without gapping: 2425
Number of HSP's successfully gapped in prelim test: 423
Number of HSP's that attempted gapping in prelim test: 5220
Number of HSP's gapped (non-prelim): 3650
length of query: 797
length of database: 9,014,727
effective HSP length: 104
effective length of query: 693
effective length of database: 5,169,223
effective search space: 3582271539
effective search space used: 3582271539
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.7 bits)
S2: 62 (28.5 bits)
Medicago: description of AC147006.6


