

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
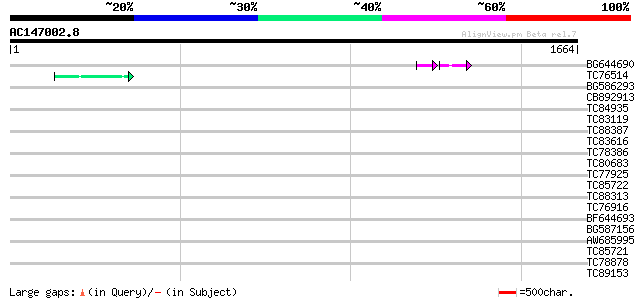
Query= AC147002.8 + phase: 0 /pseudo
(1664 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BG644690 weakly similar to GP|18542179|gb putative pol protein {... 48 7e-10
TC76514 homologue to PIR|T09648|T09648 nucleolin homolog nuM1 - ... 47 6e-05
BG586293 weakly similar to PIR|E84473|E84 probable retroelement ... 41 0.003
CB892913 similar to GP|1769897|emb lectin receptor kinase {Arabi... 41 0.004
TC84935 similar to PIR|G96631|G96631 probable RNA-binding protei... 36 0.11
TC83119 similar to GP|13357253|gb|AAK20050.1 putative zinc finge... 36 0.11
TC88387 similar to GP|13357253|gb|AAK20050.1 putative zinc finge... 36 0.11
TC83616 similar to GP|16226661|gb|AAL16226.1 At2g28910/F8N16.20 ... 35 0.19
TC78386 similar to SP|Q9NZW4|DSPP_HUMAN Dentin sialophosphoprote... 35 0.19
TC80683 homologue to GP|8777424|dbj|BAA97014.1 gb|AAF56406.1~gen... 35 0.24
TC77925 similar to GP|18700163|gb|AAL77693.1 AT4g02720/T10P11_1 ... 35 0.32
TC85722 similar to SP|P08283|H1_PEA Histone H1. [Garden pea] {Pi... 34 0.54
TC88313 weakly similar to PIR|E84776|E84776 hypothetical protein... 33 0.71
TC76916 MtN4 33 0.93
BF644693 homologue to GP|2598599|emb MtN4 {Medicago truncatula},... 33 0.93
BG587156 similar to PIR|G85055|G8 probable polyprotein [imported... 33 0.93
AW685995 similar to PIR|I51618|I516 nucleolar phosphoprotein - A... 33 0.93
TC85721 similar to SP|P08283|H1_PEA Histone H1. [Garden pea] {Pi... 33 1.2
TC78878 similar to GP|4519792|dbj|BAA75744.1 Asp1 {Arabidopsis t... 32 2.1
TC89153 similar to GP|18855061|gb|AAL79753.1 putative RNA helica... 32 2.1
>BG644690 weakly similar to GP|18542179|gb putative pol protein {Zea mays},
partial (22%)
Length = 629
Score = 48.1 bits (113), Expect(2) = 7e-10
Identities = 36/98 (36%), Positives = 51/98 (51%), Gaps = 6/98 (6%)
Frame = -1
Query: 1262 SRLQSTRRH*LH*NICSSCKIGS----NQVTSILRN*SWHNI--ISNGCQKCLS*WCH*R 1315
+R+QS RR+ L * + C+ GS N I H + + NGC++C+ *W R
Sbjct: 395 ARIQSKRRNRL**GFFTCCQNGSY*NFNSFCCI------HGVQAVPNGCEECIY*WRSQR 234
Query: 1316 RSVC*TTSWV*GS*AS*PCL*T*EITIWLETSSQSLV* 1353
VC TSW+* + C+ TIW E SS+S+V*
Sbjct: 233 GGVCQATSWI*RCRGTKSCVQIE*DTIWSEASSKSMV* 120
Score = 35.0 bits (79), Expect(2) = 7e-10
Identities = 25/62 (40%), Positives = 33/62 (52%)
Frame = -2
Query: 1194 *AQNC*RSSLR*WMDISYARRAKSIPKE*CVGSGTQTFSEEHYWNKMGIQKQAE*TRRSN 1253
*AQ C*RS +D YARR S+ K+* + G+ T + WN +G KQA R+
Sbjct: 589 *AQEC*RSIA*CRLDQFYARRTPSV*KK*GMVPGSST*RQNSNWN*VGF*KQA*VITRNK 410
Query: 1254 QK 1255
K
Sbjct: 409 SK 404
>TC76514 homologue to PIR|T09648|T09648 nucleolin homolog nuM1 - alfalfa,
partial (61%)
Length = 1288
Score = 47.0 bits (110), Expect = 6e-05
Identities = 58/238 (24%), Positives = 93/238 (38%), Gaps = 6/238 (2%)
Frame = +3
Query: 131 LQILKKSYVSSDHVSKILRSLPSRWRPKVTAIEEAKDLNTLSVE-DLVSSLKVHEMSLNE 189
L+++KK SS+ S+ P P + AK N + + S + S ++
Sbjct: 267 LKVVKKEESSSEESSESEDEQPVVKAPAPSKKTPAKKGNVKKAQPETTSEESDSDSSSSD 446
Query: 190 HETSKKSKSIALPSKGKTSKSSKAYKASESVEESPDGDSDEDQS-VKMAMLSNKLEYLAR 248
E KK S A+PSK + S+ A K S EE + SDED+ A+ S A+
Sbjct: 447 EEEVKKPVSKAVPSK---NGSAPAKKVDTSEEEDSEESSDEDKKPAAKAVPSKNGSAPAK 617
Query: 249 KQKKFLSKRGSYKNFKKEDQKGCFNC--KKPGHFIADCPDLQK--EKFKGKSKKSSFNSS 304
K + +ED+K K G A D + E + ++ ++
Sbjct: 618 KAASDEEDTDESSDEDEEDEKPAAKAVPSKNGSVPAKKADTESSDEDSESSDEEDKKPAA 797
Query: 305 KFRKQIKKSLMATWEDLDSESGSDKEEADDDAKAAVGLVATVSSEAVSEAESDSEDEN 362
K K + D ES + +E D+DAK A ++SD ED++
Sbjct: 798 KASKNVSAPTKKAASSSDEESDEESDE-DEDAKPVSKPAAVAKKSKKDSSDSDDEDDD 968
>BG586293 weakly similar to PIR|E84473|E84 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(7%)
Length = 763
Score = 41.2 bits (95), Expect = 0.003
Identities = 23/69 (33%), Positives = 36/69 (51%), Gaps = 2/69 (2%)
Frame = +1
Query: 1253 NQKQSQTCCSRLQSTRRH*LH*NICSSCKIGSNQVTSILRN*SWHNIISNGCQKCLS*W- 1311
+Q QS+ C RL+ T RH L ++C+SC ++ T + + W S+ C+ C+ W
Sbjct: 145 DQIQSKASCKRLRETTRHRLRRSVCTSCSNRNHMTTLGVSSN*WMLDPSHRCKNCIPKWT 324
Query: 1312 -CH*RRSVC 1319
C R S C
Sbjct: 325 LCGNRNSHC 351
>CB892913 similar to GP|1769897|emb lectin receptor kinase {Arabidopsis
thaliana}, partial (35%)
Length = 837
Score = 40.8 bits (94), Expect = 0.004
Identities = 28/85 (32%), Positives = 43/85 (49%)
Frame = +1
Query: 115 ESIEEMYSRFQTLVSGLQILKKSYVSSDHVSKILRSLPSRWRPKVTAIEEAKDLNTLSVE 174
E + +SR ++ + L+ + + KILRSL ++ V IEE KDL T+++E
Sbjct: 472 ERVRVYFSRVISISNQLKRNSEKLEDVRIMEKILRSLDPKFEHIVEKIEETKDLETMTIE 651
Query: 175 DLVSSLKVHEMSLNEHETSKKSKSI 199
L SL+ +E E KK K I
Sbjct: 652 KLQGSLQAYE------EKHKKKKDI 708
>TC84935 similar to PIR|G96631|G96631 probable RNA-binding protein F8A5.17
[imported] - Arabidopsis thaliana, partial (41%)
Length = 552
Score = 36.2 bits (82), Expect = 0.11
Identities = 14/29 (48%), Positives = 17/29 (58%)
Frame = +2
Query: 257 RGSYKNFKKEDQKGCFNCKKPGHFIADCP 285
RGSY + Q CF C +PGH+ DCP
Sbjct: 431 RGSYGAGDRVGQDDCFKCGRPGHWARDCP 517
>TC83119 similar to GP|13357253|gb|AAK20050.1 putative zinc finger protein
{Oryza sativa (japonica cultivar-group)}, partial (16%)
Length = 421
Score = 36.2 bits (82), Expect = 0.11
Identities = 15/45 (33%), Positives = 28/45 (61%), Gaps = 8/45 (17%)
Frame = +3
Query: 251 KKFLSKRGSYKN--FKKEDQKG------CFNCKKPGHFIADCPDL 287
+K S R SY++ ++++ +G C NCK+PGH+ +CP++
Sbjct: 264 RKIRSDRHSYRDAPYRRDSSRGFSRDNLCKNCKRPGHYARECPNV 398
>TC88387 similar to GP|13357253|gb|AAK20050.1 putative zinc finger protein
{Oryza sativa (japonica cultivar-group)}, partial (96%)
Length = 1286
Score = 36.2 bits (82), Expect = 0.11
Identities = 23/84 (27%), Positives = 42/84 (49%), Gaps = 1/84 (1%)
Frame = +1
Query: 205 GKTSKSSKAYKA-SESVEESPDGDSDEDQSVKMAMLSNKLEYLARKQKKFLSKRGSYKNF 263
GK K S ++ S+S SP S V + S + + ++ +R S + F
Sbjct: 244 GKILKMSSVSRSRSKSRSRSPMDRSRSRSPVDRRIRSERFSH-----REAPYRRDSRRGF 408
Query: 264 KKEDQKGCFNCKKPGHFIADCPDL 287
+++ C NCK+PGH++ +CP++
Sbjct: 409 SQDNL--CKNCKRPGHYVRECPNV 474
Score = 31.6 bits (70), Expect = 2.7
Identities = 10/20 (50%), Positives = 14/20 (70%)
Frame = +1
Query: 267 DQKGCFNCKKPGHFIADCPD 286
++K C NC+K GH DCP+
Sbjct: 718 NEKACNNCRKTGHLARDCPN 777
Score = 31.2 bits (69), Expect = 3.5
Identities = 9/16 (56%), Positives = 13/16 (81%)
Frame = +1
Query: 271 CFNCKKPGHFIADCPD 286
C+NCK+PGH + CP+
Sbjct: 538 CWNCKEPGHMASSCPN 585
>TC83616 similar to GP|16226661|gb|AAL16226.1 At2g28910/F8N16.20
{Arabidopsis thaliana}, partial (30%)
Length = 854
Score = 35.4 bits (80), Expect = 0.19
Identities = 31/105 (29%), Positives = 44/105 (41%), Gaps = 3/105 (2%)
Frame = +2
Query: 204 KGKTSKSSKAYKASESVEESPDG---DSDEDQSVKMAMLSNKLEYLARKQKKFLSKRGSY 260
KGK K K S EE D DS+ D ++ A++ R KK KRGSY
Sbjct: 530 KGKAEKVDKGSNVESSEEEEDDSESSDSEIDSEIERAIVK-------RSGKKVSEKRGSY 688
Query: 261 KNFKKEDQKGCFNCKKPGHFIADCPDLQKEKFKGKSKKSSFNSSK 305
+ KK+D + K +K K +G+ KK S + +
Sbjct: 689 R--KKDDSDDDESDKDSD---------RKRKKRGREKKRSVKTXR 790
>TC78386 similar to SP|Q9NZW4|DSPP_HUMAN Dentin sialophosphoprotein
precursor [Contains: Dentin phosphoprotein (Dentin
phosphophoryn) (DPP);, partial (2%)
Length = 726
Score = 35.4 bits (80), Expect = 0.19
Identities = 33/122 (27%), Positives = 53/122 (43%), Gaps = 23/122 (18%)
Frame = +3
Query: 165 AKDLNTLSVEDLVSSLKVHEMSLNEHETS---KKSKSIALPSKGKTSKSSKAY---KASE 218
+ D + S L S + S +EHETS KK K P K K + SK++ +
Sbjct: 228 SSDSDFYSSSSLSDSSRSESSSNSEHETSHRSKKHKKSDKPKKNKEKERSKSHRHKRHKH 407
Query: 219 SVEESPDGD-------------SDEDQSVKMAMLSNKLEYL----ARKQKKFLSKRGSYK 261
S++E P+G+ ++D V+ + +S K L ++ K+ SKR
Sbjct: 408 SLKEKPNGERSSSPVQLSKFLGRNKDDGVRRSAVSGKKILLKLDKTKEDKEAESKRNELL 587
Query: 262 NF 263
NF
Sbjct: 588 NF 593
>TC80683 homologue to GP|8777424|dbj|BAA97014.1
gb|AAF56406.1~gene_id:K9P8.7~strong similarity to
unknown protein {Arabidopsis thaliana}, partial (16%)
Length = 1360
Score = 35.0 bits (79), Expect = 0.24
Identities = 25/96 (26%), Positives = 45/96 (46%), Gaps = 6/96 (6%)
Frame = +1
Query: 197 KSIALPSKGKTSKSSKAYKASESVEESPDGDSDEDQSVKMAMLSNKLEYLARKQKKFLSK 256
K I+ KGK K + Y + +E++S++M++L++ + + K+++ L
Sbjct: 874 KKISRGQKGKLKKMKEKYADQD----------EEERSIRMSLLASSGKPI--KKEETLPV 1017
Query: 257 RGSYKNFKKEDQ------KGCFNCKKPGHFIADCPD 286
+ KK D K C+ CKK GH DC +
Sbjct: 1018IETSDKGKKSDSGPIDAPKICYKCKKVGHLSRDCKE 1125
>TC77925 similar to GP|18700163|gb|AAL77693.1 AT4g02720/T10P11_1
{Arabidopsis thaliana}, partial (47%)
Length = 1538
Score = 34.7 bits (78), Expect = 0.32
Identities = 46/223 (20%), Positives = 90/223 (39%)
Frame = +3
Query: 189 EHETSKKSKSIALPSKGKTSKSSKAYKASESVEESPDGDSDEDQSVKMAMLSNKLEYLAR 248
+ E +K+KS + + K S S K +S ++S + DS+ + +++E R
Sbjct: 519 DKEIKQKAKSESESEESKESDSELRKKRRKSYKKSRESDSESESE-------SEVEDRKR 677
Query: 249 KQKKFLSKRGSYKNFKKEDQKGCFNCKKPGHFIADCPDLQKEKFKGKSKKSSFNSSKFRK 308
++ + S+ S N ++ED+K + + +KSS K +
Sbjct: 678 RKSRKYSESDSDTNSEEEDRK-----------------------RRRRRKSSRRRRKSSR 788
Query: 309 QIKKSLMATWEDLDSESGSDKEEADDDAKAAVGLVATVSSEAVSEAESDSEDENEVYSKI 368
+ ++ + + D ESG D D K + S + SE S+ +E E+ S
Sbjct: 789 KKRRCSDSDESETDDESGYD----DSGRKRKRSKRSRKSKKKSSEPVSEGSEEIELGSD- 953
Query: 369 PRQELVDSLKELLSLFEHRTNELTDLKEKYVDLMKQQKSTLLE 411
+ +E+ + E + E+ K +L + QK L+
Sbjct: 954 --SAVAKINEEIDDVDEEKMAEMNAEALKLKELFESQKKPALD 1076
>TC85722 similar to SP|P08283|H1_PEA Histone H1. [Garden pea] {Pisum
sativum}, partial (61%)
Length = 897
Score = 33.9 bits (76), Expect = 0.54
Identities = 41/146 (28%), Positives = 61/146 (41%), Gaps = 17/146 (11%)
Frame = +3
Query: 525 KPEASGSQAKITSKPENLKIKVMTKSDPKSQKIKILKRSEPVHQNLIKPESKI------- 577
K ASG K+ + K+ TK + K+K ++P + ++KP +K
Sbjct: 399 KNVASGKLVKVKG---SFKLSSATKP---AAKVKAKPAAKPKAKAVVKPRTKSVTAKPKA 560
Query: 578 ----PKQKDQKNKAATASEKTIPKGVKPKVLNDQKPLSIHPKVQGRKSKTSK--ANPKGP 631
PK K KAA KT K VKP + P G+K T+K A K P
Sbjct: 561 AAAKPKSAATKPKAAVGKAKTAAK-VKPNAKVAKTTTRTSP---GKKVATAKPAAKKKTP 728
Query: 632 MKIWVPKSEL----AKNAGVLKGKRE 653
+K PK++ AK + +G R+
Sbjct: 729 VKNVKPKTKTVRSPAKKVAIKRGGRK 806
>TC88313 weakly similar to PIR|E84776|E84776 hypothetical protein At2g36070
[imported] - Arabidopsis thaliana, partial (31%)
Length = 1201
Score = 33.5 bits (75), Expect = 0.71
Identities = 45/208 (21%), Positives = 89/208 (42%), Gaps = 18/208 (8%)
Frame = +2
Query: 368 IPRQELVDSLKELLSLFEHRTNELTDLKEKYVDLMKQQKSTLLELKASEEELKGFNLISA 427
+P+Q+ V +++ +F +N++ D K + K K ELK EELKG +
Sbjct: 191 LPQQQGVKTIRRY-GVFNEFSNKVKDETVKNPEFQKSVK----ELKEKAEELKG---VKE 346
Query: 428 TYEDRLKSLCQKLQEKCD-----------KGSGNKHE--IALDDFIMAGIDRSKVASMIY 474
+++ K ++L ++ D K S N E A + + AGI + +
Sbjct: 347 GLKEKTKQTTEQLYKQFDGVWKEAEAAAKKVSHNVKEKISAATEEVKAGIGKQDSSGSTD 526
Query: 475 STYK-----NKGKGIGYSEEKSKEYSLKSYCDCIKDGLKSTFVPEGTNAKTAVQSKPEAS 529
S+ K +G+ EEK++E + + + + KSTF ++ K + +
Sbjct: 527 SSTKQDADAKQGRQTSPEEEKNQESASGNASESLFGKFKSTF----SSPKVSTSFQKLKD 694
Query: 530 GSQAKITSKPENLKIKVMTKSDPKSQKI 557
+T K ++ + ++ + PK Q +
Sbjct: 695 AKIVDMTKKGYDILKEELSGNTPKRQPV 778
>TC76916 MtN4
Length = 1850
Score = 33.1 bits (74), Expect = 0.93
Identities = 15/45 (33%), Positives = 23/45 (50%), Gaps = 4/45 (8%)
Frame = -3
Query: 667 HDWRESFVPYPYNERWRRSEVWWQPNW---KDHW-YRYYW*FLHL 707
H+WR ++ + WR +WW+ NW W YRY W F ++
Sbjct: 984 HNWRCWYIRWW*WYNWR*RNMWWENNWWLRNIRWCYRYNWWFRNI 850
Score = 32.3 bits (72), Expect = 1.6
Identities = 12/31 (38%), Positives = 16/31 (50%), Gaps = 4/31 (12%)
Frame = -3
Query: 678 YNERWRRSEVWWQPNWKDH----WYRYYW*F 704
Y WR +WW+ NW H WY +W*+
Sbjct: 1038 YRHNWR*RNMWWKNNWWFHNWRCWYIRWW*W 946
>BF644693 homologue to GP|2598599|emb MtN4 {Medicago truncatula}, partial
(79%)
Length = 621
Score = 33.1 bits (74), Expect = 0.93
Identities = 15/45 (33%), Positives = 23/45 (50%), Gaps = 4/45 (8%)
Frame = -3
Query: 667 HDWRESFVPYPYNERWRRSEVWWQPNW---KDHW-YRYYW*FLHL 707
H+WR ++ + WR +WW+ NW W YRY W F ++
Sbjct: 211 HNWRCWYIRWW*WYNWR*RNMWWENNWWLRNIRWCYRYNWWFRNI 77
Score = 32.3 bits (72), Expect = 1.6
Identities = 12/31 (38%), Positives = 16/31 (50%), Gaps = 4/31 (12%)
Frame = -3
Query: 678 YNERWRRSEVWWQPNWKDH----WYRYYW*F 704
Y WR +WW+ NW H WY +W*+
Sbjct: 265 YRHNWR*RNMWWKNNWWFHNWRCWYIRWW*W 173
>BG587156 similar to PIR|G85055|G8 probable polyprotein [imported] -
Arabidopsis thaliana, partial (17%)
Length = 618
Score = 33.1 bits (74), Expect = 0.93
Identities = 18/56 (32%), Positives = 31/56 (55%)
Frame = -3
Query: 1254 QKQSQTCCSRLQSTRRH*LH*NICSSCKIGSNQVTSILRN*SWHNIISNGCQKCLS 1309
++++ T R+ S LH*+IC+S + N L W I++NGC++C+S
Sbjct: 382 EEEN*TSSKRVYSDIWRGLH*DICTSSQATHN*NCFKLGCEPWMGIVANGCEECIS 215
>AW685995 similar to PIR|I51618|I516 nucleolar phosphoprotein - African
clawed frog, partial (5%)
Length = 516
Score = 33.1 bits (74), Expect = 0.93
Identities = 37/149 (24%), Positives = 64/149 (42%), Gaps = 2/149 (1%)
Frame = +3
Query: 226 GDSDEDQSVKMAMLSNKLEYLARKQKKFLSKRGSYKNFKKEDQKGCFNCKKPGHFIADCP 285
G S + VK A K+E + + + K +K K KK+ +K +KP +
Sbjct: 24 GKSKTSKKVK-AEEPKKVEKVEKTKGKKAAKAEEEK--KKQQKK---KEEKPAPMEVEED 185
Query: 286 DLQKEKFKGKSKKSSFNSSKFRKQIKKSLMATWEDLDSESGSDKEEADDDAKAAVGLVAT 345
E+ ++ + K+ ++ E+ S S S+ EE + AK A +
Sbjct: 186 SSSSEESSSSEDEAPAKKAAPAKKAAAKKESSSEEESSSSESESEEEEKPAKKAAAKKES 365
Query: 346 VSSE--AVSEAESDSEDENEVYSKIPRQE 372
S E + SE ESD E++NE + ++E
Sbjct: 366 SSEEESSSSEEESDEEEKNEARRR*EQEE 452
>TC85721 similar to SP|P08283|H1_PEA Histone H1. [Garden pea] {Pisum
sativum}, partial (60%)
Length = 995
Score = 32.7 bits (73), Expect = 1.2
Identities = 38/148 (25%), Positives = 60/148 (39%), Gaps = 19/148 (12%)
Frame = +1
Query: 525 KPEASGSQAKITSKPENLKIKVMTKSDPKSQKIKILKRSEPVHQNLIKPESKI------- 577
K ASG K+ + K+ TK + K+K ++P + ++KP++K
Sbjct: 433 KNVASGKLVKVKG---SFKLSSATKP---AAKVKAKPAAKPKAKAVVKPKTKSVTAKPKA 594
Query: 578 -----PKQKDQKNKAATASEKTIPKGVKPKVLNDQKPLSIHPKVQGRK---SKTSKANPK 629
PK K+KAA+ ++ KV S KV K K + A K
Sbjct: 595 AAATKPKPAATKSKAASKAKTAAKAKPNAKVAKTTARTSPGKKVAAAKPAAKKVAAAKKK 774
Query: 630 GPMKIWVPKSEL----AKNAGVLKGKRE 653
P+K PK++ AK V +G R+
Sbjct: 775 APVKSVKPKTKTVKSPAKKVSVKRGGRK 858
Score = 31.6 bits (70), Expect = 2.7
Identities = 19/69 (27%), Positives = 35/69 (50%)
Frame = +1
Query: 516 TNAKTAVQSKPEASGSQAKITSKPENLKIKVMTKSDPKSQKIKILKRSEPVHQNLIKPES 575
+ AKTA ++KP A ++ + P K + + P ++K+ K+ PV +KP++
Sbjct: 643 SKAKTAAKAKPNAKVAKTTARTSPG----KKVAAAKPAAKKVAAAKKKAPVKS--VKPKT 804
Query: 576 KIPKQKDQK 584
K K +K
Sbjct: 805 KTVKSPAKK 831
>TC78878 similar to GP|4519792|dbj|BAA75744.1 Asp1 {Arabidopsis thaliana},
partial (53%)
Length = 1839
Score = 32.0 bits (71), Expect = 2.1
Identities = 22/59 (37%), Positives = 31/59 (52%), Gaps = 2/59 (3%)
Frame = +3
Query: 139 VSSDHVSKI--LRSLPSRWRPKVTAIEEAKDLNTLSVEDLVSSLKVHEMSLNEHETSKK 195
+SSD + I L S +W+P VTAI LNT S+ L+SS + + L T+ K
Sbjct: 252 LSSDPLQWIHGLISRSRKWKPVVTAISTRSFLNTESLRKLISSPSIIQTLLRSTATAFK 428
>TC89153 similar to GP|18855061|gb|AAL79753.1 putative RNA helicase {Oryza
sativa}, partial (3%)
Length = 737
Score = 32.0 bits (71), Expect = 2.1
Identities = 13/32 (40%), Positives = 18/32 (55%)
Frame = +3
Query: 255 SKRGSYKNFKKEDQKGCFNCKKPGHFIADCPD 286
S R S N + CF+C +PGH +DCP+
Sbjct: 99 SNRSSSPNRRGSYGGACFSCGQPGHRASDCPN 194
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.357 0.156 0.559
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 49,360,586
Number of Sequences: 36976
Number of extensions: 708624
Number of successful extensions: 8351
Number of sequences better than 10.0: 60
Number of HSP's better than 10.0 without gapping: 2763
Number of HSP's successfully gapped in prelim test: 503
Number of HSP's that attempted gapping in prelim test: 5218
Number of HSP's gapped (non-prelim): 3795
length of query: 1664
length of database: 9,014,727
effective HSP length: 109
effective length of query: 1555
effective length of database: 4,984,343
effective search space: 7750653365
effective search space used: 7750653365
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.7 bits)
S2: 65 (29.6 bits)
Medicago: description of AC147002.8


