

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147000.4 + phase: 1 /pseudo
(187 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
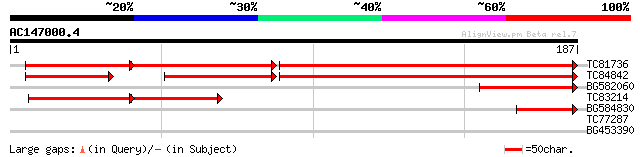
TC81736 similar to SP|O49204|KAPS_CATRO Adenylylsulfate kinase ... 199 6e-90
TC84842 similar to GP|6006852|gb|AAF00628.1| putative adenylylsu... 146 7e-37
BG582060 homologue to PIR|T26219|T262 hypothetical protein W06B1... 66 7e-12
TC83214 similar to SP|O49204|KAPS_CATRO Adenylylsulfate kinase ... 42 9e-10
BG584830 homologue to GP|546360|gb|AA ferric leghemoglobin reduc... 42 1e-04
TC77287 SP|Q02735|CAPP_MEDSA Phosphoenolpyruvate carboxylase (EC... 27 3.6
BG453390 similar to GP|22202666|dbj putative male sterility 1 pr... 27 4.6
>TC81736 similar to SP|O49204|KAPS_CATRO Adenylylsulfate kinase chloroplast
precursor (EC 2.7.1.25) (APS kinase), partial (65%)
Length = 1033
Score = 199 bits (506), Expect(3) = 6e-90
Identities = 95/98 (96%), Positives = 95/98 (96%)
Frame = +2
Query: 90 LISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPY 149
LISPYQKDRDACRALLPEGD IEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPY
Sbjct: 668 LISPYQKDRDACRALLPEGDIIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPY 847
Query: 150 EPPCCCEIILQQKGSDCKSPKDMAETVISYLEKSGHLQ 187
EPP CCEIILQQKGSDCKSP DMAETVISYLEKSGHLQ
Sbjct: 848 EPPWCCEIILQQKGSDCKSPIDMAETVISYLEKSGHLQ 961
Score = 95.9 bits (237), Expect(3) = 6e-90
Identities = 47/48 (97%), Positives = 48/48 (99%)
Frame = +1
Query: 41 QYSCMCFESKLALQRKTDLHP*R*QYSAWSKP*S*F*SRRSFRKHTKD 88
+YSCMCFESKLALQRKTDLHP*R*QYSAWSKP*S*F*SRRSFRKHTKD
Sbjct: 472 KYSCMCFESKLALQRKTDLHP*R*QYSAWSKP*S*F*SRRSFRKHTKD 615
Score = 73.9 bits (180), Expect(3) = 6e-90
Identities = 36/36 (100%), Positives = 36/36 (100%)
Frame = +3
Query: 6 TRRTFCGMIVQFKNVIDSSCFSKKDVLYG*LVSVVQ 41
TRRTFCGMIVQFKNVIDSSCFSKKDVLYG*LVSVVQ
Sbjct: 360 TRRTFCGMIVQFKNVIDSSCFSKKDVLYG*LVSVVQ 467
>TC84842 similar to GP|6006852|gb|AAF00628.1| putative adenylylsulfate
kinase {Arabidopsis thaliana}, partial (98%)
Length = 1003
Score = 146 bits (368), Expect(3) = 7e-37
Identities = 65/98 (66%), Positives = 83/98 (84%)
Frame = +3
Query: 90 LISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPY 149
LISPY++DRD CRA+LP+ +FIEV++++PL +CEARDPKGLYKLARAGKIKGFTGIDDPY
Sbjct: 522 LISPYRRDRDTCRAMLPDANFIEVYMNMPLSLCEARDPKGLYKLARAGKIKGFTGIDDPY 701
Query: 150 EPPCCCEIILQQKGSDCKSPKDMAETVISYLEKSGHLQ 187
EPP CEI +++ DC +PK MA V++YLE+ G L+
Sbjct: 702 EPPLNCEIEIKEDDGDCPTPKVMAGQVVTYLEEKGFLE 815
Score = 24.3 bits (51), Expect(3) = 7e-37
Identities = 11/29 (37%), Positives = 19/29 (64%)
Frame = +3
Query: 6 TRRTFCGMIVQFKNVIDSSCFSKKDVLYG 34
T++ + G IV +++ S +K+DVLYG
Sbjct: 213 TQQIYFGKIVN*ESLKGRSYLTKRDVLYG 299
Score = 20.4 bits (41), Expect(3) = 7e-37
Identities = 15/37 (40%), Positives = 23/37 (61%)
Frame = +2
Query: 52 ALQRKTDLHP*R*QYSAWSKP*S*F*SRRSFRKHTKD 88
AL+ K L+P* SAW+K S F*+ R K++++
Sbjct: 359 ALKGKIILYP*WR*PSAWTKQGSWF*T*RPH*KYSQN 469
>BG582060 homologue to PIR|T26219|T262 hypothetical protein W06B11.1 -
Caenorhabditis elegans, partial (4%)
Length = 802
Score = 66.2 bits (160), Expect = 7e-12
Identities = 31/32 (96%), Positives = 32/32 (99%)
Frame = +3
Query: 156 EIILQQKGSDCKSPKDMAETVISYLEKSGHLQ 187
+IILQQKGSDCKSPKDMAETVISYLEKSGHLQ
Sbjct: 432 QIILQQKGSDCKSPKDMAETVISYLEKSGHLQ 527
Score = 34.3 bits (77), Expect = 0.029
Identities = 10/14 (71%), Positives = 13/14 (92%)
Frame = +3
Query: 146 DDPYEPPCCCEIIL 159
+DPYEPPCCCE+ +
Sbjct: 3 EDPYEPPCCCEVCI 44
>TC83214 similar to SP|O49204|KAPS_CATRO Adenylylsulfate kinase chloroplast
precursor (EC 2.7.1.25) (APS kinase), partial (23%)
Length = 603
Score = 41.6 bits (96), Expect(2) = 9e-10
Identities = 22/35 (62%), Positives = 29/35 (82%)
Frame = +1
Query: 7 RRTFCGMIVQFKNVIDSSCFSKKDVLYG*LVSVVQ 41
++TF GM VQ +++IDS+ F+KK VL G*LVSVVQ
Sbjct: 403 QQTFYGMNVQSRSLIDSNYFNKKAVLSG*LVSVVQ 507
Score = 37.4 bits (85), Expect(2) = 9e-10
Identities = 18/30 (60%), Positives = 24/30 (80%)
Frame = +3
Query: 41 QYSCMCFESKLALQRKTDLHP*R*QYSAWS 70
++S M FESKLAL+RK +++P* * YS WS
Sbjct: 513 KHSSMRFESKLALERKVNIYP*W**YSTWS 602
>BG584830 homologue to GP|546360|gb|AA ferric leghemoglobin reductase; FLbR
{Glycine max}, partial (13%)
Length = 759
Score = 42.4 bits (98), Expect = 1e-04
Identities = 19/20 (95%), Positives = 20/20 (100%)
Frame = -2
Query: 168 SPKDMAETVISYLEKSGHLQ 187
+PKDMAETVISYLEKSGHLQ
Sbjct: 545 TPKDMAETVISYLEKSGHLQ 486
>TC77287 SP|Q02735|CAPP_MEDSA Phosphoenolpyruvate carboxylase (EC 4.1.1.31)
(PEPCASE). [Alfalfa] {Medicago sativa}, partial (33%)
Length = 1124
Score = 27.3 bits (59), Expect = 3.6
Identities = 11/30 (36%), Positives = 16/30 (52%), Gaps = 8/30 (26%)
Frame = +3
Query: 153 CCCEII--------LQQKGSDCKSPKDMAE 174
CCC+++ L +GSDC PK+ E
Sbjct: 393 CCCQVLFTHA*LGQLS*RGSDCAPPKEQVE 482
>BG453390 similar to GP|22202666|dbj putative male sterility 1 protein {Oryza
sativa (japonica cultivar-group)}, partial (5%)
Length = 655
Score = 26.9 bits (58), Expect = 4.6
Identities = 17/45 (37%), Positives = 20/45 (43%), Gaps = 7/45 (15%)
Frame = +3
Query: 149 YEPPC-CCEIILQQKGSDCKS------PKDMAETVISYLEKSGHL 186
Y PC CC IL S CKS D+ + V LE + HL
Sbjct: 465 YHKPCMCCGDILHLSESKCKSCNHVTTSDDVEDWVYHQLENTSHL 599
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.340 0.150 0.512
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,986,225
Number of Sequences: 36976
Number of extensions: 96766
Number of successful extensions: 613
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 610
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 613
length of query: 187
length of database: 9,014,727
effective HSP length: 91
effective length of query: 96
effective length of database: 5,649,911
effective search space: 542391456
effective search space used: 542391456
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 55 (25.8 bits)
Medicago: description of AC147000.4


