

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146972.11 + phase: 0
(362 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
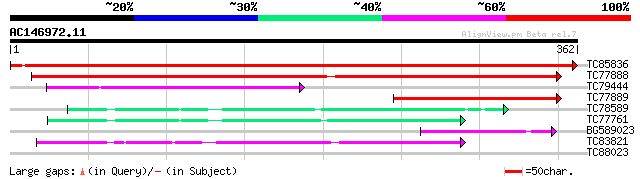
TC85836 homologue to SP|P52904|ODPB_PEA Pyruvate dehydrogenase E... 705 0.0
TC77888 homologue to PIR|E84758|E84758 probable pyruvate dehydro... 266 1e-71
TC79444 homologue to PIR|T51835|T51835 3-methyl-2-oxobutanoate d... 111 5e-25
TC77889 homologue to GP|13605515|gb|AAK32751.1 At1g30120/T2H7_8 ... 90 2e-18
TC78589 1-deoxy-D-xylulose 5-phosphate synthase 2 [Medicago trun... 67 8e-12
TC77761 1-deoxy-D-xylulose 5-phosphate synthase 1 [Medicago trun... 67 1e-11
BG589023 homologue to PIR|T51835|T518 3-methyl-2-oxobutanoate de... 63 2e-10
TC83821 similar to GP|8953400|emb|CAB96673.1 1-D-deoxyxylulose 5... 62 5e-10
TC88023 similar to PIR|T50009|T50009 hypothetical protein T31P16... 27 9.4
>TC85836 homologue to SP|P52904|ODPB_PEA Pyruvate dehydrogenase E1 component
beta subunit mitochondrial precursor (EC 1.2.4.1)
(PDHE1-B)., complete
Length = 1507
Score = 705 bits (1820), Expect = 0.0
Identities = 361/362 (99%), Positives = 361/362 (99%)
Frame = +2
Query: 1 MLGVIRNKVNLLRPSFSAFRHLSSSAKQMTVRDALNSALDEEMSADPKVFLMGEEVGEYQ 60
MLGVIRNK NLLRPSFSAFRHLSSSAKQMTVRDALNSALDEEMSADPKVFLMGEEVGEYQ
Sbjct: 113 MLGVIRNK-NLLRPSFSAFRHLSSSAKQMTVRDALNSALDEEMSADPKVFLMGEEVGEYQ 289
Query: 61 GAYKISKGLLEKYGPERVLDTPITEAGFTGIGVGAAYYGLKPVVEFMTFNFSMQAIDHII 120
GAYKISKGLLEKYGPERVLDTPITEAGFTGIGVGAAYYGLKPVVEFMTFNFSMQAIDHII
Sbjct: 290 GAYKISKGLLEKYGPERVLDTPITEAGFTGIGVGAAYYGLKPVVEFMTFNFSMQAIDHII 469
Query: 121 NSAAKSNYMSAGQINVPIVFRGPNGAAAGVGAQHSHCYASWYGSCPGLKVLAPYSSEDAR 180
NSAAKSNYMSAGQINVPIVFRGPNGAAAGVGAQHSHCYASWYGSCPGLKVLAPYSSEDAR
Sbjct: 470 NSAAKSNYMSAGQINVPIVFRGPNGAAAGVGAQHSHCYASWYGSCPGLKVLAPYSSEDAR 649
Query: 181 GLLKAAIRDPDPVVFLENELLYGESFPVSAEVLDSSFCLPIGKAKIEREGKDVTITAFSK 240
GLLKAAIRDPDPVVFLENELLYGESFPVSAEVLDSSFCLPIGKAKIEREGKDVTITAFSK
Sbjct: 650 GLLKAAIRDPDPVVFLENELLYGESFPVSAEVLDSSFCLPIGKAKIEREGKDVTITAFSK 829
Query: 241 MVGFALKAAETLEKEGISAEVINLRSIRPLDRATINASVRKTNRLVTVEEGFPQHGVGAE 300
MVGFALKAAETLEKEGISAEVINLRSIRPLDRATINASVRKTNRLVTVEEGFPQHGVGAE
Sbjct: 830 MVGFALKAAETLEKEGISAEVINLRSIRPLDRATINASVRKTNRLVTVEEGFPQHGVGAE 1009
Query: 301 ICASVIEESFGYLDAPVERIAGADVPMPYAANLERLAVPQIEDIVRAAKRACHRSVPMAA 360
ICASVIEESFGYLDAPVERIAGADVPMPYAANLERLAVPQIEDIVRAAKRACHRSVPMAA
Sbjct: 1010ICASVIEESFGYLDAPVERIAGADVPMPYAANLERLAVPQIEDIVRAAKRACHRSVPMAA 1189
Query: 361 TA 362
TA
Sbjct: 1190TA 1195
>TC77888 homologue to PIR|E84758|E84758 probable pyruvate dehydrogenase E1
beta subunit [imported] - Arabidopsis thaliana, partial
(85%)
Length = 1707
Score = 266 bits (679), Expect = 1e-71
Identities = 138/338 (40%), Positives = 208/338 (60%)
Frame = +2
Query: 15 SFSAFRHLSSSAKQMTVRDALNSALDEEMSADPKVFLMGEEVGEYQGAYKISKGLLEKYG 74
S SA S ++ + +AL L+EEM DP V +MGE+VG Y G+YK+++ L EK+G
Sbjct: 389 SSSATSAASKPGHELLLFEALREGLEEEMERDPCVCVMGEDVGHYGGSYKVTRNLAEKFG 568
Query: 75 PERVLDTPITEAGFTGIGVGAAYYGLKPVVEFMTFNFSMQAIDHIINSAAKSNYMSAGQI 134
RVLDTPI E FTG+G+GAA GL+P++E M F + A + I N+ +Y S GQ
Sbjct: 569 DLRVLDTPIAENAFTGMGIGAAMTGLRPIIEGMNMGFLLLAFNQISNNCGMLHYTSGGQF 748
Query: 135 NVPIVFRGPNGAAAGVGAQHSHCYASWYGSCPGLKVLAPYSSEDARGLLKAAIRDPDPVV 194
+P+V RGP G +GA+HS S++ S PG++++A + +A+GL+KAAIR +PV+
Sbjct: 749 KIPVVIRGPGGVGRQLGAEHSQRLESYFQSIPGIQMVACSTPYNAKGLMKAAIRSDNPVI 928
Query: 195 FLENELLYGESFPVSAEVLDSSFCLPIGKAKIEREGKDVTITAFSKMVGFALKAAETLEK 254
E+ LLY + + D + L + +A++ R G+ VTI +S+M ++AA+TL
Sbjct: 929 LFEHVLLYN----LKERIPDEEYVLSLEEAEMVRPGEHVTILTYSRMRYHVMQAAKTLVN 1096
Query: 255 EGISAEVINLRSIRPLDRATINASVRKTNRLVTVEEGFPQHGVGAEICASVIEESFGYLD 314
+G EVI++RS++P D TI S++KT+R++ VEE G+GA + A++ E YLD
Sbjct: 1097KGYDPEVIDIRSLKPFDLHTIGNSIKKTHRVLIVEECMRTGGIGASLTAAITENFHDYLD 1276
Query: 315 APVERIAGADVPMPYAANLERLAVPQIEDIVRAAKRAC 352
AP+ ++ DVP PYA LE V Q IV A ++ C
Sbjct: 1277APIICLSSQDVPTPYAGTLEEWTVVQPAQIVTAVEQLC 1390
>TC79444 homologue to PIR|T51835|T51835 3-methyl-2-oxobutanoate
dehydrogenase (lipoamide) (EC 1.2.4.4) beta chain
[imported] - Arabidopsis, partial (46%)
Length = 849
Score = 111 bits (277), Expect = 5e-25
Identities = 57/166 (34%), Positives = 92/166 (55%), Gaps = 1/166 (0%)
Frame = +2
Query: 24 SSAKQMTVRDALNSALDEEMSADPKVFLMGEEVGEYQGAYKISKGLLEKYGPERVLDTPI 83
+ K + + A+N AL + DP+ ++ GE+VG + G ++ + GL +++G +RV +TP+
Sbjct: 353 NGVKSLNLYSAVNQALHIALDTDPRAYVFGEDVG-FGGVFRCTTGLADRFGKKRVFNTPL 529
Query: 84 TEAGFTGIGVGAAYYGLKPVVEFMTFNFSMQAIDHIINSAAKSNYMSAGQINV-PIVFRG 142
E G G G+G A G + + E ++ A D I+N AAK Y S Q N + R
Sbjct: 530 CEQGIVGFGIGLAAMGNRAIAEIQFADYIFPAFDQIVNEAAKFRYRSGNQFNCGGLTIRT 709
Query: 143 PNGAAAGVGAQHSHCYASWYGSCPGLKVLAPYSSEDARGLLKAAIR 188
P GA G HS +++ PG+KV+ P S ++A+GLL + IR
Sbjct: 710 PYGAVGHGGHYHSQSPEAFFCHVPGIKVVIPRSPKEAKGLLVSCIR 847
>TC77889 homologue to GP|13605515|gb|AAK32751.1 At1g30120/T2H7_8
{Arabidopsis thaliana}, partial (28%)
Length = 653
Score = 89.7 bits (221), Expect = 2e-18
Identities = 45/107 (42%), Positives = 67/107 (62%)
Frame = +3
Query: 246 LKAAETLEKEGISAEVINLRSIRPLDRATINASVRKTNRLVTVEEGFPQHGVGAEICASV 305
++AA+TL +G EVI++RS++P D TI S++KT+R++ VEE G+GA + A++
Sbjct: 21 MQAAKTLVNKGYDPEVIDIRSLKPFDLHTIGNSIKKTHRVLIVEECMRTGGIGASLTAAI 200
Query: 306 IEESFGYLDAPVERIAGADVPMPYAANLERLAVPQIEDIVRAAKRAC 352
E YLDAP+ ++ DVP PYA LE V Q IV A ++ C
Sbjct: 201 TENFHDYLDAPIICLSSQDVPTPYAGTLEEWTVVQPAQIVTAVEQLC 341
>TC78589 1-deoxy-D-xylulose 5-phosphate synthase 2 [Medicago truncatula]
Length = 2760
Score = 67.4 bits (163), Expect = 8e-12
Identities = 73/284 (25%), Positives = 111/284 (38%), Gaps = 3/284 (1%)
Frame = +3
Query: 38 ALDEEMSADPKVFLMGEEVGEYQGAYKISKGLLEKYGPERVLDTPITEAGFTGIGVGAAY 97
+L +E D K+ + +G G +K P+R D I E G A
Sbjct: 1386 SLIKEAEMDNKIVAIHAAMGGGTGL-----NYFQKRFPDRCFDVGIAEQHAVTFAAGLAA 1550
Query: 98 YGLKPVVEFMTFNFSMQAIDHIINSAAKSNYMSAGQINVPIVFRGPNGAAAGV-GAQHSH 156
GLKP + +F + D +++ +P+ F G G H
Sbjct: 1551 EGLKPFCAIYS-SFLQRGYDQVVHDVDLQK--------LPVRFAMDRAGLVGADGPTHCG 1703
Query: 157 CYASWYGSC-PGLKVLAPYSSEDARGLLK-AAIRDPDPVVFLENELLYGESFPVSAEVLD 214
+ + +C P + V+AP + ++ AA D P F G + + +
Sbjct: 1704 AFDITFMACLPNMIVMAPSDEAELMNMVATAAAIDDRPSCF---RFPRGNGIGANLPLNN 1874
Query: 215 SSFCLPIGKAKIEREGKDVTITAFSKMVGFALKAAETLEKEGISAEVINLRSIRPLDRAT 274
L IGK +I EG V I + MV +KAAE L G+ V + R +PLD
Sbjct: 1875 KGTPLEIGKGRILLEGSRVAILGYGCMVQQCMKAAEMLRAVGVYVTVADARFCKPLDTDL 2054
Query: 275 INASVRKTNRLVTVEEGFPQHGVGAEICASVIEESFGYLDAPVE 318
I R+ L+TVEEG G G+ + S G LD P++
Sbjct: 2055 IRLLAREHEILITVEEG-SIGGFGSHV--SQFLSLAGLLDGPLK 2177
>TC77761 1-deoxy-D-xylulose 5-phosphate synthase 1 [Medicago truncatula]
Length = 2595
Score = 66.6 bits (161), Expect = 1e-11
Identities = 69/270 (25%), Positives = 104/270 (37%), Gaps = 3/270 (1%)
Frame = +3
Query: 25 SAKQMTVRDALNSALDEEMSADPKVFLMGEEVGEYQGAYKISKGLLEKYGPERVLDTPIT 84
+AK + AL E AD + + +G G + + P R D I
Sbjct: 1299 AAKTQSYTTYFAEALIAEAKADKDIIAIHAAMGGGTGM-----NIFHRRFPTRCFDVGIA 1463
Query: 85 EAGFTGIGVGAAYYGLKPVVEFMTFNFSMQAIDHIINSAAKSNYMSAGQINVPIVFRGPN 144
E G A GLKP + +F +A D +++ +P+ F
Sbjct: 1464 EQHAVTFAAGLACEGLKPFCAIYS-SFLQRAYDQVVHDVDLQK--------LPVRFAMDR 1616
Query: 145 GAAAGV-GAQHSHCYASWYGSC-PGLKVLAPYS-SEDARGLLKAAIRDPDPVVFLENELL 201
G G HS + + +C P + V+AP +E + AA D P F
Sbjct: 1617 AGLVGSDGPTHSGSFDVTFMACLPNMVVMAPSDEAELCHMVATAAAIDDRPSCFRYPR-- 1790
Query: 202 YGESFPVSAEVLDSSFCLPIGKAKIEREGKDVTITAFSKMVGFALKAAETLEKEGISAEV 261
G V L IGK +I EG+ V + + V L AA +E+ G+ V
Sbjct: 1791 -GNGIGVELPTEYKGIPLEIGKGRILIEGERVALLGYGSAVQNCLAAASLVEQHGLRLTV 1967
Query: 262 INLRSIRPLDRATINASVRKTNRLVTVEEG 291
+ R +PLDR+ I + + L+TVEEG
Sbjct: 1968 ADARFCKPLDRSLIRSLAKSHEVLITVEEG 2057
>BG589023 homologue to PIR|T51835|T518 3-methyl-2-oxobutanoate dehydrogenase
(lipoamide) (EC 1.2.4.4) beta chain [imported] -
Arabidopsis, partial (25%)
Length = 486
Score = 63.2 bits (152), Expect = 2e-10
Identities = 35/87 (40%), Positives = 49/87 (56%)
Frame = +2
Query: 263 NLRSIRPLDRATINASVRKTNRLVTVEEGFPQHGVGAEICASVIEESFGYLDAPVERIAG 322
+L+++ P D+ T+ ASV+KT RL+ E G GAEI AS++E F L+APV R+ G
Sbjct: 2 DLKTLIPWDKETVEASVKKTGRLLISHEAPVTGGFGAEISASILERCFSRLEAPVARVCG 181
Query: 323 ADVPMPYAANLERLAVPQIEDIVRAAK 349
D P P E +P I+ A K
Sbjct: 182 LDTPFPLV--FEPFYMPTKNKILDAIK 256
>TC83821 similar to GP|8953400|emb|CAB96673.1 1-D-deoxyxylulose 5-phosphate
synthase-like protein {Arabidopsis thaliana}, partial
(46%)
Length = 1172
Score = 61.6 bits (148), Expect = 5e-10
Identities = 70/278 (25%), Positives = 112/278 (40%), Gaps = 4/278 (1%)
Frame = +1
Query: 18 AFRHLSSSAKQMTVRDALNSALDEEMSADPKVFLMGEEVGEYQGAYKISKGLLEKYGPER 77
+F L ++ + T D +L E D + ++ + K +EK+ P+R
Sbjct: 205 SFDLLDNAGRLQTYGDCFVESLVAEAEKDKDIVVVHAGITTEPSL----KLFMEKF-PDR 369
Query: 78 VLDTPITEAGFTGIGVGAAYYGLKPVVEFMTFNFSMQAIDHIINSAAKSNYMSAGQINVP 137
+ + I E G + GLKP + +F +A D +++ Q VP
Sbjct: 370 IFNVGIAEQHAVTFASGLSCGGLKPFC-IIPSSFLQRAYDQVVHDV--------DQQKVP 522
Query: 138 IVFRGPNGAAAGV-GAQHSHCYASWYGSC-PGLKVLAPYSSEDARGLLKAAIRDPD-PVV 194
+ F + G G + + SC P + V+AP + ++ A D PV
Sbjct: 523 VRFVITSARLVGSDGPLQCGAFDITFMSCLPNMIVMAPSDEAELVHMVATAAHINDQPVC 702
Query: 195 F-LENELLYGESFPVSAEVLDSSFCLPIGKAKIEREGKDVTITAFSKMVGFALKAAETLE 253
F L G+ E + + IGK +I EGKDV + + MV LKA L
Sbjct: 703 FRYPRGALVGKD-----EAILDGIPIEIGKGRILVEGKDVALLGYGSMVQNCLKAYSLLA 867
Query: 254 KEGISAEVINLRSIRPLDRATINASVRKTNRLVTVEEG 291
GI V + R +PLD + + + L+TVEEG
Sbjct: 868 NLGIEVTVADARFCKPLDIELLRQLCKHHSFLITVEEG 981
>TC88023 similar to PIR|T50009|T50009 hypothetical protein T31P16.40 -
Arabidopsis thaliana, partial (35%)
Length = 559
Score = 27.3 bits (59), Expect = 9.4
Identities = 22/85 (25%), Positives = 39/85 (45%), Gaps = 2/85 (2%)
Frame = +1
Query: 10 NLLRPSFSA--FRHLSSSAKQMTVRDALNSALDEEMSADPKVFLMGEEVGEYQGAYKISK 67
N L SF+A R +++S + T+ D + DPK FL +V + ++
Sbjct: 166 NALARSFAANSCRVVATSRSRSTMAD---------LDQDPKFFLQELDVQSDESVNRVVN 318
Query: 68 GLLEKYGPERVLDTPITEAGFTGIG 92
+L+K+G +D + AG +G
Sbjct: 319 NVLDKFGR---IDVLVNNAGVPCVG 384
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.318 0.134 0.383
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,969,580
Number of Sequences: 36976
Number of extensions: 104443
Number of successful extensions: 466
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 462
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 462
length of query: 362
length of database: 9,014,727
effective HSP length: 97
effective length of query: 265
effective length of database: 5,428,055
effective search space: 1438434575
effective search space used: 1438434575
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 59 (27.3 bits)
Medicago: description of AC146972.11


