

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
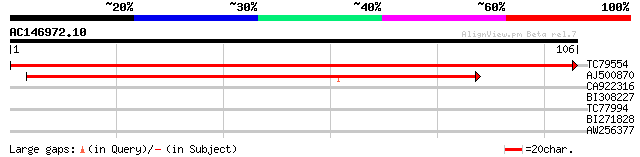
Query= AC146972.10 - phase: 0
(106 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC79554 homologue to SP|Q39236|T2AG_ARATH Transcription initiati... 213 9e-57
AJ500870 similar to SP|Q39236|T2AG_ Transcription initiation fac... 113 1e-26
CA922316 similar to PIR|F96807|F968 unknown protein T32E8.10 [im... 25 3.6
BI308227 similar to GP|17978937|gb AT4g32850/T16I18_60 {Arabidop... 25 4.7
TC77994 weakly similar to PIR|T39903|T39903 serine-rich protein ... 25 4.7
BI271828 similar to GP|10177162|dbj contains similarity to myb-r... 25 6.1
AW256377 24 8.0
>TC79554 homologue to SP|Q39236|T2AG_ARATH Transcription initiation factor
IIA gamma chain (TFIIA-gamma). [Mouse-ear cress],
complete
Length = 708
Score = 213 bits (542), Expect = 9e-57
Identities = 106/106 (100%), Positives = 106/106 (100%)
Frame = +1
Query: 1 MATFELYRRSTIGMCLTETLDEMVQNGTLSPEIAIQVLVQFDKSMTEALETQVKSKVSIK 60
MATFELYRRSTIGMCLTETLDEMVQNGTLSPEIAIQVLVQFDKSMTEALETQVKSKVSIK
Sbjct: 85 MATFELYRRSTIGMCLTETLDEMVQNGTLSPEIAIQVLVQFDKSMTEALETQVKSKVSIK 264
Query: 61 GHLHTYRFCDNVWTFILQDALFKNEDNQENVGRVKIVACDSKLLSQ 106
GHLHTYRFCDNVWTFILQDALFKNEDNQENVGRVKIVACDSKLLSQ
Sbjct: 265 GHLHTYRFCDNVWTFILQDALFKNEDNQENVGRVKIVACDSKLLSQ 402
>AJ500870 similar to SP|Q39236|T2AG_ Transcription initiation factor IIA
gamma chain (TFIIA-gamma). [Mouse-ear cress], partial
(57%)
Length = 357
Score = 113 bits (283), Expect = 1e-26
Identities = 51/86 (59%), Positives = 74/86 (85%), Gaps = 1/86 (1%)
Frame = +1
Query: 4 FELYRRSTIGMCLTETLDEMVQNGTLSPEIAIQVLVQFDKSMTEALETQVKSKVSIK-GH 62
+ELYRRS+IG+ LT++LDE++Q ++P++A++VLVQFDKS+TEAL+T+V++K + K GH
Sbjct: 88 YELYRRSSIGIALTDSLDELIQKEKITPQLAMKVLVQFDKSITEALDTKVRNKATFKAGH 267
Query: 63 LHTYRFCDNVWTFILQDALFKNEDNQ 88
LHTYRFCD VWTFI+++ FK E +Q
Sbjct: 268 LHTYRFCDEVWTFIIKNPNFKLESDQ 345
>CA922316 similar to PIR|F96807|F968 unknown protein T32E8.10 [imported] -
Arabidopsis thaliana, partial (7%)
Length = 746
Score = 25.4 bits (54), Expect = 3.6
Identities = 9/18 (50%), Positives = 14/18 (77%)
Frame = -3
Query: 66 YRFCDNVWTFILQDALFK 83
+RFC ++WT+I QD + K
Sbjct: 537 FRFCCSLWTWIPQDTVGK 484
>BI308227 similar to GP|17978937|gb AT4g32850/T16I18_60 {Arabidopsis
thaliana}, partial (27%)
Length = 761
Score = 25.0 bits (53), Expect = 4.7
Identities = 13/35 (37%), Positives = 20/35 (57%), Gaps = 3/35 (8%)
Frame = -1
Query: 2 ATFELYRRSTIG---MCLTETLDEMVQNGTLSPEI 33
AT + ++ T+G +C TET D +GT+ EI
Sbjct: 686 ATLQPFKLRTVGSSILCRTETSDISNSSGTIKQEI 582
>TC77994 weakly similar to PIR|T39903|T39903 serine-rich protein - fission
yeast (Schizosaccharomyces pombe), partial (5%)
Length = 1571
Score = 25.0 bits (53), Expect = 4.7
Identities = 20/67 (29%), Positives = 26/67 (37%)
Frame = -1
Query: 34 AIQVLVQFDKSMTEALETQVKSKVSIKGHLHTYRFCDNVWTFILQDALFKNEDNQENVGR 93
A +VL +FDK + +ET N T L D LFKN+ + V
Sbjct: 710 AYEVLQKFDKMASGGIETS-----------------SNRETLTLLDRLFKNKPPSDPVSD 582
Query: 94 VKIVACD 100
ACD
Sbjct: 581 EVTRACD 561
>BI271828 similar to GP|10177162|dbj contains similarity to myb-related
transcription factor~gene_id:K21P3.23 {Arabidopsis
thaliana}, partial (5%)
Length = 607
Score = 24.6 bits (52), Expect = 6.1
Identities = 12/44 (27%), Positives = 20/44 (45%)
Frame = +1
Query: 29 LSPEIAIQVLVQFDKSMTEALETQVKSKVSIKGHLHTYRFCDNV 72
+ PE+ I D +TE + V+ K + TY CD++
Sbjct: 271 IEPEVQIPRTPSLDCEVTEKIIDDVEVKTTHDEKETTYELCDDI 402
>AW256377
Length = 607
Score = 24.3 bits (51), Expect = 8.0
Identities = 11/38 (28%), Positives = 23/38 (59%)
Frame = +1
Query: 29 LSPEIAIQVLVQFDKSMTEALETQVKSKVSIKGHLHTY 66
L PE+ I +L +FD + ++T+++S ++ L +Y
Sbjct: 460 LPPELKILILYRFDLI*IDFIKTKIQSSDNMPSLLSSY 573
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.320 0.133 0.377
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,760,271
Number of Sequences: 36976
Number of extensions: 27346
Number of successful extensions: 150
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 148
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 149
length of query: 106
length of database: 9,014,727
effective HSP length: 82
effective length of query: 24
effective length of database: 5,982,695
effective search space: 143584680
effective search space used: 143584680
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 50 (23.9 bits)
Medicago: description of AC146972.10


