

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146971.7 - phase: 0 /pseudo
(2360 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
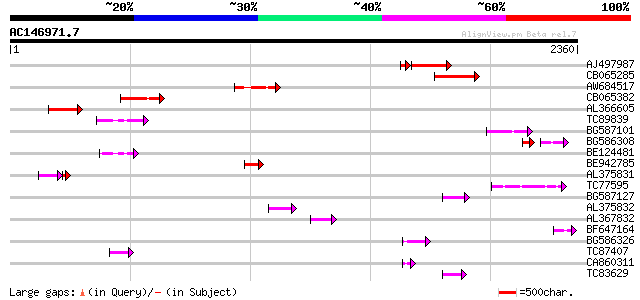
Score E
Sequences producing significant alignments: (bits) Value
AJ497987 weakly similar to GP|9927273|dbj Similar to Arabidopsis... 350 3e-96
CB065285 weakly similar to PIR|A84500|A84 probable retroelement ... 302 1e-81
AW684517 similar to GP|9927273|dbj Similar to Arabidopsis thalia... 183 8e-46
CB065382 similar to PIR|T09671|T096 RPE15 protein - alfalfa (fra... 163 6e-40
AL366605 136 8e-32
TC89839 homologue to SP|P23466|CYAA_SACKL Adenylate cyclase (EC ... 115 2e-25
BG587101 similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana... 109 1e-23
BG586308 weakly similar to PIR|F84528|F8 probable retroelement p... 65 5e-19
BE124481 94 6e-19
BE942785 92 2e-18
AL375831 75 5e-18
TC77595 weakly similar to PIR|T18350|T18350 probable pol polypro... 79 2e-14
BG587127 weakly similar to PIR|H84506|H84 probable retroelement ... 68 5e-11
AL375832 59 2e-08
AL367832 weakly similar to GP|14091845|gb Putative retroelement ... 54 9e-07
BF647164 weakly similar to GP|6691191|gb F7F22.15 {Arabidopsis t... 50 1e-05
BG586326 similar to PIR|G84493|G8 probable retroelement pol poly... 48 5e-05
TC87407 similar to PIR|T49945|T49945 periaxin-like protein - Ara... 47 7e-05
CA860311 weakly similar to GP|7289872|gb|A CG17427 gene product ... 46 1e-04
TC83629 weakly similar to GP|6692261|gb|AAF24611.1| putative RNa... 45 3e-04
>AJ497987 weakly similar to GP|9927273|dbj Similar to Arabidopsis thaliana
chromosome II BAC F26H6; putative retroelement pol
polyprotein, partial (1%)
Length = 636
Score = 350 bits (899), Expect = 3e-96
Identities = 164/167 (98%), Positives = 165/167 (98%)
Frame = -2
Query: 1672 KTCCALAWAAKRLRHYMINHTTWLISKMDPIKYIFEKPALIGRIARWQMLLSEYDIEYRS 1731
KTCCALAWAAKRLRHYMINHTTWL+SKMDPIKYIFEKPAL GRIARWQMLLSEYDIEYRS
Sbjct: 635 KTCCALAWAAKRLRHYMINHTTWLVSKMDPIKYIFEKPALTGRIARWQMLLSEYDIEYRS 456
Query: 1732 QKAIKGSILADHLAHQPLEDYRPIKFDFPDEEIMYLKMKDCDEPLFGEGPDPDSVWGLIF 1791
QKAIKGSILADHLAHQPLEDYRPIKFDFPDEEIMYLKMKDCDEPLFGEGPDPDSVWGLIF
Sbjct: 455 QKAIKGSILADHLAHQPLEDYRPIKFDFPDEEIMYLKMKDCDEPLFGEGPDPDSVWGLIF 276
Query: 1792 DGAVNVYGNGIGAVLLTPKGTHIPFTARLRFDCTNNIAEYEACIMGI 1838
DGAVNVYGNGIGAVLLTPKG HIPFTARLRFDCTNNIAEYEACIMGI
Sbjct: 275 DGAVNVYGNGIGAVLLTPKGAHIPFTARLRFDCTNNIAEYEACIMGI 135
Score = 53.5 bits (127), Expect = 9e-07
Identities = 30/44 (68%), Positives = 33/44 (74%)
Frame = +2
Query: 1625 *SYT*RC*ENSMGCVLGQQDETGRKEHAIYYLSKKFTECESRYS 1668
*S T*+C* N CVLGQQDETGRKEHAIYYLS+K + R S
Sbjct: 2 *SCT*QC*-NPWVCVLGQQDETGRKEHAIYYLSRKLSSTVWRIS 130
>CB065285 weakly similar to PIR|A84500|A84 probable retroelement gag/pol
polyprotein [imported] - Arabidopsis thaliana, partial
(2%)
Length = 592
Score = 302 bits (773), Expect = 1e-81
Identities = 143/196 (72%), Positives = 168/196 (84%), Gaps = 8/196 (4%)
Frame = -2
Query: 1768 KMKDCDEPLFGEGPDPDSVWGLIFDGAVNVYGNGIGAVLLTPKGTHIPFTARLRFDCTNN 1827
K KDC+EPL GEGPDP+S WGL+FDGAVN YG GIGAV+++P+G +IPFTAR+ F+CTNN
Sbjct: 591 KSKDCEEPLIGEGPDPNSKWGLVFDGAVNAYGKGIGAVIVSPQGHYIPFTARILFECTNN 412
Query: 1828 IAEYEACIMGIEEAIDLRIKNIEIYGDSALVINQIKGKWETLHAGLIPYRDYARRLLTFF 1887
+AEYEACI GIEEAID+RIK+++IYGDSALVINQIKG+WET HA LIPYRDYARRLLT+F
Sbjct: 411 MAEYEACIFGIEEAIDMRIKHLDIYGDSALVINQIKGEWETHHANLIPYRDYARRLLTYF 232
Query: 1888 NKVELHHIPRDENQMADALATLSSMIKVNHHNDVPLISVKFLDRPAYVFA--------AE 1939
KVELHHIPRDENQMADALATLSSM +VNH NDVP+I V+ L+RP++VFA E
Sbjct: 231 TKVELHHIPRDENQMADALATLSSMFRVNHWNDVPIIKVQRLERPSHVFAIGNVIDQTGE 52
Query: 1940 VVFDDKPWFHDIKVFL 1955
V D KPW++DIK FL
Sbjct: 51 NVVDYKPWYYDIKQFL 4
>AW684517 similar to GP|9927273|dbj Similar to Arabidopsis thaliana chromosome
II BAC F26H6; putative retroelement pol polyprotein,
partial (1%)
Length = 488
Score = 183 bits (464), Expect = 8e-46
Identities = 97/190 (51%), Positives = 124/190 (65%), Gaps = 1/190 (0%)
Frame = +1
Query: 936 VNFQVMDIQASYSCLLGRPWIHEAGAVTSTLHQKLKFVKNGKLVTVNGEEALLVSHLSSF 995
+ FQVMDI ASYSCLLGRPWIH+AGAVTSTLHQKLKFVKNG
Sbjct: 1 ITFQVMDINASYSCLLGRPWIHDAGAVTSTLHQKLKFVKNG------------------- 123
Query: 996 SFIGADDVEGTPFQGFTIEDKNTKRNEASISSLKDAQKVIQAGGSTSWGKLIELPENKHR 1055
EGT FQG ++E K+ A+++SLKDAQK +Q G + WGKLI+L ENK +
Sbjct: 124 ------SAEGTAFQGLSMEGAEPKKVGAAMASLKDAQKAVQEGQAADWGKLIQLCENKRK 285
Query: 1056 EGLGFFPSTGLSTAKKGTFHSSGFIHAIIEDDPESVPRG-FITPGVSSHNWVAVDVPFVA 1114
EGL F P++G+ST GTFHS+GF++ + E+ VPR F+ PG + +W AVDVP +
Sbjct: 286 EGLRFSPTSGVST---GTFHSAGFVNTLAEEVARFVPRPLFVIPGGIAKDWDAVDVPSIM 456
Query: 1115 HLSK*CICLV 1124
H+S *C+ L+
Sbjct: 457 HVSX*CVLLL 486
>CB065382 similar to PIR|T09671|T096 RPE15 protein - alfalfa (fragment),
partial (9%)
Length = 624
Score = 163 bits (413), Expect = 6e-40
Identities = 87/182 (47%), Positives = 114/182 (61%)
Frame = -2
Query: 461 PQQNFQQQPYQQRPQQPRPPRMPINPIPVTYAELLPGLLKKNLVQTRTAPPIPEKLPSWY 520
P +N +QQR Q P P+ IP+ YAE LP LL + R P P+ LP +
Sbjct: 539 PPRNNHPPVHQQRQ*QQARPTFPL--IPMLYAE*LPTLLLRGHCTIRQGKPPPDPLPPRF 366
Query: 521 RLDQTCDFHEGGRGHNIETCYAFKSTVQRLINDGKITFTDSAPNVQTNPLPNHGAATVNM 580
R D CDFH+G GH++E CYA K V++LIN GK+TF ++ P+V NPLPNH A VNM
Sbjct: 365 RSDLKCDFHQGALGHDVEGCYALKHIVKKLINQGKLTFENNVPHVLDNPLPNH--AAVNM 192
Query: 581 IEDCQKTRPILNVQHIRTPLVPLHAKLCKVDLFEHDHDLCEICLMNSGGCQKVRNDIQGL 640
IE ++ P L+V+++ TPLVPLH KLC+ LF+HDH C+ N GC V+NDIQ L
Sbjct: 191 IEVYEEA-PGLDVRNVTTPLVPLHIKLCQASLFDHDHANCQE*FYNPLGCCVVQNDIQSL 15
Query: 641 LN 642
+N
Sbjct: 14 MN 9
>AL366605
Length = 422
Score = 136 bits (343), Expect = 8e-32
Identities = 64/140 (45%), Positives = 93/140 (65%)
Frame = -3
Query: 161 DMKALRDKGKFGKTAYDLCLVPSVQVPHKFKIPDFEKYKGSSCPEEHLKMYVRRMPAYAQ 220
++KA R KT DLCLV V VP KFKIPDF++Y G +CP+ H+ YVR+M Y
Sbjct: 417 EIKANRGNADSFKTQ-DLCLVSKVDVPKKFKIPDFDRYNGLTCPQNHIIKYVRKMGNYKD 241
Query: 221 DDQILIYYFQESLTGPASKWYTNLDKTRVQTFRDLCEAFVEQYSYNVDMTPDRSDLQAMT 280
+D ++I+ FQ+SL A++WYT+L K + TF +L AF Y +N + P+R L++++
Sbjct: 240 NDSLMIHCFQDSLMEDAAEWYTSLSKNDIHTFDELAAAFKSHYGFNTRLKPNREFLRSLS 61
Query: 281 QGDKETFKEYAQRWRDTAAQ 300
Q +E+F+EYAQRWR AA+
Sbjct: 60 QKKEESFREYAQRWRGAAAR 1
>TC89839 homologue to SP|P23466|CYAA_SACKL Adenylate cyclase (EC 4.6.1.1)
(ATP pyrophosphate-lyase) (Adenylyl cyclase). [Yeast],
partial (0%)
Length = 665
Score = 115 bits (289), Expect = 2e-25
Identities = 74/221 (33%), Positives = 99/221 (44%), Gaps = 7/221 (3%)
Frame = +3
Query: 363 TKKTGNHFPR----KKEQEVGMVTHGMPQQTYPAYQHIAAITPTSHPFQQTNNHSQIPQY 418
+ K G+H +K ++ H + + + +T TN+ + Q
Sbjct: 69 SSKFGSHMQTLVQTQKMDQLEQEVHELRGEVTTLRAEVEKLTSLVSSLMATNDPPLVQQR 248
Query: 419 PQIPQYPQIPQYPQFPQNPSPQNTQQQNFQQQPYQQFPYQQYPQQNFQQQPYQQRPQQPR 478
PQ P P PQ P+ QQ P Q P Q PQ+ Q Q+ Q
Sbjct: 249 PQSPYQPMCPQKPR---------------QQAPRQSIPQNQVPQKFIPQNQVQKASQ--- 374
Query: 479 PPRMPINPIPVTYAELLPGLLKKNLVQTRTAPPIPEKLPSWYRLDQTCDFHEGGRGHNIE 538
+PIPV YA+LLP LLKKNL+QT P +P LP WYR D C FH+G GH+ E
Sbjct: 375 -----CDPIPVKYADLLPILLKKNLIQTLPLPRVPNSLPPWYRPDLNCVFHQGAPGHDTE 539
Query: 539 TCYAFKSTVQRLINDGKITFTDSAPNV---QTNPLPNHGAA 576
CY K VQ+LI + +F D V Q + P+ GAA
Sbjct: 540 QCYPLKEEVQKLIENNVWSFDDQDIKVLLQQQHLAPHSGAA 662
>BG587101 similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana}, partial
(10%)
Length = 624
Score = 109 bits (273), Expect = 1e-23
Identities = 60/188 (31%), Positives = 95/188 (49%)
Frame = +2
Query: 1986 LYKRNFDTVLLRCVDKYEADLLIHEIHEGSFGIHPNGHTMAKKILRAGYYWMTMESDCYK 2045
LYK+ D + +RCV + E ++ H ++ H KI +AG++W TM D +
Sbjct: 41 LYKQCADNIYIRCVAEEEIPGILFHCHGSNYAGHFAVSKTVSKIQQAGFWWPTMFKDAHS 220
Query: 2046 HTRKCHKCQIYADKIHMPPTTLNLLSSPWPFSMWGIDMIGRIEPKASNGHRFILVAIDYF 2105
KC CQ + N + F +WGID +G +N ++ILVA+DY
Sbjct: 221 FISKCDPCQRQGNIS*RNEMPQNFILEVEVFDVWGIDFMGPFPSSYNN--KYILVAVDYV 394
Query: 2106 TKWVEAASYANVTKQVVVKFIKNHIICRYGIPNRIITDNGTNLNNKMMKELCDDFKIEHH 2165
+KWVEA + VVVK K+ I R+G+P +I+D G++ NK+ ++L + H
Sbjct: 395 SKWVEAIASPTNDATVVVKMFKSVIFPRFGVPRVVISDGGSHFINKVFEKLLKKNGVRHK 574
Query: 2166 NSSPYRPQ 2173
++ Y PQ
Sbjct: 575 VATAYHPQ 598
>BG586308 weakly similar to PIR|F84528|F8 probable retroelement pol polyprotein
[imported] - Arabidopsis thaliana, partial (7%)
Length = 686
Score = 65.1 bits (157), Expect(2) = 5e-19
Identities = 41/122 (33%), Positives = 69/122 (55%), Gaps = 4/122 (3%)
Frame = -3
Query: 2207 LHGYRTSVRTSIGATPFSLVYGMEAVLPVEVEIPSL---RVLMEVDLSEAEWVQNRYDQL 2263
L +RT+ R + +TPFS+ + +EA+ P EV + SL R+ ++L+ ++ L
Sbjct: 462 LWSHRTNPRGATKSTPFSMAHRVEAMAPAEVNVTSL*RSRMPQNIELNN----DRLFNAL 295
Query: 2264 NLIEEKRMAALCHGQLYQKRMKQAFDKKVRPREFKEGDLVLKKIFSFQPD-SRGKWAPNY 2322
IEE+R AL Q YQ +++ ++K V+ + K GD+VL K+F + + GK N+
Sbjct: 294 ETIEERRDQALLRIQNYQHQIESYYNKTVKSQPLKLGDIVLCKVFENTKELNAGKLGTNW 115
Query: 2323 EG 2324
EG
Sbjct: 114 EG 109
Score = 49.7 bits (117), Expect(2) = 5e-19
Identities = 21/52 (40%), Positives = 34/52 (65%)
Frame = -2
Query: 2134 YGIPNRIITDNGTNLNNKMMKELCDDFKIEHHNSSPYRPQMNGAVEAANKNI 2185
+G+P I+TDNG++ + +E C+ ++I + +SP PQ NG EA+NK I
Sbjct: 685 HGLPYEIVTDNGSHFISNKFREFCERWRIRLNTASPRYPQSNGQAEASNKII 530
>BE124481
Length = 538
Score = 94.0 bits (232), Expect = 6e-19
Identities = 58/162 (35%), Positives = 73/162 (44%)
Frame = +3
Query: 375 EQEVGMVTHGMPQQTYPAYQHIAAITPTSHPFQQTNNHSQIPQYPQIPQYPQIPQYPQFP 434
EQEV H + + + +T TN+ + Q PQ P P PQ P+
Sbjct: 132 EQEV----HELRGEVTTLRAEVEKLTSLVSSLMATNDPPLVQQRPQSPYQPMCPQKPR-- 293
Query: 435 QNPSPQNTQQQNFQQQPYQQFPYQQYPQQNFQQQPYQQRPQQPRPPRMPINPIPVTYAEL 494
QQ P Q P Q PQ+ Q Q+ Q +PIPV YA+L
Sbjct: 294 -------------QQAPRQSIPQNQVPQKFIPQNQVQKASQ--------CDPIPVKYADL 410
Query: 495 LPGLLKKNLVQTRTAPPIPEKLPSWYRLDQTCDFHEGGRGHN 536
LP LLKKNL+QT P +P LP WYR D C FH+G GH+
Sbjct: 411 LPILLKKNLIQTLPLPRVPNSLPPWYRPDLNCVFHQGAPGHD 536
>BE942785
Length = 460
Score = 92.0 bits (227), Expect = 2e-18
Identities = 43/82 (52%), Positives = 59/82 (71%)
Frame = -2
Query: 975 NGKLVTVNGEEALLVSHLSSFSFIGADDVEGTPFQGFTIEDKNTKRNEASISSLKDAQKV 1034
+G LVT++GEEA L+S LSSFS I A EGT FQG T+E KR+ +++SLKDAQ+
Sbjct: 378 SGXLVTIHGEEAYLISQLSSFSCIEAGSAEGTAFQGLTVEGTEPKRDGTAMASLKDAQRA 199
Query: 1035 IQAGGSTSWGKLIELPENKHRE 1056
+Q + WG+LI+L ENKH++
Sbjct: 198 VQESQAAGWGRLIQLRENKHKD 133
>AL375831
Length = 467
Score = 74.7 bits (182), Expect(2) = 5e-18
Identities = 39/102 (38%), Positives = 59/102 (57%), Gaps = 1/102 (0%)
Frame = +2
Query: 118 TMTYSAPVIHTIPQTEEPIFHSGNVEAYEEVSDLQEKY-DELRRDMKALRDKGKFGKTAY 176
T + P++HT+PQ ++ + + ++ E+ ELR +++A R KT
Sbjct: 56 TAAVTEPLVHTLPQGININTQHRSIPVTKTMEEMMEELAKELRHEIQANRGNADSVKTQ- 232
Query: 177 DLCLVPSVQVPHKFKIPDFEKYKGSSCPEEHLKMYVRRMPAY 218
DLCLV V VP KFKIPDF++Y G +CP+ H+ YVR+M Y
Sbjct: 233 DLCLVSKVDVPKKFKIPDFDRYNGLTCPQNHIIKYVRKMGNY 358
Score = 36.6 bits (83), Expect(2) = 5e-18
Identities = 14/32 (43%), Positives = 23/32 (71%)
Frame = +3
Query: 221 DDQILIYYFQESLTGPASKWYTNLDKTRVQTF 252
+D ++I+ FQ+SL A++WYT+L K + TF
Sbjct: 366 NDSLMIHCFQDSLMEDAAEWYTSLSKNDIHTF 461
>TC77595 weakly similar to PIR|T18350|T18350 probable pol polyprotein - rice
blast fungus gypsy retroelement (fragment), partial (14%)
Length = 1708
Score = 79.0 bits (193), Expect = 2e-14
Identities = 74/325 (22%), Positives = 136/325 (41%), Gaps = 13/325 (4%)
Frame = +2
Query: 2007 LIHEIHEGSFGIHPNGHTMAKKILRAGYYWMTMESDCYKHTRKCHKCQIYADKIHMPPTT 2066
L+ E H+ + HP G +I+ ++W + R C C IH+
Sbjct: 179 LVQESHDSTAAGHP-GRNGTLEIVSRKFFWPGQSQTVRRFVRNCDVC----GGIHIWRQA 343
Query: 2067 LNLLSSPWPF-----SMWGIDMIGRIEPKASNGHRFILVAIDYFTKWVEAASYANVTKQV 2121
P P S +D I + P G +++ V +D +K V + +
Sbjct: 344 KRGFLKPLPVPNRLHSDLSMDFITSLPPTRGRGSQYLWVIVDRLSKSVTLEEMDTMEAEA 523
Query: 2122 VVKFIKNHIICRY---GIPNRIITDNGTNLNNKMMKELCDDFKIEHHNSSPYRPQMNGAV 2178
+ + C Y G+P I++D G+N + +E C + S+ Y PQ +G
Sbjct: 524 CAQ---RFLSCHYRFHGMPQSIVSDRGSNWVGRFWREFCRLTGVTQLLSTSYHPQTDGGT 694
Query: 2179 EAANKNIKRIVQKMVVTYKD-WHEMLPFALHGYRTSVRTSIGATPFSLVYGMEAVLPVEV 2237
E N+ I+ +++ V +D W ++LP R +SIGATPF + +G V P+
Sbjct: 695 ERWNQEIQAVLRAYVCWSQDNWGDLLPTVQLALRNRHNSSIGATPFFVEHGYH-VDPIPT 871
Query: 2238 EIPSLRVLMEVDLSEAEWVQNRYDQLNLIEEKRMAALCHGQLYQKRMKQAFDKKVRPRE- 2296
+ V+ E + + V+ D I+ + +AA Q+R + + +K+ P +
Sbjct: 872 VEDTGGVVSEGEAAAQLLVKRMKDVTGFIQAEIVAA-------QQRSEASANKRRCPADR 1030
Query: 2297 FKEGDLVLKKIFSF---QPDSRGKW 2318
++ GD V + ++ +P + W
Sbjct: 1031 YQVGDKVWLNVSNYKSPRPSKKLDW 1105
>BG587127 weakly similar to PIR|H84506|H84 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(13%)
Length = 415
Score = 67.8 bits (164), Expect = 5e-11
Identities = 41/109 (37%), Positives = 56/109 (50%)
Frame = +3
Query: 1803 GAVLLTPKGTHIPFTARLRFDCTNNIAEYEACIMGIEEAIDLRIKNIEIYGDSALVINQI 1862
G L +P + + RL F +NN YEA I G+ A L+I+NI Y DS LV +Q
Sbjct: 3 GIRLTSPTNEILKQSFRLEFHASNNETNYEALIAGVRLAHGLKIRNIHAYCDSQLVASQF 182
Query: 1863 KGKWETLHAGLIPYRDYARRLLTFFNKVELHHIPRDENQMADALATLSS 1911
G++E + Y ++L + L IPR EN ADALA L+S
Sbjct: 183 SGEYEARDELMDTYLKLVQKLAQKLDYFALTRIPRSENVQADALAALAS 329
>AL375832
Length = 535
Score = 59.3 bits (142), Expect = 2e-08
Identities = 45/123 (36%), Positives = 64/123 (51%), Gaps = 7/123 (5%)
Frame = +3
Query: 1079 FIHAIIEDDPESVPR-GFITPGVSSHNWVAVDVPFVAHLSK*CICLVYAFLFKKKKNNIL 1137
F++AI E+ S R F+TPG + +W A+D+P + H+S+*C+ L+ F+F K N +
Sbjct: 3 FVNAISEEATGSGLRPAFVTPGGIASDWDAIDIPSIMHVSE*CVLLL*RFVF-KNNNPLA 179
Query: 1138 SPAPSESELI*GFLFQEIIINKIKTSFFSRCFVSFFLCFSEKW*-YKKPN-----MNFQK 1191
P + LF+ + FF CF F F EKW* KKP +NF K
Sbjct: 180 LPKARVIFSVRACLFRHCRLMNKNCRFFHDCFF-FSAFFLEKW*CQKKPKRLYFLINF-K 353
Query: 1192 NLH 1194
NLH
Sbjct: 354 NLH 362
>AL367832 weakly similar to GP|14091845|gb Putative retroelement {Oryza
sativa}, partial (2%)
Length = 384
Score = 53.5 bits (127), Expect = 9e-07
Identities = 39/108 (36%), Positives = 59/108 (54%)
Frame = -2
Query: 1250 KKRRPFSLMGTN*K*LTWAPKKTRKKSRLGHRSEQVLKSK**SSSKNMLMCLPGPTKICR 1309
KK +PFSL T P++T ++SR + LK + SSS+N + L KIC+
Sbjct: 383 KKGKPFSLTRKRLSLST*VPRRTSERSRSVLL*RKGLKGRSSSSSENTWIFLHARMKICQ 204
Query: 1310 V*TLILWYITYL*NLSVRRSNKS*EEHVLTWLSKSKKRYRNRSTLVSS 1357
V* L LW I +L+V S ++*E + WLS+ + R+++R V S
Sbjct: 203 V*ILKLWNIGSQQSLNVLPSGRN*EGLIQIWLSRLRVRFKSRLMRVFS 60
>BF647164 weakly similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana},
partial (6%)
Length = 469
Score = 49.7 bits (117), Expect = 1e-05
Identities = 33/109 (30%), Positives = 58/109 (52%), Gaps = 12/109 (11%)
Frame = +1
Query: 2262 QLNLIEEKRMAALCHGQLYQKRMKQAFDKKVRPREFKEGDLVL-----KKIFSFQPDSRG 2316
+LN +EE R+ A + ++Y++R K+ D+++ REF+EG+LVL K+F G
Sbjct: 103 ELNELEELRLDAYENAKIYKERTKKWHDRRIIRREFREGELVLLFNSRLKLFP------G 264
Query: 2317 KWAPNYEGPYVVKKAFSGGAMTLQ-------TMDGEELPRPVNTDTVKK 2358
K ++ GP+ VK GA+ + T++G+ L + + V K
Sbjct: 265 KLRSHWSGPFQVKNVMPSGAVEVWSESTGPFTVNGQRLKHYTSGENVDK 411
>BG586326 similar to PIR|G84493|G8 probable retroelement pol polyprotein
[imported] - Arabidopsis thaliana, partial (13%)
Length = 736
Score = 47.8 bits (112), Expect = 5e-05
Identities = 31/115 (26%), Positives = 55/115 (46%)
Frame = +2
Query: 1636 MGCVLGQQDETGRKEHAIYYLSKKFTECESRYSMLEKTCCALAWAAKRLRHYMINHTTWL 1695
+GCVL Q E I Y S++ + E Y + A+ +A K R Y+ +
Sbjct: 80 LGCVLTQH------EKVIAYASRQLRKHEGNYPTHDLEMAAVVFALKIWRSYLYGAKVQI 241
Query: 1696 ISKMDPIKYIFEKPALIGRIARWQMLLSEYDIEYRSQKAIKGSILADHLAHQPLE 1750
+ +KYIF +P L R RW +++YD++ K +++AD L+ + ++
Sbjct: 242 HTDHKSLKYIFTQPELNLRQRRWMEFVADYDLDITYYPG-KANLVADALSRRRVD 403
>TC87407 similar to PIR|T49945|T49945 periaxin-like protein - Arabidopsis
thaliana, partial (44%)
Length = 1559
Score = 47.4 bits (111), Expect = 7e-05
Identities = 22/98 (22%), Positives = 45/98 (45%)
Frame = +3
Query: 417 QYPQIPQYPQIPQYPQFPQNPSPQNTQQQNFQQQPYQQFPYQQYPQQNFQQQPYQQRPQQ 476
++P++P+ P++P+ P+FP+ P P+ + + P P+ P + +Q
Sbjct: 366 EFPKLPELPKVPELPKFPELPKPELPKVPELSMPEIPKIPELPKPELPKLNAPELPKLEQ 545
Query: 477 PRPPRMPINPIPVTYAELLPGLLKKNLVQTRTAPPIPE 514
P+ P +P + +P P + K + P +PE
Sbjct: 546 PKVPELPKHELPKVSELPKPDIPKVPELPKPELPKVPE 659
>CA860311 weakly similar to GP|7289872|gb|A CG17427 gene product {Drosophila
melanogaster}, partial (20%)
Length = 192
Score = 46.2 bits (108), Expect = 1e-04
Identities = 21/53 (39%), Positives = 30/53 (55%)
Frame = +1
Query: 1636 MGCVLGQQDETGRKEHAIYYLSKKFTECESRYSMLEKTCCALAWAAKRLRHYM 1688
+G L Q+DE EH I Y S+ T E Y+++E+ C A WA + RHY+
Sbjct: 16 IGAGLSQKDEENH-EHPIAYASRLLTAAERNYTVVERECLAAIWAIRNFRHYL 171
>TC83629 weakly similar to GP|6692261|gb|AAF24611.1| putative RNase H
{Arabidopsis thaliana}, partial (21%)
Length = 1071
Score = 45.4 bits (106), Expect = 3e-04
Identities = 32/100 (32%), Positives = 45/100 (45%)
Frame = +2
Query: 1801 GIGAVLLTPKGTHIPFTARLRFDCTNNIAEYEACIMGIEEAIDLRIKNIEIYGDSALVIN 1860
G GA+LL G+ + + T AEY A ++G++ A K + GDS LVIN
Sbjct: 602 GAGALLLAEDGSLLYGFRQGLGHQTKESAEYRALLLGLKHASMKGFKYVTAKGDSELVIN 781
Query: 1861 QIKGKWETLHAGLIPYRDYARRLLTFFNKVELHHIPRDEN 1900
QI W+ L A L F+ + HI R+ N
Sbjct: 782 QILDPWKIKDEHLKKLCAEALELSDNFHSFRIQHISRERN 901
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.327 0.140 0.434
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 74,114,229
Number of Sequences: 36976
Number of extensions: 1139846
Number of successful extensions: 10969
Number of sequences better than 10.0: 141
Number of HSP's better than 10.0 without gapping: 6568
Number of HSP's successfully gapped in prelim test: 313
Number of HSP's that attempted gapping in prelim test: 1799
Number of HSP's gapped (non-prelim): 9031
length of query: 2360
length of database: 9,014,727
effective HSP length: 112
effective length of query: 2248
effective length of database: 4,873,415
effective search space: 10955436920
effective search space used: 10955436920
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 66 (30.0 bits)
Medicago: description of AC146971.7


