

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146941.2 - phase: 0 /pseudo
(240 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
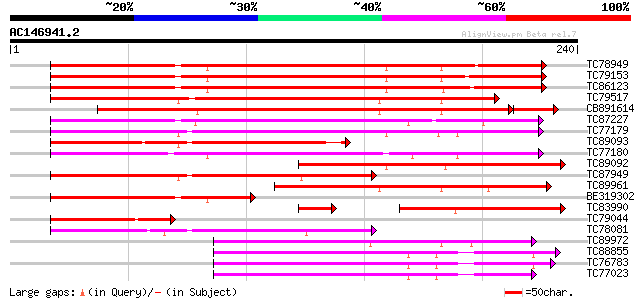
Score E
Sequences producing significant alignments: (bits) Value
TC78949 similar to GP|16215471|emb|CAC82999. calcium-dependent p... 196 5e-51
TC79153 similar to SP|P28583|CDPK_SOYBN Calcium-dependent protei... 186 5e-48
TC86123 homologue to PIR|S71770|S71770 calcium-dependent protein... 186 5e-48
TC79517 similar to PIR|T08874|T08874 calcium-dependent protein k... 175 2e-44
CB891614 similar to GP|13561063|em protein kinase {Medicago sati... 159 7e-42
TC87227 similar to GP|17064926|gb|AAL32617.1 calcium-dependent p... 147 5e-36
TC77179 similar to GP|2665890|gb|AAB88537.1| calcium-dependent p... 142 1e-34
TC89093 similar to GP|15011837|gb|AAK26164.2 calcium-dependent c... 134 2e-32
TC77180 similar to GP|2665890|gb|AAB88537.1| calcium-dependent p... 132 9e-32
TC89092 similar to GP|15011837|gb|AAK26164.2 calcium-dependent c... 119 8e-28
TC87949 similar to GP|16904224|gb|AAL30819.1 calcium/calmodulin-... 108 2e-24
TC89961 GP|13561063|emb|CAA65500. protein kinase {Medicago sativ... 106 9e-24
BE319302 homologue to SP|P28583|CDPK Calcium-dependent protein k... 95 3e-20
TC83990 similar to GP|15011837|gb|AAK26164.2 calcium-dependent c... 71 7e-15
TC79044 homologue to GP|13561063|emb|CAA65500. protein kinase {M... 75 3e-14
TC78081 similar to GP|16904226|gb|AAL30820.1 calcium/calmodulin-... 72 1e-13
TC89972 similar to GP|22296346|dbj|BAC10116. putative caltractin... 72 2e-13
TC88855 PIR|A30900|MCBH calmodulin - barley, complete 70 6e-13
TC76783 calmodulin 1 [Medicago truncatula] 70 7e-13
TC77023 calmodulin 2 [Medicago truncatula] 70 9e-13
>TC78949 similar to GP|16215471|emb|CAC82999. calcium-dependent protein
kinase 3 {Nicotiana tabacum}, partial (44%)
Length = 1015
Score = 196 bits (499), Expect = 5e-51
Identities = 116/220 (52%), Positives = 147/220 (66%), Gaps = 10/220 (4%)
Frame = +1
Query: 18 WPSISPSAKDLVRKMLNSDPKQRISAYEVLNHPWIKEDGEAPDTPLDNAVLNRNSLEQ*T 77
WP+IS SAKDLVRKML DPK+R++A++VL HPWI+ DG APD PLD+AVL+R L+Q +
Sbjct: 58 WPAISESAKDLVRKMLVRDPKRRMTAHQVLCHPWIQVDGVAPDKPLDSAVLSR--LKQFS 231
Query: 78 DSRKL-------L*SCLSEEEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQ 130
KL + LSEEE+ GLK+MFK +DTDNSG IT EELK GL K G L E +
Sbjct: 232 AMNKLKKMALIVIAESLSEEELAGLKEMFKMIDTDNSGQITFEELKVGLKKVGANLKESE 411
Query: 131 VKQLMEAADADGNGIIDYDEFITASQQL--CTRTD*TEKNMFILPSNTSIKIT-TELEQA 187
+ LM+AAD D +G IDY EFI A+ + R D + S IT EL+QA
Sbjct: 412 IYDLMQAADVDNSGTIDYGEFIAATLHINKIEREDHLFAAFSYFDKDGSGYITQEELQQA 591
Query: 188 LHEYNMHDGRYIKEIISEVDADNDGRINYDEFVAMMRKGN 227
E+ + D R ++EII E+D DNDGRI+Y+EF AMM+KGN
Sbjct: 592 CDEFGIKDVR-LEEIIKEIDEDNDGRIDYNEFAAMMQKGN 708
>TC79153 similar to SP|P28583|CDPK_SOYBN Calcium-dependent protein kinase SK5
(EC 2.7.1.-) (CDPK). [Soybean] {Glycine max}, partial
(95%)
Length = 1751
Score = 186 bits (473), Expect = 5e-48
Identities = 108/220 (49%), Positives = 144/220 (65%), Gaps = 10/220 (4%)
Frame = +3
Query: 18 WPSISPSAKDLVRKMLNSDPKQRISAYEVLNHPWIKEDGEAPDTPLDNAVLNRNSLEQ*T 77
WP IS SAKDL+RKML+ +PK R +A++VL HPWI +D APD PLD+AVL+R L+Q +
Sbjct: 822 WPGISDSAKDLIRKMLDRNPKTRFTAHQVLCHPWIVDDNIAPDKPLDSAVLSR--LKQFS 995
Query: 78 DSRKL-------L*SCLSEEEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQ 130
KL + LSEEEI GLK++F+ +D DNSGTIT+EELK GL + G+ L E +
Sbjct: 996 AMNKLKKMALRVIAERLSEEEIGGLKELFRMLDADNSGTITLEELKEGLKRVGSELMESE 1175
Query: 131 VKQLMEAADADGNGIIDYDEFITASQQL--CTRTD*TEKNMFILPSNTSIKIT-TELEQA 187
+K LM+AAD D NG +DY EFI A+ L R + + S IT E++ A
Sbjct: 1176 IKDLMDAADIDNNGTLDYGEFIAATVHLNKLEREENLLSAFSYFDKDCSGYITIDEIQAA 1355
Query: 188 LHEYNMHDGRYIKEIISEVDADNDGRINYDEFVAMMRKGN 227
E+ + D +I E++ E+D DNDG+I+Y EF AMMRKGN
Sbjct: 1356 CKEFGL-DDVHIDEMVKEIDQDNDGQIDYGEFAAMMRKGN 1472
>TC86123 homologue to PIR|S71770|S71770 calcium-dependent protein kinase (EC
2.7.1.-) - mung bean, complete
Length = 2448
Score = 186 bits (473), Expect = 5e-48
Identities = 113/220 (51%), Positives = 144/220 (65%), Gaps = 10/220 (4%)
Frame = +3
Query: 18 WPSISPSAKDLVRKMLNSDPKQRISAYEVLNHPWIKEDGEAPDTPLDNAVLNRNSLEQ*T 77
WP IS S KDL+RKML S P R++A+EVL HPWI E+G APD LD AVL+R L+Q +
Sbjct: 1206 WPLISDSGKDLIRKMLCSRPSDRLTAHEVLCHPWICENGVAPDRSLDPAVLSR--LKQFS 1379
Query: 78 DSRKL-------L*SCLSEEEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQ 130
KL + LSEEEI GL++MF+ MDTDNSG IT +ELK GL + G+ L + +
Sbjct: 1380 AMNKLKKMALRVIAESLSEEEIAGLREMFQTMDTDNSGAITFDELKAGLRRYGSTLKDIE 1559
Query: 131 VKQLMEAADADGNGIIDYDEFITASQQL--CTRTD*TEKNMFILPSNTSIKITT-ELEQA 187
++ LMEAAD D +G IDY EFI A+ L R + + S IT EL+QA
Sbjct: 1560 IRDLMEAADVDNSGTIDYGEFIAATVHLNKLEREEHLVAAFQYFDKDGSGYITVDELQQA 1739
Query: 188 LHEYNMHDGRYIKEIISEVDADNDGRINYDEFVAMMRKGN 227
E+NM D ++++II EVD DNDGRI+Y EFVAMM+KGN
Sbjct: 1740 CTEHNMTD-VFLEDIIKEVDQDNDGRIDYGEFVAMMQKGN 1856
>TC79517 similar to PIR|T08874|T08874 calcium-dependent protein kinase (EC
2.7.1.-) gamma - soybean, partial (80%)
Length = 1930
Score = 175 bits (443), Expect = 2e-44
Identities = 105/200 (52%), Positives = 133/200 (66%), Gaps = 10/200 (5%)
Frame = +1
Query: 18 WPSISPSAKDLVRKMLNSDPKQRISAYEVLNHPWIKEDGEAPDTPLDNAVLNR------- 70
WPSIS SAKDLV +ML DPK+RI+A +VL+HPW+KE G A D P+D+AVL+R
Sbjct: 1306 WPSISNSAKDLVSRMLMQDPKKRITASQVLDHPWLKEGGNASDKPIDSAVLSRMKQFRAM 1485
Query: 71 NSLEQ*TDSRKLL*SCLSEEEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQ 130
N L++ + K++ LS EEI GLK MF MDTD SGTIT EEL+ GL + G++L+E +
Sbjct: 1486 NKLKKL--ALKVIAENLSSEEIQGLKAMFTNMDTDKSGTITYEELRAGLQRLGSKLTEAE 1659
Query: 131 VKQLMEAADADGNGIIDYDEFITAS--QQLCTRTD*TEKNMFILPSNTSIKIT-TELEQA 187
V+QLMEAAD DGNG IDY EFITA+ + R + K + S IT ELE A
Sbjct: 1660 VRQLMEAADVDGNGTIDYIEFITATMHRHRLERDEHLYKAFQYFDKDNSGFITRDELETA 1839
Query: 188 LHEYNMHDGRYIKEIISEVD 207
+ EY M D I+EIISEV+
Sbjct: 1840 MKEYGMGDEETIREIISEVE 1899
>CB891614 similar to GP|13561063|em protein kinase {Medicago sativa}, partial
(37%)
Length = 637
Score = 159 bits (403), Expect(2) = 7e-42
Identities = 97/184 (52%), Positives = 122/184 (65%), Gaps = 8/184 (4%)
Frame = +3
Query: 38 KQRISAYEVLNHPWIKEDGEAPDTPLDNAVLNRNSLEQ*TD-----SRKLL*SCLSEEEI 92
K+RI++ EVL HPW++E GEA D P+D+AVL+R + + + K++ LSEEEI
Sbjct: 3 KKRITSTEVLEHPWMREGGEASDKPIDSAVLSRMKQFRAMNKFKKLALKVMAENLSEEEI 182
Query: 93 MGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQVKQLMEAADADGNGIIDYDEFI 152
GLK MF MDTD+SGTIT EELK GL + G+RLSE +VKQLMEAAD DGNG IDY EFI
Sbjct: 183 KGLKAMFANMDTDSSGTITYEELKTGLARIGSRLSEAEVKQLMEAADVDGNGSIDYLEFI 362
Query: 153 TAS--QQLCTRTD*TEKNMFILPSNTSIKIT-TELEQALHEYNMHDGRYIKEIISEVDAD 209
+A+ + R + K + S IT ELE A+ ++ M D IKEIISEVD D
Sbjct: 363 SATMHRHRLERDEHLYKAFQYFDKDNSGHITREELETAMTKHGMGDEATIKEIISEVDTD 542
Query: 210 NDGR 213
NDGR
Sbjct: 543 NDGR 554
Score = 28.1 bits (61), Expect(2) = 7e-42
Identities = 12/19 (63%), Positives = 13/19 (68%)
Frame = +2
Query: 214 INYDEFVAMMRKGNPEAHT 232
INY+EF AMMR G P T
Sbjct: 554 INYEEFCAMMRSGMPHQGT 610
>TC87227 similar to GP|17064926|gb|AAL32617.1 calcium-dependent protein kinase
{Arabidopsis thaliana}, partial (95%)
Length = 2319
Score = 147 bits (370), Expect = 5e-36
Identities = 90/221 (40%), Positives = 129/221 (57%), Gaps = 12/221 (5%)
Frame = +1
Query: 18 WPSISPSAKDLVRKMLNSDPKQRISAYEVLNHPWIKEDGEAPDTPLDNAVLNRNSLEQ*T 77
WPSIS SAK LV++ML DPK R++A +VL HPW++ +AP+ PL + V +R L+Q +
Sbjct: 1042 WPSISESAKSLVKQMLEPDPKLRLTAKQVLEHPWLQNAKKAPNVPLGDVVKSR--LKQFS 1215
Query: 78 -------DSRKLL*SCLSEEEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQ 130
+ +++ LS EE+ +K+MFK MDTDN G ++IEELK G ++L+E +
Sbjct: 1216 MMNRFKRKALRVIADFLSNEEVEDIKEMFKKMDTDNDGIVSIEELKVGFRNHQSQLAESE 1395
Query: 131 VKQLMEAADADGNGIIDYDEFITASQQLCTRTD*TEK----NMFILPSNTSIKITTELEQ 186
V+ +EA D +G G +DY EF+ S L + + F N I+ EL
Sbjct: 1396 VQMFIEAVDNNGKGTLDYGEFVAISLHLKRMANDEHLHKAFSYFDKDGNGYIE-PEELRN 1572
Query: 187 ALHEYNMHDGRYI-KEIISEVDADNDGRINYDEFVAMMRKG 226
AL E D + +I EVD D DG+I+YDEFVAMM+ G
Sbjct: 1573 ALMEDGTDDCTDVANDIFQEVDTDKDGKISYDEFVAMMKTG 1695
>TC77179 similar to GP|2665890|gb|AAB88537.1| calcium-dependent protein kinase
{Fragaria x ananassa}, partial (92%)
Length = 2243
Score = 142 bits (358), Expect = 1e-34
Identities = 84/220 (38%), Positives = 129/220 (58%), Gaps = 11/220 (5%)
Frame = +1
Query: 18 WPSISPSAKDLVRKMLNSDPKQRISAYEVLNHPWIKEDGEAPDTPLDNAVLNR------- 70
WP +S +AKDL++KML+ DPK+R++A EVL+HPW++ AP+ L V R
Sbjct: 1066 WPKVSDNAKDLIKKMLDPDPKRRLTAQEVLDHPWLQNAKTAPNVSLGETVRARLMQFSVM 1245
Query: 71 NSLEQ*TDSRKLL*SCLSEEEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQ 130
N L++ + +++ LS EE+ G+K+ F+ MDT+N G I ++EL+ GL+K G ++ E
Sbjct: 1246 NKLKK--TALRIIADHLSVEEVAGIKEGFQVMDTENKGKINLDELRVGLLKLGHQIPEGD 1419
Query: 131 VKQLMEAADADGNGIIDYDEFITASQQL--CTRTD*TEKNMFILPSNTSIKI-TTELEQA 187
V+ LMEA D D +G +DY EF+ S L + + ++ N S I EL A
Sbjct: 1420 VQILMEAGDVDKDGFLDYGEFVAISIHLRKISHDEHLQRAFQFFDKNESGFIELEELRNA 1599
Query: 188 L-HEYNMHDGRYIKEIISEVDADNDGRINYDEFVAMMRKG 226
L E + + I I+ +VD D DG+I+Y+EF MM+ G
Sbjct: 1600 LADEVDTNSEEVINAIMHDVDTDKDGKISYEEFATMMKAG 1719
>TC89093 similar to GP|15011837|gb|AAK26164.2 calcium-dependent
calmodulin-independent protein kinase 5 {Cucumis
sativus}, partial (42%)
Length = 690
Score = 134 bits (338), Expect = 2e-32
Identities = 77/134 (57%), Positives = 96/134 (71%), Gaps = 7/134 (5%)
Frame = +1
Query: 18 WPSISPSAKDLVRKMLNSDPKQRISAYEVLNHPWIKEDGEAPDTPLDNAVLNR------- 70
WP IS AKDL++KML +DPK+RISA EVL+H W+KEDG A D PLD AVL R
Sbjct: 295 WPKISSIAKDLIKKMLRADPKERISAVEVLDHSWMKEDG-ASDKPLDIAVLTRMKQFRAM 471
Query: 71 NSLEQ*TDSRKLL*SCLSEEEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQ 130
N L++ + K++ LSEEEI+GLK+MFK MDTDNSGTIT EELK GL K GT++SE +
Sbjct: 472 NKLKK--VALKVIAENLSEEEIIGLKEMFKSMDTDNSGTITFEELKAGLPKLGTKISESE 645
Query: 131 VKQLMEAADADGNG 144
V+Q+ DG+G
Sbjct: 646 VRQI------DGSG 669
>TC77180 similar to GP|2665890|gb|AAB88537.1| calcium-dependent protein
kinase {Fragaria x ananassa}, partial (41%)
Length = 681
Score = 132 bits (333), Expect = 9e-32
Identities = 86/222 (38%), Positives = 123/222 (54%), Gaps = 13/222 (5%)
Frame = +1
Query: 18 WPSISPSAKDLVRKMLNSDPKQRISAYEVLNHPWIKEDGEAPDTPLDNAVLNRNSLEQ*T 77
WP +S +AKDLV+ MLN DPK+R++A EVL+HPW+ +AP+ L V R L+Q +
Sbjct: 4 WPKVSDNAKDLVKNMLNPDPKRRLTAQEVLDHPWLINAKKAPNVSLGETV--RARLKQFS 177
Query: 78 DSRKL-------L*SCLSEEEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQ 130
KL + LS EE GLK+ F MDT N G I I+EL+ GL K G ++ +
Sbjct: 178 VMNKLKKRALRVIAEHLSVEEAAGLKEGFNLMDTTNRGKINIDELRTGLHKLGHQVPDSD 357
Query: 131 VKQLMEAADADGNGIIDYDEFITASQQLCTRTD*TEKNM-----FILPSNTSIKITTELE 185
++ LMEA D D +G +DY E++ S L R ++++ F + T EL
Sbjct: 358 LQILMEAGDIDRDGYLDYGEYVAISVHL--RKMGNDEHLHKAFDFFDQNQTGYIEIEELR 531
Query: 186 QAL-HEYNMHDGRYIKEIISEVDADNDGRINYDEFVAMMRKG 226
AL E + I I+ +VD D DG+I+Y+EF MM+ G
Sbjct: 532 NALSDEIETNSEEVISAIMHDVDTDKDGKISYEEFANMMKAG 657
>TC89092 similar to GP|15011837|gb|AAK26164.2 calcium-dependent
calmodulin-independent protein kinase 5 {Cucumis
sativus}, partial (22%)
Length = 632
Score = 119 bits (299), Expect = 8e-28
Identities = 68/116 (58%), Positives = 80/116 (68%), Gaps = 3/116 (2%)
Frame = +3
Query: 123 GTRLSEQQVKQLMEAADADGNGIIDYDEFITASQQL--CTRTD*TEKNMFILPSNTSIKI 180
G ++SE +V+QLMEAAD DGNG IDY FITA+ L R D K ++ S I
Sbjct: 3 GIKISESEVRQLMEAADVDGNGTIDYINFITATMHLNRMEREDHLYKAFEYFDNDKSGYI 182
Query: 181 TTE-LEQALHEYNMHDGRYIKEIISEVDADNDGRINYDEFVAMMRKGNPEAHTKKR 235
T E LE AL +YNM D + IKEII EVD+DNDGRINY+EFVAMMRKGNP+ T KR
Sbjct: 183 TKEELESALTKYNMGDEKTIKEIIDEVDSDNDGRINYEEFVAMMRKGNPDLITNKR 350
>TC87949 similar to GP|16904224|gb|AAL30819.1 calcium/calmodulin-dependent
protein kinase CaMK2 {Nicotiana tabacum}, partial (62%)
Length = 1413
Score = 108 bits (269), Expect = 2e-24
Identities = 55/146 (37%), Positives = 95/146 (64%), Gaps = 8/146 (5%)
Frame = +1
Query: 18 WPSISPSAKDLVRKMLNSDPKQRISAYEVLNHPWIKEDGEAPDTPLDNAVLNR------- 70
WP+IS +AKD V+K+L DP+ R++A + L+HPW++E GEA + P+D +VLN
Sbjct: 523 WPTISNAAKDFVKKLLVKDPRARLTAAQALSHPWVREGGEASEIPIDISVLNNMRQFVKY 702
Query: 71 NSLEQ*TDSRKLL*SCLSEEEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQ-GTRLSEQ 129
+ L+Q + + L S L+E E+ LK F +D D +G I++EE++ L K +L E
Sbjct: 703 SRLKQ--FALRALASTLNEGELSDLKDQFDAIDVDKNGAISLEEMRQALAKDLPWKLKES 876
Query: 130 QVKQLMEAADADGNGIIDYDEFITAS 155
+V ++++A D++ +G++D+ EF+ A+
Sbjct: 877 RVLEILQAIDSNTDGLVDFTEFVAAT 954
>TC89961 GP|13561063|emb|CAA65500. protein kinase {Medicago sativa}, partial
(24%)
Length = 719
Score = 106 bits (264), Expect = 9e-24
Identities = 68/127 (53%), Positives = 80/127 (62%), Gaps = 10/127 (7%)
Frame = +3
Query: 113 EELK*GLVKQGTRLSEQQVKQLMEAADADGNGIIDYDEFITAS--QQLCTRTD*TEKNMF 170
EELK GL + G++LSE +VKQLMEAAD DGNG ID EFITA+ + R D K
Sbjct: 6 EELKAGLQRLGSKLSEAEVKQLMEAADVDGNGTIDCIEFITATMHRHKLERDDHLYKAFQ 185
Query: 171 ILPSNTSIKIT-TELEQALHEYNMHDGRYIKE-------IISEVDADNDGRINYDEFVAM 222
++S IT ELE A+ EY M D IKE IISEVD D+DGRINY+EF AM
Sbjct: 186 YFDKDSSGFITRDELETAMKEYGMGDDATIKEIISEVDTIISEVDTDHDGRINYEEFCAM 365
Query: 223 MRKGNPE 229
MR GN +
Sbjct: 366 MRSGNQQ 386
>BE319302 homologue to SP|P28583|CDPK Calcium-dependent protein kinase SK5
(EC 2.7.1.-) (CDPK). [Soybean] {Glycine max}, partial
(35%)
Length = 546
Score = 94.7 bits (234), Expect = 3e-20
Identities = 51/94 (54%), Positives = 68/94 (72%), Gaps = 7/94 (7%)
Frame = +3
Query: 18 WPSISPSAKDLVRKMLNSDPKQRISAYEVLNHPWIKEDGEAPDTPLDNAVLNRNSLEQ*T 77
WPSIS SAKDL+RKML+ +P+ R++A+EVL HPWI +D APD P+D+AVL+R L+Q +
Sbjct: 162 WPSISDSAKDLIRKMLDQNPRTRLTAHEVLRHPWIVDDNIAPDKPIDSAVLSR--LKQFS 335
Query: 78 DSRKL-------L*SCLSEEEIMGLKQMFKGMDT 104
KL + LSEEEI GLK++FK +DT
Sbjct: 336 AMNKLKKMALRVIAERLSEEEIGGLKELFKMIDT 437
>TC83990 similar to GP|15011837|gb|AAK26164.2 calcium-dependent
calmodulin-independent protein kinase 5 {Cucumis
sativus}, partial (21%)
Length = 679
Score = 71.2 bits (173), Expect(2) = 7e-15
Identities = 39/76 (51%), Positives = 47/76 (61%), Gaps = 6/76 (7%)
Frame = +3
Query: 166 EKNMFILPSNTSIKITTELEQA------LHEYNMHDGRYIKEIISEVDADNDGRINYDEF 219
E+ +I P + KI Q L +YNM D IKEII+EVD DND RINYDEF
Sbjct: 126 ERITYIKPLSILTKIRAGTSQRKSWNLPLRKYNMGDENTIKEIIAEVDTDNDARINYDEF 305
Query: 220 VAMMRKGNPEAHTKKR 235
VAMMRKGNP+ ++R
Sbjct: 306 VAMMRKGNPDITHRRR 353
Score = 32.3 bits (72), Expect = 0.17
Identities = 24/53 (45%), Positives = 27/53 (50%), Gaps = 3/53 (5%)
Frame = +2
Query: 139 DADGNGIIDYDEFITASQQL--CTRTD*TEKNMFILPSNTSIKIT-TELEQAL 188
D DGNG IDY EFITA+ + R D K + S IT ELE AL
Sbjct: 53 DVDGNGTIDYIEFITATMHMNRMEREDHLYKAFEYFDQDKSGYITKEELESAL 211
Score = 26.6 bits (57), Expect = 9.2
Identities = 9/25 (36%), Positives = 16/25 (64%)
Frame = +3
Query: 128 EQQVKQLMEAADADGNGIIDYDEFI 152
E +K+++ D D + I+YDEF+
Sbjct: 234 ENTIKEIIAEVDTDNDARINYDEFV 308
Score = 25.8 bits (55), Expect(2) = 7e-15
Identities = 10/16 (62%), Positives = 15/16 (93%)
Frame = +1
Query: 123 GTRLSEQQVKQLMEAA 138
G+++SE +V+QLMEAA
Sbjct: 4 GSKMSESEVRQLMEAA 51
>TC79044 homologue to GP|13561063|emb|CAA65500. protein kinase {Medicago
sativa}, partial (67%)
Length = 1408
Score = 74.7 bits (182), Expect = 3e-14
Identities = 36/53 (67%), Positives = 43/53 (80%)
Frame = +3
Query: 18 WPSISPSAKDLVRKMLNSDPKQRISAYEVLNHPWIKEDGEAPDTPLDNAVLNR 70
WP IS SAKDLVRKML +PK+RI+A +VL HPWIK DG A D P+D+AVL+R
Sbjct: 1233 WPKISDSAKDLVRKMLIQEPKKRITAAQVLEHPWIK-DGNASDKPIDSAVLSR 1388
>TC78081 similar to GP|16904226|gb|AAL30820.1 calcium/calmodulin-dependent
protein kinase CaMK3 {Nicotiana tabacum}, partial (58%)
Length = 1066
Score = 72.4 bits (176), Expect = 1e-13
Identities = 42/146 (28%), Positives = 79/146 (53%), Gaps = 8/146 (5%)
Frame = +2
Query: 18 WPSISPSAKDLVRKMLNSDPKQRISAYEVLNHPWIKEDGEAPDTPLD-------NAVLNR 70
WPS+S AKD V+++LN DP++RISA + L+HPWI+ + PLD +
Sbjct: 614 WPSLSSEAKDFVKRLLNKDPRKRISAAQALSHPWIRNYNDV-KVPLDILIFKLMKTYMRS 790
Query: 71 NSLEQ*TDSRKLL*SCLSEEEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGT-RLSEQ 129
+SL + + + L L+ +E+ L++ F ++ +G+I++E + L K T + E
Sbjct: 791 SSLRK--AALRALSKTLTADELYYLREQFALLEPSKNGSISLENINKALKKHATDAMKES 964
Query: 130 QVKQLMEAADADGNGIIDYDEFITAS 155
++ + + + +D++EF A+
Sbjct: 965 RITDFLSSLSSLQYRRMDFEEFCAAA 1042
>TC89972 similar to GP|22296346|dbj|BAC10116. putative caltractin {Oryza
sativa (japonica cultivar-group)}, partial (33%)
Length = 801
Score = 72.0 bits (175), Expect = 2e-13
Identities = 44/142 (30%), Positives = 81/142 (56%), Gaps = 5/142 (3%)
Frame = +1
Query: 87 LSEEEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQVKQLMEAADADGNGII 146
L++++ +K+ F+ DTD SGTI +EL + G ++E+Q++Q++ D DG+G I
Sbjct: 115 LTQQKRQEIKEAFELFDTDGSGTIDAKELNVAMRALGFEMTEEQIEQMIADVDKDGSGAI 294
Query: 147 DYDEF---ITASQQLCTRTD*TEKNMFILPSNTSIKIT-TELEQALHEYNMH-DGRYIKE 201
DYDEF +TA + K I+ + + KI+ +++++ E + R I+E
Sbjct: 295 DYDEFEHMMTAKIGERDTKEELMKAFHIIDKDKNGKISASDIKRIAKELGQNFTDREIQE 474
Query: 202 IISEVDADNDGRINYDEFVAMM 223
++ E D +ND ++ +EF+ MM
Sbjct: 475 MVDEADQNNDREVDPEEFIMMM 540
>TC88855 PIR|A30900|MCBH calmodulin - barley, complete
Length = 889
Score = 70.5 bits (171), Expect = 6e-13
Identities = 50/159 (31%), Positives = 76/159 (47%), Gaps = 12/159 (7%)
Frame = +1
Query: 87 LSEEEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQVKQLMEAADADGNGII 146
L++++I K+ F D D G IT +EL + G +E +++ ++ DADGNG I
Sbjct: 88 LTDDQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTI 267
Query: 147 DYDEFITASQQLCTRTD*TEK-----NMFILPSNTSIK------ITTELEQALHEYNMHD 195
D+ EF+ + TD E+ +F N I + T L + L +
Sbjct: 268 DFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTD----- 432
Query: 196 GRYIKEIISEVDADNDGRINYDEFV-AMMRKGNPEAHTK 233
+ E+I E D D DG+INY+EFV MM K P H +
Sbjct: 433 -EEVDEMIREADVDGDGQINYEEFVKVMMAK*GPINHNR 546
>TC76783 calmodulin 1 [Medicago truncatula]
Length = 957
Score = 70.1 bits (170), Expect = 7e-13
Identities = 50/157 (31%), Positives = 76/157 (47%), Gaps = 12/157 (7%)
Frame = +3
Query: 87 LSEEEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQVKQLMEAADADGNGII 146
L++++I K+ F D D G IT +EL + G +E +++ ++ DADGNG I
Sbjct: 132 LTDDQISEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTI 311
Query: 147 DYDEFITASQQLCTRTD*TEK-----NMFILPSNTSIK------ITTELEQALHEYNMHD 195
D+ EF+ + TD E+ +F N I + T L + L +
Sbjct: 312 DFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTD----- 476
Query: 196 GRYIKEIISEVDADNDGRINYDEFV-AMMRKGNPEAH 231
+ E+I E D D DG+INY+EFV MM K +P H
Sbjct: 477 -EEVDEMIREADVDGDGQINYEEFVKVMMAK*SPFFH 584
>TC77023 calmodulin 2 [Medicago truncatula]
Length = 999
Score = 69.7 bits (169), Expect = 9e-13
Identities = 47/148 (31%), Positives = 72/148 (47%), Gaps = 11/148 (7%)
Frame = +1
Query: 87 LSEEEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQVKQLMEAADADGNGII 146
L++E+I K+ F D D G IT +EL + G +E +++ ++ DADGNG I
Sbjct: 97 LTDEQISEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTI 276
Query: 147 DYDEFITASQQLCTRTD*TEK-----NMFILPSNTSIK------ITTELEQALHEYNMHD 195
D+ EF+ + TD E+ +F N I + T L + L +
Sbjct: 277 DFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTD----- 441
Query: 196 GRYIKEIISEVDADNDGRINYDEFVAMM 223
+ E+I E D D DG+INY+EFV +M
Sbjct: 442 -EEVDEMIREADVDGDGQINYEEFVKVM 522
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.323 0.137 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,501,192
Number of Sequences: 36976
Number of extensions: 57596
Number of successful extensions: 471
Number of sequences better than 10.0: 152
Number of HSP's better than 10.0 without gapping: 399
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 426
length of query: 240
length of database: 9,014,727
effective HSP length: 93
effective length of query: 147
effective length of database: 5,575,959
effective search space: 819665973
effective search space used: 819665973
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 57 (26.6 bits)
Medicago: description of AC146941.2


