

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146940.4 + phase: 1 /pseudo
(674 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
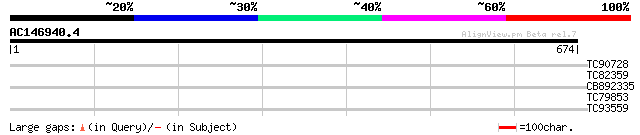
Score E
Sequences producing significant alignments: (bits) Value
TC90728 similar to GP|7715599|gb|AAF68117.1| F20B17.17 {Arabidop... 33 0.27
TC82359 weakly similar to GP|22655127|gb|AAM98154.1 putative pro... 32 0.79
CB892335 similar to GP|7715599|gb| F20B17.17 {Arabidopsis thalia... 30 3.9
TC79853 similar to GP|13605811|gb|AAK32891.1 AT5g36940/MLF18_60 ... 29 5.1
TC93559 homologue to PIR|F86426|F86426 hypothetical protein AAG3... 29 5.1
>TC90728 similar to GP|7715599|gb|AAF68117.1| F20B17.17 {Arabidopsis
thaliana}, partial (58%)
Length = 1171
Score = 33.5 bits (75), Expect = 0.27
Identities = 21/62 (33%), Positives = 34/62 (53%), Gaps = 7/62 (11%)
Frame = +3
Query: 569 KKRNRLEHQRLNDLVFVRYNLMLENRNNKIRNYDPINDELLD----DHHDNWVLED---S 621
+K+N+++ + LNDLV++ YNL L N+ + + +LL D WV E+ S
Sbjct: 720 EKKNKIDRETLNDLVYINYNLKL----NRQMSAKSLEVDLLQFDDIDMTSEWVEENETVS 887
Query: 622 PP 623
PP
Sbjct: 888 PP 893
>TC82359 weakly similar to GP|22655127|gb|AAM98154.1 putative protein
{Arabidopsis thaliana}, partial (14%)
Length = 907
Score = 32.0 bits (71), Expect = 0.79
Identities = 18/49 (36%), Positives = 27/49 (54%), Gaps = 2/49 (4%)
Frame = +3
Query: 571 RNRLEHQRLNDLVFVRYNLMLEN--RNNKIRNYDPINDELLDDHHDNWV 617
RN +E Q DLVFV YNL L N + + DP++ + + + D W+
Sbjct: 399 RNYIERQHFTDLVFVHYNLRLRQMFMNKEQESSDPLSFDNICNVED-WI 542
>CB892335 similar to GP|7715599|gb| F20B17.17 {Arabidopsis thaliana}, partial
(27%)
Length = 788
Score = 29.6 bits (65), Expect = 3.9
Identities = 20/66 (30%), Positives = 32/66 (48%)
Frame = +1
Query: 178 YWLYSYGRWVD*RG*ENSDKHFSLLP*RNDFYQIC*CFWCFKNW*DVV*AFQGSSVIYWP 237
YWL+ Y R +D* + SL+ ++ F Q+ C F+ +F + WP
Sbjct: 460 YWLHHYCRHMD*L*IKGHY*FLSLITIQDIFPQVSGCVCIFQEHKMACGSF*FCNSGVWP 639
Query: 238 *KCCSD 243
*KCC++
Sbjct: 640 *KCCTN 657
>TC79853 similar to GP|13605811|gb|AAK32891.1 AT5g36940/MLF18_60
{Arabidopsis thaliana}, partial (23%)
Length = 782
Score = 29.3 bits (64), Expect = 5.1
Identities = 15/33 (45%), Positives = 19/33 (57%), Gaps = 3/33 (9%)
Frame = +2
Query: 345 CT*SHGNISRMDNFCLCKRC---QGQTICGTSL 374
C+*SHGN R +N C C R G+ C TS+
Sbjct: 101 CS*SHGNWCRCNNRCWCIRSSWNSGKGTCWTSI 199
>TC93559 homologue to PIR|F86426|F86426 hypothetical protein AAG30958.1
[imported] - Arabidopsis thaliana, partial (1%)
Length = 894
Score = 29.3 bits (64), Expect = 5.1
Identities = 17/39 (43%), Positives = 22/39 (55%), Gaps = 1/39 (2%)
Frame = +1
Query: 96 KCVANQGSCRKV*SCTC-KMVHCCIYSLQCSKFTIFSVC 133
KCV+ +GSC C C + CCI CS F +FS+C
Sbjct: 451 KCVSEKGSC-----CNCHRFAICCI---ACS-FGLFSIC 540
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.366 0.162 0.645
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 26,920,937
Number of Sequences: 36976
Number of extensions: 480003
Number of successful extensions: 8194
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 2577
Number of HSP's successfully gapped in prelim test: 339
Number of HSP's that attempted gapping in prelim test: 5056
Number of HSP's gapped (non-prelim): 3555
length of query: 674
length of database: 9,014,727
effective HSP length: 103
effective length of query: 571
effective length of database: 5,206,199
effective search space: 2972739629
effective search space used: 2972739629
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.6 bits)
S2: 62 (28.5 bits)
Medicago: description of AC146940.4


