

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146866.2 + phase: 0
(504 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
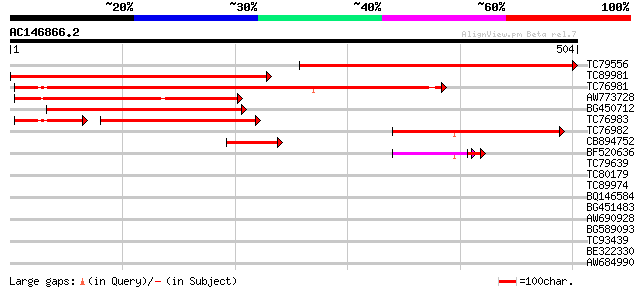
Score E
Sequences producing significant alignments: (bits) Value
TC79556 similar to GP|5882292|gb|AAD55269.1| sucrose transporter... 494 e-140
TC89981 similar to GP|9957218|gb|AAG09270.1| sucrose transporter... 468 e-132
TC76981 similar to PIR|T12198|T12198 sucrose transport protein -... 420 e-118
AW773728 similar to GP|10998390|gb sucrose transporter SUT4 {Sol... 290 7e-79
BG450712 similar to GP|17645762|em unnamed protein product {Glyc... 255 2e-68
TC76983 similar to PIR|T12198|T12198 sucrose transport protein -... 190 2e-62
TC76982 similar to PIR|T12198|T12198 sucrose transport protein -... 147 7e-36
CB894752 62 7e-10
BF520636 homologue to GP|5230818|gb|A sucrose transport protein ... 46 2e-05
TC79639 similar to GP|10728064|gb|AAF50455.2 CG7060 gene product... 32 0.75
TC80179 weakly similar to GP|20161298|dbj|BAB90223. hypothetical... 31 1.3
TC89974 similar to GP|7767653|gb|AAF69150.1| F27F5.2 {Arabidopsi... 31 1.3
BQ146584 similar to PIR|T06751|T0675 hypothetical protein F15B8.... 29 4.8
BG451483 weakly similar to PIR|PN0109|PN01 keratin-like protein ... 28 6.3
AW690928 similar to GP|22093675|d putative pre-mRNA splicing fac... 28 8.3
BG589093 similar to PIR|F71608|F716 hypothetical protein PFB0700... 28 8.3
TC93439 similar to GP|15081735|gb|AAK82522.1 AT3g06980/F17A9_13 ... 28 8.3
BE322330 28 8.3
AW684990 similar to PIR|F96703|F96 unknown protein 35653-33693 ... 28 8.3
>TC79556 similar to GP|5882292|gb|AAD55269.1| sucrose transporter {Vitis
vinifera}, partial (45%)
Length = 1209
Score = 494 bits (1272), Expect = e-140
Identities = 247/247 (100%), Positives = 247/247 (100%)
Frame = +3
Query: 258 SGAGESGSAEEAFMWELFGTFKYFSMPVWIVLSVTALTWIGWFPFNLFDTDWMGREIYGG 317
SGAGESGSAEEAFMWELFGTFKYFSMPVWIVLSVTALTWIGWFPFNLFDTDWMGREIYGG
Sbjct: 9 SGAGESGSAEEAFMWELFGTFKYFSMPVWIVLSVTALTWIGWFPFNLFDTDWMGREIYGG 188
Query: 318 DPEGGLIYDTGVRMGALGLLLNSVVLAVTSLLMERLCRKRGAGFVWGISNIFMAICFIAM 377
DPEGGLIYDTGVRMGALGLLLNSVVLAVTSLLMERLCRKRGAGFVWGISNIFMAICFIAM
Sbjct: 189 DPEGGLIYDTGVRMGALGLLLNSVVLAVTSLLMERLCRKRGAGFVWGISNIFMAICFIAM 368
Query: 378 LVLTYAANSIGYVSKGQPPPTGIVIAALAIFTILGFPMAITYSVPYALISTHIEPLGLGQ 437
LVLTYAANSIGYVSKGQPPPTGIVIAALAIFTILGFPMAITYSVPYALISTHIEPLGLGQ
Sbjct: 369 LVLTYAANSIGYVSKGQPPPTGIVIAALAIFTILGFPMAITYSVPYALISTHIEPLGLGQ 548
Query: 438 GLSMGVLNLAIVVPQIVVSLGSGPWDQLFGGGNSPAFAVAAVAALLSGLLALLAIPRTRT 497
GLSMGVLNLAIVVPQIVVSLGSGPWDQLFGGGNSPAFAVAAVAALLSGLLALLAIPRTRT
Sbjct: 549 GLSMGVLNLAIVVPQIVVSLGSGPWDQLFGGGNSPAFAVAAVAALLSGLLALLAIPRTRT 728
Query: 498 QKPRVRI 504
QKPRVRI
Sbjct: 729 QKPRVRI 749
>TC89981 similar to GP|9957218|gb|AAG09270.1| sucrose transporter
{Lycopersicon esculentum}, partial (39%)
Length = 717
Score = 468 bits (1204), Expect = e-132
Identities = 230/232 (99%), Positives = 230/232 (99%)
Frame = +3
Query: 1 MPNPTTTNPHRSRTRSSTSTSTSRPVQPVQPRTPLRQLLRVASVASGIQFGWALQLSLLT 60
MPNPTTTNPHRSRTRSSTSTSTSRPVQPVQPRTPLRQLLRVASVASGIQFGWALQLSLLT
Sbjct: 21 MPNPTTTNPHRSRTRSSTSTSTSRPVQPVQPRTPLRQLLRVASVASGIQFGWALQLSLLT 200
Query: 61 PYVQQLGIPHKWASIIWLCGPVSGLFVQPLVGHLSDRCSSRFGRRRPFILVGAASIVVAV 120
PYVQQLGIPHKWASIIWLCGPVSGLFVQPLVGHLSDRCSSRFGRRRPFILVGAASIVVAV
Sbjct: 201 PYVQQLGIPHKWASIIWLCGPVSGLFVQPLVGHLSDRCSSRFGRRRPFILVGAASIVVAV 380
Query: 121 VIIGYAADIGYLIGDDITQNYRPFAIVVFVIGFWILDVANNVTQGPCRALLADLTCNDAR 180
VIIGYAADIGYLIGDDITQNYRPFAIVVFVIGFWILDVANNVTQGPCRALLADLTCNDAR
Sbjct: 381 VIIGYAADIGYLIGDDITQNYRPFAIVVFVIGFWILDVANNVTQGPCRALLADLTCNDAR 560
Query: 181 RTRVANAYFSLFMAVGNILGYATGSYSGWYKIFTFTLTPACSISCANLKSAF 232
RTRVANAYFSLFMAVGNILGYATGSYSGWYKIFTFTLTPACSI CAN KSAF
Sbjct: 561 RTRVANAYFSLFMAVGNILGYATGSYSGWYKIFTFTLTPACSIXCANXKSAF 716
>TC76981 similar to PIR|T12198|T12198 sucrose transport protein - fava bean,
partial (71%)
Length = 1224
Score = 420 bits (1079), Expect = e-118
Identities = 211/394 (53%), Positives = 269/394 (67%), Gaps = 10/394 (2%)
Frame = +2
Query: 5 TTTNPHRSRTRSSTSTSTSRPVQPVQPRTPLRQLLRVASVASGIQFGWALQLSLLTPYVQ 64
T N + + T + S P P++P +PLR+++ VAS+A+G+QFGWALQLSLLTPYVQ
Sbjct: 56 TKQNHNNNNTLTKPSLHVESP--PLEP-SPLRKIIVVASIAAGVQFGWALQLSLLTPYVQ 226
Query: 65 QLGIPHKWASIIWLCGPVSGLFVQPLVGHLSDRCSSRFGRRRPFILVGAASIVVAVVIIG 124
LGIPH WA+ IWLCGP+SG+ VQP+VG+ SDRC+SRFGRRRPFI G+ ++ +AV +IG
Sbjct: 227 LLGIPHTWAAYIWLCGPISGMLVQPIVGYHSDRCTSRFGRRRPFIAAGSFAVAIAVFLIG 406
Query: 125 YAADIGYLIGDDITQNYRPFAIVVFVIGFWILDVANNVTQGPCRALLADLTCNDARRTRV 184
YAAD+G+ GDD+++ RP AI +FV+GFWILDVANN+ QGPCRALL DL + ++TR
Sbjct: 407 YAADLGHSFGDDLSKKVRPRAIGIFVVGFWILDVANNMLQGPCRALLGDLCAGNHQKTRN 586
Query: 185 ANAYFSLFMAVGNILGYATGSYSGWYKIFTFTLTPACSISCANLKSAFFLDVAFIVVTTY 244
ANA+FS FMAVGNILGYA G+YS + +F FT T AC I CANLKS FFL +A +
Sbjct: 587 ANAFFSFFMAVGNILGYAAGAYSKLFHVFPFTKTKACDIYCANLKSCFFLSIALLTAVAT 766
Query: 245 LSIVSAHEVPLSSSGAGESGSAEE---------AFMWELFGTFKYFSMPVWIVLSVTALT 295
+++ E+PLS +G +E EL G F+ P+WI+L VT L
Sbjct: 767 AALIYVKEIPLSPEKVTGNGVTDEDGNVTKSSNPCFGELSGAFRELKRPMWILLLVTCLN 946
Query: 296 WIGWFPFNLFDTDWMGREIYGGDPEGGLIYDTGVRMGALGLLLNSVVLAVTSLLMERLCR 355
WI WFPF LFDTDWMG+E+YGG G YD GVR GALGL+LNSVVL TSL ++ L R
Sbjct: 947 WIAWFPFLLFDTDWMGKEVYGGTVGEGHAYDKGVRAGALGLMLNSVVLGATSLXVDVLAR 1126
Query: 356 -KRGAGFVWGISNIFMAICFIAMLVLTYAANSIG 388
G +WGI N +AIC L +T A +G
Sbjct: 1127GVGGVKRLWGIVNFLLAIC----LAMTVXATKLG 1216
>AW773728 similar to GP|10998390|gb sucrose transporter SUT4 {Solanum
tuberosum}, partial (34%)
Length = 620
Score = 290 bits (743), Expect = 7e-79
Identities = 148/203 (72%), Positives = 168/203 (81%)
Frame = +2
Query: 5 TTTNPHRSRTRSSTSTSTSRPVQPVQPRTPLRQLLRVASVASGIQFGWALQLSLLTPYVQ 64
T + HRS+ R S S++ +P + R L +LLRVASVA GIQFGWALQLSLLTPYVQ
Sbjct: 5 THRHHHRSKLRPSPSSTVRIKPRP-KDRVLLTKLLRVASVAGGIQFGWALQLSLLTPYVQ 181
Query: 65 QLGIPHKWASIIWLCGPVSGLFVQPLVGHLSDRCSSRFGRRRPFILVGAASIVVAVVIIG 124
QLGIPH WASIIWLCGP+SGL VQPLVGHLSDRC+SRFGRRRPFIL GA SIV++V+IIG
Sbjct: 182 QLGIPHAWASIIWLCGPLSGLLVQPLVGHLSDRCTSRFGRRRPFILGGAVSIVISVLIIG 361
Query: 125 YAADIGYLIGDDITQNYRPFAIVVFVIGFWILDVANNVTQGPCRALLADLTCNDARRTRV 184
+AAD+G+ GD T+N+R A+ FV GFWILDVANNVTQGPCRALL DLT D R TRV
Sbjct: 362 HAADLGWKFGD--TKNHRHSAVAFFVFGFWILDVANNVTQGPCRALLGDLTGKDHRMTRV 535
Query: 185 ANAYFSLFMAVGNILGYATGSYS 207
ANAYFSLFMA+GNILG+ATGSYS
Sbjct: 536 ANAYFSLFMAIGNILGFATGSYS 604
>BG450712 similar to GP|17645762|em unnamed protein product {Glycine max},
partial (35%)
Length = 555
Score = 255 bits (652), Expect = 2e-68
Identities = 116/178 (65%), Positives = 142/178 (79%)
Frame = +2
Query: 33 TPLRQLLRVASVASGIQFGWALQLSLLTPYVQQLGIPHKWASIIWLCGPVSGLFVQPLVG 92
TPLR++ VAS+A+GIQFGWALQLSLLTPYVQ LG+PH+WA+ IWLCGP+SG+ +QPLVG
Sbjct: 5 TPLRKMAAVASIAAGIQFGWALQLSLLTPYVQLLGVPHQWAANIWLCGPISGMIIQPLVG 184
Query: 93 HLSDRCSSRFGRRRPFILVGAASIVVAVVIIGYAADIGYLIGDDITQNYRPFAIVVFVIG 152
+ SDR SRFGRRRPFI GA ++ +AV +IGYAAD+G+ +GDD+T+ RP A+V+FV G
Sbjct: 185 YYSDRSHSRFGRRRPFIFFGAIAVAIAVFLIGYAADLGHSMGDDLTKKTRPRAVVIFVFG 364
Query: 153 FWILDVANNVTQGPCRALLADLTCNDARRTRVANAYFSLFMAVGNILGYATGSYSGWY 210
FWILDVANN+ QGPCRA + DL D R R+ N FS FMA GN+LGYA GSY Y
Sbjct: 365 FWILDVANNMLQGPCRAFIGDLAGGDHRXMRIGNGMFSFFMAXGNVLGYAAGSYXKLY 538
>TC76983 similar to PIR|T12198|T12198 sucrose transport protein - fava bean,
partial (40%)
Length = 674
Score = 190 bits (482), Expect(2) = 2e-62
Identities = 88/143 (61%), Positives = 113/143 (78%)
Frame = +2
Query: 81 PVSGLFVQPLVGHLSDRCSSRFGRRRPFILVGAASIVVAVVIIGYAADIGYLIGDDITQN 140
P+SG+ VQP+VG+ SDRC+SRFGRRRPFI G+ ++ +AV +IGYAAD+G+ GDD+++
Sbjct: 227 PISGMLVQPIVGYHSDRCTSRFGRRRPFIAAGSFAVAIAVFLIGYAADLGHSFGDDLSKK 406
Query: 141 YRPFAIVVFVIGFWILDVANNVTQGPCRALLADLTCNDARRTRVANAYFSLFMAVGNILG 200
RP AI +FV+GFWILDVANN+ QGPCRALL DL + ++TR ANA+FS FMAVGNILG
Sbjct: 407 VRPRAIGIFVVGFWILDVANNMLQGPCRALLGDLCAGNHQKTRNANAFFSFFMAVGNILG 586
Query: 201 YATGSYSGWYKIFTFTLTPACSI 223
YA G+YS + +F FT T A I
Sbjct: 587 YAAGAYSKLFHVFPFTKTKAXDI 655
Score = 67.8 bits (164), Expect(2) = 2e-62
Identities = 35/65 (53%), Positives = 47/65 (71%)
Frame = +3
Query: 5 TTTNPHRSRTRSSTSTSTSRPVQPVQPRTPLRQLLRVASVASGIQFGWALQLSLLTPYVQ 64
T N + + T + S P P++P +PLR+++ VAS+A+G+QFGWALQLSLLTPYVQ
Sbjct: 42 TKQNHNNNNTLTKPSLHVESP--PLEP-SPLRKIIVVASIAAGVQFGWALQLSLLTPYVQ 212
Query: 65 QLGIP 69
LGIP
Sbjct: 213 LLGIP 227
>TC76982 similar to PIR|T12198|T12198 sucrose transport protein - fava bean,
partial (40%)
Length = 1085
Score = 147 bits (372), Expect = 7e-36
Identities = 83/159 (52%), Positives = 106/159 (66%), Gaps = 6/159 (3%)
Frame = +2
Query: 341 VVLAVTSLLMERLCRK-RGAGFVWGISNIFMAICF-IAMLVLTYAANSIGYVSKGQ---- 394
VVL TSL ++ L R G +WGI N +AIC + +LV A +S Y
Sbjct: 140 VVLGATSLGVDVLARGVGGVKRLWGIVNFPLAICLAMTVLVTKLAQHSRVYADASHTDPL 319
Query: 395 PPPTGIVIAALAIFTILGFPMAITYSVPYALISTHIEPLGLGQGLSMGVLNLAIVVPQIV 454
PP GI ALA+F++LG P+AITYS+P+AL S G GQGLS+GVLNLAIV+PQ++
Sbjct: 320 PPSGGITAGALALFSVLGIPLAITYSIPFALASIFSSSSGAGQGLSLGVLNLAIVIPQMI 499
Query: 455 VSLGSGPWDQLFGGGNSPAFAVAAVAALLSGLLALLAIP 493
VS+ SGPWD LFGGGN PAF V AVAAL SG+L+++ +P
Sbjct: 500 VSVLSGPWDALFGGGNLPAFVVGAVAALASGILSVVLLP 616
>CB894752
Length = 152
Score = 61.6 bits (148), Expect = 7e-10
Identities = 35/50 (70%), Positives = 37/50 (74%)
Frame = +1
Query: 193 MAVGNILGYATGSYSGWYKIFTFTLTPACSISCANLKSAFFLDVAFIVVT 242
MAVG I+GYA GS+SGW KI T LT A IS AN KSAFF DVA IVVT
Sbjct: 1 MAVGTIVGYALGSFSGWDKIRTPILTVASFISGANHKSAFFPDVAAIVVT 150
>BF520636 homologue to GP|5230818|gb|A sucrose transport protein SUT1 {Pisum
sativum}, partial (16%)
Length = 334
Score = 45.8 bits (107), Expect(2) = 2e-05
Identities = 32/81 (39%), Positives = 43/81 (52%), Gaps = 6/81 (7%)
Frame = +1
Query: 341 VVLAVTSLLMERLCRK-RGAGFVWGISNIFMAICF-IAMLVLTYAANSIGYVSKGQ---- 394
VVL TSL ++ L R G +WGI N +AIC + +LV A +S Y
Sbjct: 67 VVLGATSLGVDVLARGVGGVKRLWGIVNFPLAICLAMTVLVTKLAQHSRVYADASHTDPF 246
Query: 395 PPPTGIVIAALAIFTILGFPM 415
PP GI ALA+F++LG P+
Sbjct: 247 PPSGGITAGALALFSVLGIPL 309
Score = 20.4 bits (41), Expect(2) = 2e-05
Identities = 8/16 (50%), Positives = 10/16 (62%)
Frame = +2
Query: 408 FTILGFPMAITYSVPY 423
F F AITYS+P+
Sbjct: 287 FPFSEFLFAITYSIPF 334
>TC79639 similar to GP|10728064|gb|AAF50455.2 CG7060 gene product
{Drosophila melanogaster}, partial (2%)
Length = 757
Score = 31.6 bits (70), Expect = 0.75
Identities = 17/65 (26%), Positives = 33/65 (50%)
Frame = -3
Query: 170 LLADLTCNDARRTRVANAYFSLFMAVGNILGYATGSYSGWYKIFTFTLTPACSISCANLK 229
L+A++ + R++ ++ L +GN +A+ S S + F F +CS+ +K
Sbjct: 404 LIANIAS*ETRKSLISPFNIVLIRFIGNTCVFASSSSSSSFACFFFLPLSSCSLEFPFVK 225
Query: 230 SAFFL 234
+FFL
Sbjct: 224 LSFFL 210
>TC80179 weakly similar to GP|20161298|dbj|BAB90223. hypothetical protein
{Oryza sativa (japonica cultivar-group)}, partial (28%)
Length = 1438
Score = 30.8 bits (68), Expect = 1.3
Identities = 20/54 (37%), Positives = 27/54 (49%), Gaps = 7/54 (12%)
Frame = +2
Query: 54 LQLSLLTPYVQQLGIPHKWASIIWLCGPVS-------GLFVQPLVGHLSDRCSS 100
L +L+ P + Q+G W ++IW PV GL +Q L HLS RC S
Sbjct: 950 LGFTLVGPKLIQIGSLRYWLALIWT-SPVLPTKK*MFGLLMQTLTLHLSRRCKS 1108
>TC89974 similar to GP|7767653|gb|AAF69150.1| F27F5.2 {Arabidopsis
thaliana}, partial (12%)
Length = 1479
Score = 30.8 bits (68), Expect = 1.3
Identities = 28/96 (29%), Positives = 41/96 (42%), Gaps = 11/96 (11%)
Frame = +1
Query: 167 CRALLADLTCNDARRTRVANAYFSLFMAVGN-----------ILGYATGSYSGWYKIFTF 215
CRA + + C DA +R A A FS VG+ I GY YS K T
Sbjct: 652 CRACRSFI*CRDACYSRSAIAVFSTNATVGSKTNSARSSCFIITGYTYAIYSN--KQATD 825
Query: 216 TLTPACSISCANLKSAFFLDVAFIVVTTYLSIVSAH 251
+ C+ SC++ K ++ A++ S +S H
Sbjct: 826 ICSTTCTTSCSSHKQSY----AWLACFRSTSSISIH 921
>BQ146584 similar to PIR|T06751|T0675 hypothetical protein F15B8.120 -
Arabidopsis thaliana, partial (37%)
Length = 415
Score = 28.9 bits (63), Expect = 4.8
Identities = 18/74 (24%), Positives = 30/74 (40%)
Frame = +3
Query: 203 TGSYSGWYKIFTFTLTPACSISCANLKSAFFLDVAFIVVTTYLSIVSAHEVPLSSSGAGE 262
T +S YK+ T + ++CA L +A + A + + HEVP + G
Sbjct: 15 TTQFSSLYKVHTHFIMNMKKVTCAVLVAALSMSAA---------LAATHEVPSPAPAPGP 167
Query: 263 SGSAEEAFMWELFG 276
+ A + L G
Sbjct: 168 ASGASTTVVGSLVG 209
>BG451483 weakly similar to PIR|PN0109|PN01 keratin-like protein - rat
(fragment), partial (15%)
Length = 673
Score = 28.5 bits (62), Expect = 6.3
Identities = 16/51 (31%), Positives = 25/51 (48%), Gaps = 5/51 (9%)
Frame = +1
Query: 1 MPNPTTTNPH-----RSRTRSSTSTSTSRPVQPVQPRTPLRQLLRVASVAS 46
MP P + R R +++TST+RP QP P+R+ R+ A+
Sbjct: 211 MPRPASARSSPRTWLRPRRSCTSATSTARPQQPRSTTRPMRRSTRLPQPAT 363
>AW690928 similar to GP|22093675|d putative pre-mRNA splicing factor
ATP-dependent RNA helicase {Oryza sativa (japonica
cultivar-group), partial (10%)
Length = 638
Score = 28.1 bits (61), Expect = 8.3
Identities = 12/32 (37%), Positives = 18/32 (55%)
Frame = +3
Query: 205 SYSGWYKIFTFTLTPACSISCANLKSAFFLDV 236
+YS + FTF+L P S+SC+N F +
Sbjct: 180 AYSSFIL*FTFSLGPCFSVSCSNTSFRLFSQI 275
>BG589093 similar to PIR|F71608|F716 hypothetical protein PFB0700c - malaria
parasite (Plasmodium falciparum), partial (2%)
Length = 657
Score = 28.1 bits (61), Expect = 8.3
Identities = 15/34 (44%), Positives = 20/34 (58%), Gaps = 6/34 (17%)
Frame = +3
Query: 7 TNPHRSRTRSSTST------STSRPVQPVQPRTP 34
+NP+ SRT S+TST TS P +P + TP
Sbjct: 219 SNPNSSRTTSTTSTFFTTHNKTSLPTRPSRQHTP 320
>TC93439 similar to GP|15081735|gb|AAK82522.1 AT3g06980/F17A9_13
{Arabidopsis thaliana}, partial (3%)
Length = 600
Score = 28.1 bits (61), Expect = 8.3
Identities = 18/60 (30%), Positives = 27/60 (45%), Gaps = 5/60 (8%)
Frame = -2
Query: 236 VAFIVVTTYLSIVSAHEVPLSSSGAGESGSAEE-----AFMWELFGTFKYFSMPVWIVLS 290
V +VV + +V+ + LSSS + S +W TF FS+P +VLS
Sbjct: 368 VVVVVVVVVVXVVAVADAELSSSSSSGCSSCLRFVVFCLVLWMRAFTFSTFSLPKLVVLS 189
>BE322330
Length = 603
Score = 28.1 bits (61), Expect = 8.3
Identities = 8/24 (33%), Positives = 17/24 (70%)
Frame = -1
Query: 274 LFGTFKYFSMPVWIVLSVTALTWI 297
+ G F + S+ VW+ +++T L+W+
Sbjct: 240 IMGQFSFHSIIVWVEVTITGLSWL 169
>AW684990 similar to PIR|F96703|F96 unknown protein 35653-33693 [imported]
- Arabidopsis thaliana, partial (23%)
Length = 517
Score = 28.1 bits (61), Expect = 8.3
Identities = 13/31 (41%), Positives = 20/31 (63%)
Frame = -1
Query: 14 TRSSTSTSTSRPVQPVQPRTPLRQLLRVASV 44
++SST+TS S P P P+T ++L + SV
Sbjct: 94 SKSSTNTSLSNPCLPHPPKTTSQELFKQESV 2
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.325 0.140 0.437
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,407,139
Number of Sequences: 36976
Number of extensions: 247746
Number of successful extensions: 1858
Number of sequences better than 10.0: 38
Number of HSP's better than 10.0 without gapping: 1823
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1849
length of query: 504
length of database: 9,014,727
effective HSP length: 100
effective length of query: 404
effective length of database: 5,317,127
effective search space: 2148119308
effective search space used: 2148119308
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 60 (27.7 bits)
Medicago: description of AC146866.2


