

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146866.12 + phase: 0
(273 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
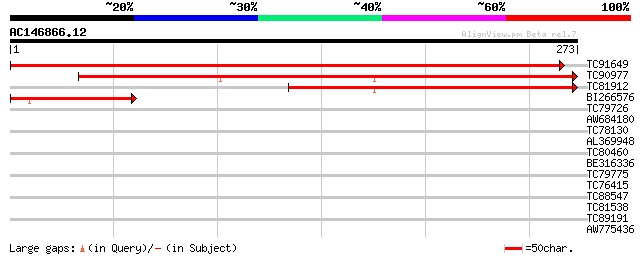
Score E
Sequences producing significant alignments: (bits) Value
TC91649 similar to GP|21700765|gb|AAG38144.1 unknown {Glycine ma... 563 e-161
TC90977 similar to GP|10178227|dbj|BAB11607. ZW10-like protein {... 340 3e-94
TC81912 similar to GP|21700765|gb|AAG38144.1 unknown {Glycine ma... 214 3e-56
BI266576 similar to GP|21700765|gb| unknown {Glycine max}, parti... 102 1e-22
TC79726 similar to GP|17064808|gb|AAL32558.1 Unknown protein {Ar... 32 0.34
AW684180 similar to GP|11993473|gb ubiquitin-specific protease 1... 30 1.3
TC78130 similar to GP|22652125|gb|AAN03626.1 BEL1-related homeot... 29 1.7
AL369948 similar to GP|14587221|db contains ESTs AU108104(C12321... 28 2.9
TC80460 similar to GP|21537277|gb|AAM61618.1 unknown {Arabidopsi... 28 4.9
BE316336 weakly similar to PIR|T05537|T05 probable serine/threon... 27 6.4
TC79775 similar to GP|18175886|gb|AAL59945.1 unknown protein {Ar... 27 6.4
TC76415 similar to PIR|T05479|T05479 hypothetical protein T8O5.1... 27 6.4
TC88547 similar to PIR|F96592|F96592 probable zinc finger protei... 27 8.4
TC81538 similar to GP|9758506|dbj|BAB08914.1 senescence-associat... 27 8.4
TC89191 similar to GP|14272369|emb|CAC40022. bZIP transcription ... 27 8.4
AW775436 27 8.4
>TC91649 similar to GP|21700765|gb|AAG38144.1 unknown {Glycine max}, partial
(93%)
Length = 1143
Score = 563 bits (1452), Expect = e-161
Identities = 267/267 (100%), Positives = 267/267 (100%)
Frame = +3
Query: 1 MDTAPGQSLYPLHHSKTIHLVRHAQGVHNVEGEKNHDAYLSYDFFDANLTPLGWQQVENL 60
MDTAPGQSLYPLHHSKTIHLVRHAQGVHNVEGEKNHDAYLSYDFFDANLTPLGWQQVENL
Sbjct: 60 MDTAPGQSLYPLHHSKTIHLVRHAQGVHNVEGEKNHDAYLSYDFFDANLTPLGWQQVENL 239
Query: 61 QKHVKAIGLSKKIELVVVSPLLRTMQTAVGVFGGEANTDGVNKPPLMIENVGHSDHPAVS 120
QKHVKAIGLSKKIELVVVSPLLRTMQTAVGVFGGEANTDGVNKPPLMIENVGHSDHPAVS
Sbjct: 240 QKHVKAIGLSKKIELVVVSPLLRTMQTAVGVFGGEANTDGVNKPPLMIENVGHSDHPAVS 419
Query: 121 SLNCPPFVAVELCREQMGLHPCDKRRTVSEYRHMFPGIDFSLIETDDDTWWKPEREKKEE 180
SLNCPPFVAVELCREQMGLHPCDKRRTVSEYRHMFPGIDFSLIETDDDTWWKPEREKKEE
Sbjct: 420 SLNCPPFVAVELCREQMGLHPCDKRRTVSEYRHMFPGIDFSLIETDDDTWWKPEREKKEE 599
Query: 181 VTGRGLKFLEWLCTRKEKEIAVVTHSSFLFNTLSAFGNDCHPNIKTEMCAHFANCELRSM 240
VTGRGLKFLEWLCTRKEKEIAVVTHSSFLFNTLSAFGNDCHPNIKTEMCAHFANCELRSM
Sbjct: 600 VTGRGLKFLEWLCTRKEKEIAVVTHSSFLFNTLSAFGNDCHPNIKTEMCAHFANCELRSM 779
Query: 241 VIVDKCMIGSNNSTTNYPGKIPHGPDL 267
VIVDKCMIGSNNSTTNYPGKIPHGPDL
Sbjct: 780 VIVDKCMIGSNNSTTNYPGKIPHGPDL 860
>TC90977 similar to GP|10178227|dbj|BAB11607. ZW10-like protein {Arabidopsis
thaliana}, partial (84%)
Length = 779
Score = 340 bits (872), Expect = 3e-94
Identities = 161/242 (66%), Positives = 196/242 (80%), Gaps = 2/242 (0%)
Frame = +3
Query: 34 KNHDAYLSYDFFDANLTPLGWQQVENLQKHVKAIGLSKKIELVVVSPLLRTMQTAVGVFG 93
KN+ AYL+ D+FDA+LTPLGW+QV+NL+KHV + GL KI+LV+ SPL+RT+QTAVGVFG
Sbjct: 3 KNYKAYLNPDYFDAHLTPLGWEQVDNLRKHVHSSGLINKIDLVIASPLMRTLQTAVGVFG 182
Query: 94 GEANTDG-VNKPPLMIENVGHSDHPAVSSLNCPPFVAVELCREQMGLHPCDKRRTVSEYR 152
GE TD + PLM+ N G+S H A+SS NCPP VA ELCRE +G+HPCDKRR+VSEY+
Sbjct: 183 GEGYTDDKTDVLPLMVANAGNSFHGAISSHNCPPIVAGELCREHLGVHPCDKRRSVSEYQ 362
Query: 153 HMFPGIDFSLIETDDDTWWKPE-REKKEEVTGRGLKFLEWLCTRKEKEIAVVTHSSFLFN 211
+FP +DFSLI++D+D WWK RE KEE+ RG++FL WL TRKEKEIA+VTHS FLF+
Sbjct: 363 FLFPAVDFSLIDSDEDVWWKDNVRETKEELAARGVEFLNWLWTRKEKEIAIVTHSGFLFH 542
Query: 212 TLSAFGNDCHPNIKTEMCAHFANCELRSMVIVDKCMIGSNNSTTNYPGKIPHGPDLPSDA 271
TL+ FGNDCHP +K E+ HFANCELRSMV+VD+ MIGS STTNYPGKIP G D PSDA
Sbjct: 543 TLTTFGNDCHPLVKKEISKHFANCELRSMVLVDRNMIGSEASTTNYPGKIPSGLDKPSDA 722
Query: 272 TD 273
D
Sbjct: 723 VD 728
>TC81912 similar to GP|21700765|gb|AAG38144.1 unknown {Glycine max}, partial
(48%)
Length = 758
Score = 214 bits (545), Expect = 3e-56
Identities = 98/140 (70%), Positives = 117/140 (83%), Gaps = 1/140 (0%)
Frame = +2
Query: 135 EQMGLHPCDKRRTVSEYRHMFPGIDFSLIETDDDTWWKPE-REKKEEVTGRGLKFLEWLC 193
E +G+HPCD RR +SEY +MFP IDFSLIE D+D WKP+ REK EEV RGLKFLEWL
Sbjct: 2 EHLGVHPCDNRRGISEYHNMFPAIDFSLIERDEDILWKPDIREKNEEVAARGLKFLEWLW 181
Query: 194 TRKEKEIAVVTHSSFLFNTLSAFGNDCHPNIKTEMCAHFANCELRSMVIVDKCMIGSNNS 253
TRK+KEIAVV+HS FLF+TLSAFGNDCH N+K+E+C HFANCELRS+VI+D+ IGS+ S
Sbjct: 182 TRKQKEIAVVSHSGFLFHTLSAFGNDCHANVKSEICTHFANCELRSVVIIDRGTIGSDES 361
Query: 254 TTNYPGKIPHGPDLPSDATD 273
+TN+PGKIP G DLPSD +
Sbjct: 362 STNFPGKIPQGLDLPSDVAE 421
>BI266576 similar to GP|21700765|gb| unknown {Glycine max}, partial (18%)
Length = 427
Score = 102 bits (255), Expect = 1e-22
Identities = 48/62 (77%), Positives = 54/62 (86%), Gaps = 1/62 (1%)
Frame = +1
Query: 1 MDTAPGQS-LYPLHHSKTIHLVRHAQGVHNVEGEKNHDAYLSYDFFDANLTPLGWQQVEN 59
MDT+ GQS LYPLH KT+HLVRHAQG HNVEG+KN DAYLSYD FDA+LTPLGW+QV N
Sbjct: 241 MDTSSGQSILYPLHRCKTLHLVRHAQGFHNVEGDKNPDAYLSYDLFDASLTPLGWKQVXN 420
Query: 60 LQ 61
L+
Sbjct: 421 LR 426
>TC79726 similar to GP|17064808|gb|AAL32558.1 Unknown protein {Arabidopsis
thaliana}, partial (36%)
Length = 777
Score = 31.6 bits (70), Expect = 0.34
Identities = 27/83 (32%), Positives = 38/83 (45%)
Frame = +3
Query: 11 PLHHSKTIHLVRHAQGVHNVEGEKNHDAYLSYDFFDANLTPLGWQQVENLQKHVKAIGLS 70
PL +K + LVRH Q N EG S DF + LT G Q E ++ + L
Sbjct: 240 PLKVAKRVVLVRHGQSTWNAEGRIQG----SSDF--SVLTKKGESQAETSRQML----LE 389
Query: 71 KKIELVVVSPLLRTMQTAVGVFG 93
+ SPL R+ +TA ++G
Sbjct: 390 DNFDACFASPLARSKKTAEIIWG 458
>AW684180 similar to GP|11993473|gb ubiquitin-specific protease 14
{Arabidopsis thaliana}, partial (15%)
Length = 527
Score = 29.6 bits (65), Expect = 1.3
Identities = 27/97 (27%), Positives = 41/97 (41%), Gaps = 9/97 (9%)
Frame = -2
Query: 154 MFPGIDFSLIETDDDTWWKPEREKKEEVTGRGLKFLEWLCTRKEKEIA---------VVT 204
+FP ++ S TW + R+ + R ++ WL K I V
Sbjct: 454 VFPALERS-------TWSRK*RKNXKASCCRIVENSGWLAATKVLNIRGGIPSCSADVGV 296
Query: 205 HSSFLFNTLSAFGNDCHPNIKTEMCAHFANCELRSMV 241
++FL ++ S G + P+ K CA F NC LRS V
Sbjct: 295 VATFLLSSFSTCGTEYLPDSKP--CASFVNCMLRSTV 191
>TC78130 similar to GP|22652125|gb|AAN03626.1 BEL1-related homeotic protein 29
{Solanum tuberosum}, partial (58%)
Length = 2740
Score = 29.3 bits (64), Expect = 1.7
Identities = 11/27 (40%), Positives = 17/27 (62%)
Frame = -3
Query: 188 FLEWLCTRKEKEIAVVTHSSFLFNTLS 214
FL W+ T EI++V+H + +TLS
Sbjct: 2018 FLHWILTETSNEISLVSHETITISTLS 1938
>AL369948 similar to GP|14587221|db contains ESTs AU108104(C12321)
C26435(C12321)~similar to Arabidopsis thaliana
chromosome 3 F24P17., partial (5%)
Length = 510
Score = 28.5 bits (62), Expect = 2.9
Identities = 13/26 (50%), Positives = 16/26 (61%)
Frame = +3
Query: 184 RGLKFLEWLCTRKEKEIAVVTHSSFL 209
RGL FLEW E E+A +TH+ L
Sbjct: 93 RGL*FLEWTYALLEIELAAITHNQAL 170
>TC80460 similar to GP|21537277|gb|AAM61618.1 unknown {Arabidopsis
thaliana}, partial (52%)
Length = 1867
Score = 27.7 bits (60), Expect = 4.9
Identities = 12/16 (75%), Positives = 13/16 (81%)
Frame = +1
Query: 238 RSMVIVDKCMIGSNNS 253
RS+V VD C IGSNNS
Sbjct: 307 RSVVPVDSCFIGSNNS 354
>BE316336 weakly similar to PIR|T05537|T05 probable serine/threonine-specific
protein kinase (EC 2.7.1.-) F13M23.300 - Arabidopsis
thaliana, partial (21%)
Length = 614
Score = 27.3 bits (59), Expect = 6.4
Identities = 14/43 (32%), Positives = 23/43 (52%)
Frame = -1
Query: 11 PLHHSKTIHLVRHAQGVHNVEGEKNHDAYLSYDFFDANLTPLG 53
P +HS IH QG+H+ + + +A L +F +LT +G
Sbjct: 491 PSYHSPCIH----TQGIHDQDMHQEEEANLDDEFCSISLTLVG 375
>TC79775 similar to GP|18175886|gb|AAL59945.1 unknown protein {Arabidopsis
thaliana}, partial (82%)
Length = 1147
Score = 27.3 bits (59), Expect = 6.4
Identities = 11/26 (42%), Positives = 19/26 (72%)
Frame = -3
Query: 75 LVVVSPLLRTMQTAVGVFGGEANTDG 100
L +VSPLLR + +++G+ GG+ + G
Sbjct: 839 LSLVSPLLRLLASSIGIGGGKVSPAG 762
>TC76415 similar to PIR|T05479|T05479 hypothetical protein T8O5.180 -
Arabidopsis thaliana, partial (30%)
Length = 1114
Score = 27.3 bits (59), Expect = 6.4
Identities = 11/42 (26%), Positives = 21/42 (49%)
Frame = -1
Query: 9 LYPLHHSKTIHLVRHAQGVHNVEGEKNHDAYLSYDFFDANLT 50
L+P HH K +H H+ +H + ++ Y ++ +LT
Sbjct: 634 LHPHHHDKQLHHHLHSNHIHYT*KPS*YSCFVYYYYYHKSLT 509
>TC88547 similar to PIR|F96592|F96592 probable zinc finger protein
[imported] - Arabidopsis thaliana, partial (43%)
Length = 1765
Score = 26.9 bits (58), Expect = 8.4
Identities = 11/24 (45%), Positives = 15/24 (61%)
Frame = -1
Query: 39 YLSYDFFDANLTPLGWQQVENLQK 62
+LSYDF D + L WQ + +L K
Sbjct: 829 FLSYDFIDVSFL*LPWQIMSSLMK 758
>TC81538 similar to GP|9758506|dbj|BAB08914.1 senescence-associated protein
5-like protein {Arabidopsis thaliana}, partial (92%)
Length = 1073
Score = 26.9 bits (58), Expect = 8.4
Identities = 12/32 (37%), Positives = 18/32 (55%)
Frame = +1
Query: 21 VRHAQGVHNVEGEKNHDAYLSYDFFDANLTPL 52
+R N+ GE N ++ DFF+A LTP+
Sbjct: 439 IRSCLSQTNMCGELNQSFRMAQDFFNARLTPM 534
>TC89191 similar to GP|14272369|emb|CAC40022. bZIP transcription factor
{Arabidopsis thaliana}, partial (42%)
Length = 1158
Score = 26.9 bits (58), Expect = 8.4
Identities = 12/40 (30%), Positives = 21/40 (52%)
Frame = +2
Query: 152 RHMFPGIDFSLIETDDDTWWKPEREKKEEVTGRGLKFLEW 191
+ +FP LI+ D W K E+ +++ V G+ L + W
Sbjct: 689 KELFPHQKRHLIQRH*DVWHKTEKLQRKAV*GKRLMYSSW 808
>AW775436
Length = 659
Score = 26.9 bits (58), Expect = 8.4
Identities = 18/56 (32%), Positives = 25/56 (44%), Gaps = 2/56 (3%)
Frame = +1
Query: 193 CTRKEKEIAVVTHSSFLFNTLSAFGNDCHPNIKTEMCAH--FANCELRSMVIVDKC 246
C E+ I V + FN+L F C PN+ + C+ F L VIV +C
Sbjct: 250 CNSLEEVITGVENVDIAFNSLEVFKLKCLPNL-VKFCSSKCFMKFPLMEEVIVREC 414
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.318 0.135 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,696,863
Number of Sequences: 36976
Number of extensions: 144229
Number of successful extensions: 713
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 708
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 710
length of query: 273
length of database: 9,014,727
effective HSP length: 95
effective length of query: 178
effective length of database: 5,502,007
effective search space: 979357246
effective search space used: 979357246
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 57 (26.6 bits)
Medicago: description of AC146866.12


