

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
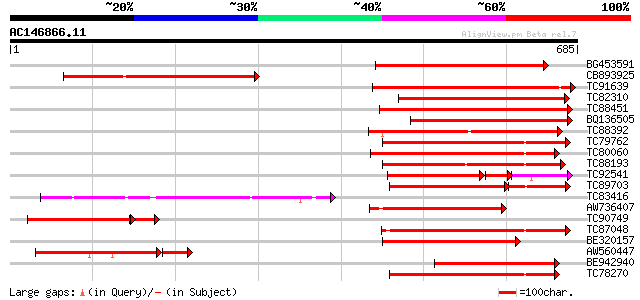
Query= AC146866.11 - phase: 0 /pseudo
(685 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BG453591 similar to PIR|T05181|T05 S-receptor kinase (EC 2.7.1.-... 332 3e-91
CB893925 weakly similar to GP|1272349|gb|A secreted glycoprotein... 316 2e-86
TC91639 similar to PIR|T05181|T05181 S-receptor kinase (EC 2.7.1... 301 5e-82
TC82310 similar to PIR|T05181|T05181 S-receptor kinase (EC 2.7.1... 283 2e-76
TC88451 similar to GP|836954|gb|AAC23542.1|| receptor protein ki... 278 6e-75
BQ136505 similar to PIR|T05754|T05 S-receptor kinase (EC 2.7.1.-... 266 2e-71
TC88392 similar to GP|11275527|dbj|BAB18292. putative receptor-l... 254 1e-67
TC79762 similar to PIR|H96731|H96731 hypothetical protein F5A18.... 252 3e-67
TC80060 similar to PIR|D96574|D96574 hypothetical protein T3F20.... 233 2e-61
TC88193 similar to PIR|G96602|G96602 probable receptor protein k... 232 3e-61
TC92541 similar to PIR|T02153|T02153 protein kinase homolog T1F1... 145 4e-58
TC89703 similar to GP|11994595|dbj|BAB02650. receptor-like serin... 172 7e-58
TC83416 weakly similar to GP|3056594|gb|AAC13905.1| T1F9.15 {Ara... 218 8e-57
AW736407 similar to GP|4008008|gb| receptor-like protein kinase ... 215 5e-56
TC90749 similar to PIR|T14450|T14450 serine/threonine kinase (EC... 191 3e-55
TC87048 similar to GP|8778594|gb|AAF79602.1| F5M15.3 {Arabidopsi... 208 6e-54
BE320157 weakly similar to PIR|T04835|T04 probable serine/threon... 206 3e-53
AW560447 similar to PIR|T14520|T14 probable S-receptor kinase (E... 179 5e-53
BE942940 weakly similar to GP|13506749|gb receptor-like protein ... 204 7e-53
TC78270 similar to GP|16649103|gb|AAL24403.1 Unknown protein {Ar... 203 2e-52
>BG453591 similar to PIR|T05181|T05 S-receptor kinase (EC 2.7.1.-) T6K22.120
precursor - Arabidopsis thaliana, partial (24%)
Length = 640
Score = 332 bits (851), Expect = 3e-91
Identities = 160/211 (75%), Positives = 186/211 (87%), Gaps = 1/211 (0%)
Frame = +3
Query: 442 EHEDFDLPFFDLATIIKATDNFSTNNKLGEGGFGPVYKATLQ-DGHVIAVKRLSGNSEQG 500
+ +DF+LPFF+L+TII AT++FS +NKLGEGGFGPVYK TL D IAVKRLSG+S+QG
Sbjct: 6 DEQDFELPFFNLSTIIDATNDFSNDNKLGEGGFGPVYKGTLVLDRREIAVKRLSGSSKQG 185
Query: 501 SKEFKNEVILCVKLQHRNLVKVLGCCIEGDEKLLIYEYMPNKSLDSFLFDPTQSKLLSWS 560
++EFKNEVILC KLQHRNLVKVLGCCI+G+EK+LIYEYMPN+SLDSFLFD Q KLL WS
Sbjct: 186 TREFKNEVILCSKLQHRNLVKVLGCCIQGEEKMLIYEYMPNRSLDSFLFDQAQKKLLDWS 365
Query: 561 MRLNILNAIARGIQYLHQDSRLRIIHRDLKASNILLDNEMDPKISDFGMARMCGGDQIEG 620
R NI+ IARG+ YLHQDSRLRIIHRDLK SNILLDN+M+PKISDFG+A++CG DQ+EG
Sbjct: 366 KRFNIICGIARGLIYLHQDSRLRIIHRDLKPSNILLDNDMNPKISDFGLAKICGDDQVEG 545
Query: 621 KTRRIVGTYGYMAPEYVIHGLFSIKSDVFSF 651
T R+VGT+GYMAPEY I GLFSIKSDVFSF
Sbjct: 546 NTNRVVGTHGYMAPEYAIDGLFSIKSDVFSF 638
>CB893925 weakly similar to GP|1272349|gb|A secreted glycoprotein 3 {Ipomoea
trifida}, partial (19%)
Length = 715
Score = 316 bits (809), Expect = 2e-86
Identities = 152/239 (63%), Positives = 186/239 (77%), Gaps = 2/239 (0%)
Frame = +2
Query: 66 VGIWYKNIPVRRVVWVANRNNPTKDDSSKLIISQDGNLVLLNHNDSLVWSTNAS-RKASS 124
VGIWYKNIPVRRVVWVANR+NP KD SSKLIISQD NLVLL+ N SL+WSTNA+ K SS
Sbjct: 2 VGIWYKNIPVRRVVWVANRDNPIKDTSSKLIISQDRNLVLLDKNQSLLWSTNATIEKVSS 181
Query: 125 PVVQLLNNGNLVLRDEKDNNEESFLWQGFDHPCDTLLPGMTFGYNRKLDFYWNLTAWKNE 184
P+ QLL++GNLVL K+ EE FLWQ FD+PCDT+L GM G++++ D +L AWKN
Sbjct: 182 PIAQLLDDGNLVL---KNGGEEHFLWQSFDYPCDTILSGMKAGWDKRKDLNRSLVAWKNW 352
Query: 185 DDPSSGDLYASVVFTSNPESMIWKGSTKICRSGPWN-PLSSGVVGMKPNPLYDYKVVNNE 243
DDPSS D +++V T NPES IWKG TK+ R+GPW P SSGV+G+ NPLYDY+ VNN+
Sbjct: 353 DDPSSSDFTSAMVLTPNPESFIWKGLTKLYRTGPWTGPRSSGVIGLTENPLYDYEFVNNQ 532
Query: 244 DEVYYQFVLRNSSVTSIAVLNQTLLIRQRLVYVPESKIWSVYQIMPSDTCEYYNVCGAN 302
DEVYY F L+NSSV S VLN+TL +R RL+++ ESK W+VYQ +P D+C+ YNVC N
Sbjct: 533 DEVYYLFTLKNSSVVSFVVLNRTLSVRLRLIWISESKTWNVYQTLPQDSCDVYNVCREN 709
>TC91639 similar to PIR|T05181|T05181 S-receptor kinase (EC 2.7.1.-)
T6K22.120 precursor - Arabidopsis thaliana, partial
(28%)
Length = 1313
Score = 301 bits (771), Expect = 5e-82
Identities = 150/245 (61%), Positives = 185/245 (75%)
Frame = +3
Query: 439 DGGEHEDFDLPFFDLATIIKATDNFSTNNKLGEGGFGPVYKATLQDGHVIAVKRLSGNSE 498
+G + +LPFF+ + + AT+NFS NKLG+GGFGPVYK L G IAVKRLS S
Sbjct: 336 EGNQLSKVELPFFNFSCMSSATNNFSEENKLGQGGFGPVYKGKLPSGEEIAVKRLSRRSG 515
Query: 499 QGSKEFKNEVILCVKLQHRNLVKVLGCCIEGDEKLLIYEYMPNKSLDSFLFDPTQSKLLS 558
QG EFKNE+ L +LQHRNLVK++GC IEGDEKLL+YE+M NKSLD FLFDP + L
Sbjct: 516 QGLDEFKNEMRLFAQLQHRNLVKLMGCSIEGDEKLLVYEFMLNKSLDRFLFDPIKKTQLD 695
Query: 559 WSMRLNILNAIARGIQYLHQDSRLRIIHRDLKASNILLDNEMDPKISDFGMARMCGGDQI 618
W+ R I+ IARG+ YLH+DSRLRIIHRDLKASNILLD M+PKISDFG+AR+ GG+Q
Sbjct: 696 WARRYEIIEGIARGLLYLHRDSRLRIIHRDLKASNILLDENMNPKISDFGLARIFGGNQN 875
Query: 619 EGKTRRIVGTYGYMAPEYVIHGLFSIKSDVFSFGVLLLETISGKKNRTLTYHEHDHNLIW 678
E ++VGTYGYM+PEY + GL S+KSDV+SFGVLLLE +SG++N T H D +LI
Sbjct: 876 EENATKVVGTYGYMSPEYAMEGLVSVKSDVYSFGVLLLEIVSGRRN-TSFRHSDDSSLIG 1052
Query: 679 HVSTN 683
+VS +
Sbjct: 1053YVSVS 1067
Score = 34.7 bits (78), Expect = 0.12
Identities = 18/44 (40%), Positives = 28/44 (62%)
Frame = +3
Query: 401 GCSLWFNDLIDLRLSQSSEGDDLYIRVDRDSNFGKKERDGGEHE 444
GC +W+ DL+D+ Q EG+ L+IR+ S+ G DGG++E
Sbjct: 24 GCMVWYGDLVDILHFQHGEGNALHIRL-AYSDLG----DGGKNE 140
>TC82310 similar to PIR|T05181|T05181 S-receptor kinase (EC 2.7.1.-)
T6K22.120 precursor - Arabidopsis thaliana, partial
(24%)
Length = 696
Score = 283 bits (724), Expect = 2e-76
Identities = 140/207 (67%), Positives = 161/207 (77%)
Frame = +1
Query: 470 GEGGFGPVYKATLQDGHVIAVKRLSGNSEQGSKEFKNEVILCVKLQHRNLVKVLGCCIEG 529
G+GGFG VYK L D IAVKRLS S QG +E KNE+IL KLQHRNLV++L CCIE
Sbjct: 1 GKGGFGTVYKGVLADEKEIAVKRLSKTSSQGVEELKNEIILIAKLQHRNLVRLLACCIEQ 180
Query: 530 DEKLLIYEYMPNKSLDSFLFDPTQSKLLSWSMRLNILNAIARGIQYLHQDSRLRIIHRDL 589
+EKLLIYEY+PN SLD LFD + L+W RLNI+N IA+G+ YLH+DSRLR+IHRDL
Sbjct: 181 NEKLLIYEYLPNSSLDFHLFDMVKGAQLAWRQRLNIINGIAKGLLYLHEDSRLRVIHRDL 360
Query: 590 KASNILLDNEMDPKISDFGMARMCGGDQIEGKTRRIVGTYGYMAPEYVIHGLFSIKSDVF 649
KASNILLD EM+PKISDFG+AR GGDQ E T R+VGTYGYMAPEY + GLFS+KSDVF
Sbjct: 361 KASNILLDQEMNPKISDFGLARTFGGDQDEANTIRVVGTYGYMAPEYAMEGLFSVKSDVF 540
Query: 650 SFGVLLLETISGKKNRTLTYHEHDHNL 676
SFGVLLLE ISG+KN EH +L
Sbjct: 541 SFGVLLLEIISGRKNSKFYLSEHGQSL 621
>TC88451 similar to GP|836954|gb|AAC23542.1|| receptor protein kinase
{Ipomoea trifida}, partial (28%)
Length = 1276
Score = 278 bits (710), Expect = 6e-75
Identities = 139/233 (59%), Positives = 172/233 (73%)
Frame = +3
Query: 447 DLPFFDLATIIKATDNFSTNNKLGEGGFGPVYKATLQDGHVIAVKRLSGNSEQGSKEFKN 506
D+ F+ +I++AT FS NKLG+GG+GPVYK L G IAVKRLS S QG EFKN
Sbjct: 180 DIKVFNFTSILEATMEFSPENKLGQGGYGPVYKGILATGQEIAVKRLSKTSGQGIVEFKN 359
Query: 507 EVILCVKLQHRNLVKVLGCCIEGDEKLLIYEYMPNKSLDSFLFDPTQSKLLSWSMRLNIL 566
E++L +LQH+NLV++LGCCI +E++LIYEYMPNKSLD +LFD T+ KLL W R NI+
Sbjct: 360 ELLLICELQHKNLVQLLGCCIHEEERILIYEYMPNKSLDFYLFDCTKKKLLDWKKRFNII 539
Query: 567 NAIARGIQYLHQDSRLRIIHRDLKASNILLDNEMDPKISDFGMARMCGGDQIEGKTRRIV 626
IA+G+ YLH+ SRL+IIHRDLKASNILLD M+PKI+DFGMARM + T RIV
Sbjct: 540 EGIAQGLLYLHKYSRLKIIHRDLKASNILLDENMNPKIADFGMARMFTQQESVVNTNRIV 719
Query: 627 GTYGYMAPEYVIHGLFSIKSDVFSFGVLLLETISGKKNRTLTYHEHDHNLIWH 679
GTYGYM+PEY + G+ S KSDV+SFGVLLLE + G KN + + NLI H
Sbjct: 720 GTYGYMSPEYAMEGVCSTKSDVYSFGVLLLEIVCGIKNNSFYDVDRPLNLIGH 878
>BQ136505 similar to PIR|T05754|T05 S-receptor kinase (EC 2.7.1.-) M4I22.110
precursor - Arabidopsis thaliana, partial (23%)
Length = 618
Score = 266 bits (680), Expect = 2e-71
Identities = 130/195 (66%), Positives = 156/195 (79%)
Frame = +2
Query: 485 GHVIAVKRLSGNSEQGSKEFKNEVILCVKLQHRNLVKVLGCCIEGDEKLLIYEYMPNKSL 544
G +A+KR S S+QG +EFKNEV+L KLQHRNLVK+LGCCI +EKLLIYEYMPN+SL
Sbjct: 20 GKEVAIKRNSKMSDQGLEEFKNEVLLIAKLQHRNLVKLLGCCIHREEKLLIYEYMPNRSL 199
Query: 545 DSFLFDPTQSKLLSWSMRLNILNAIARGIQYLHQDSRLRIIHRDLKASNILLDNEMDPKI 604
D F+FD T+SKLL WS R +I+ +ARG+ YLHQDSRLRIIHRDLK SNILLD M+PKI
Sbjct: 200 DYFIFDETRSKLLDWSKRSHIIAGVARGLLYLHQDSRLRIIHRDLKLSNILLDALMNPKI 379
Query: 605 SDFGMARMCGGDQIEGKTRRIVGTYGYMAPEYVIHGLFSIKSDVFSFGVLLLETISGKKN 664
SDFG+AR GDQ+E KTR++VGTYGYM PEY +HG +S+KSDVFSFGV++LE ISGKK
Sbjct: 380 SDFGLARTFCGDQVEAKTRKLVGTYGYMPPEYAVHGRYSMKSDVFSFGVIVLEIISGKKI 559
Query: 665 RTLTYHEHDHNLIWH 679
+ H NL+ H
Sbjct: 560 KVFYDPXHSLNLLGH 604
>TC88392 similar to GP|11275527|dbj|BAB18292. putative receptor-like protein
kinase {Oryza sativa (japonica cultivar-group)}, partial
(61%)
Length = 1017
Score = 254 bits (648), Expect = 1e-67
Identities = 126/241 (52%), Positives = 171/241 (70%), Gaps = 7/241 (2%)
Frame = +1
Query: 434 GKKERDGGEHEDFDLP-------FFDLATIIKATDNFSTNNKLGEGGFGPVYKATLQDGH 486
G DGG+ ++ + F+L T+ AT+ FS N+LG GGFGPV+K + +G
Sbjct: 109 GTGSEDGGDDDEDTVDPGDSGNLLFELNTLQLATNFFSELNQLGRGGFGPVFKGLMPNGE 288
Query: 487 VIAVKRLSGNSEQGSKEFKNEVILCVKLQHRNLVKVLGCCIEGDEKLLIYEYMPNKSLDS 546
+A+K+LS S QG +EF NEV L +++QH+NLV +LGCC EG EK+L+YEY+PNKSLD
Sbjct: 289 EVAIKKLSMESRQGIREFTNEVRLLLRIQHKNLVTLLGCCAEGPEKMLVYEYLPNKSLDH 468
Query: 547 FLFDPTQSKLLSWSMRLNILNAIARGIQYLHQDSRLRIIHRDLKASNILLDNEMDPKISD 606
FLFD +S L W R I+ IARG+ YLH+++ RIIHRD+KASNILLD +++PKISD
Sbjct: 469 FLFDKKRS--LDWMTRFRIVTGIARGLLYLHEEAPERIIHRDIKASNILLDEKLNPKISD 642
Query: 607 FGMARMCGGDQIEGKTRRIVGTYGYMAPEYVIHGLFSIKSDVFSFGVLLLETISGKKNRT 666
FG+AR+ G+ +T RI T+GYMAPEY + G S+K+DVFS+GVL+LE +SG+KN
Sbjct: 643 FGLARLFPGEDTHVQTFRISRTHGYMAPEYALRGYLSVKTDVFSYGVLVLEIVSGRKNHD 822
Query: 667 L 667
L
Sbjct: 823 L 825
>TC79762 similar to PIR|H96731|H96731 hypothetical protein F5A18.8
[imported] - Arabidopsis thaliana, partial (55%)
Length = 1564
Score = 252 bits (644), Expect = 3e-67
Identities = 121/228 (53%), Positives = 164/228 (71%), Gaps = 1/228 (0%)
Frame = +3
Query: 451 FDLATIIKATDNFSTNNKLGEGGFGPVYKATLQDGHVIAVKRLSGNSEQGSKEFKNEVIL 510
F T++ AT NF+ +KLGEGGFGPVYK L DG +AVK+LS S QG KEF NE L
Sbjct: 348 FSYETLLSATKNFNATHKLGEGGFGPVYKGKLSDGREVAVKKLSQTSNQGKKEFMNEAKL 527
Query: 511 CVKLQHRNLVKVLGCCIEGDEKLLIYEYMPNKSLDSFLFDPTQSK-LLSWSMRLNILNAI 569
++QH+N+V +LG C+ G EK+L+YEY+P++SLD FLF + + L W R I+ +
Sbjct: 528 LARVQHKNVVNLLGYCVHGTEKILVYEYVPHESLDKFLFKEAEKREQLDWKRRFGIITGV 707
Query: 570 ARGIQYLHQDSRLRIIHRDLKASNILLDNEMDPKISDFGMARMCGGDQIEGKTRRIVGTY 629
A+G+ YLH+DS IIHRD+KASNILLD++ KI+DFGMAR+ DQ + KT R+ GT
Sbjct: 708 AKGLLYLHEDSHNCIIHRDIKASNILLDDKWTAKIADFGMARLFPEDQSQVKT-RVAGTN 884
Query: 630 GYMAPEYVIHGLFSIKSDVFSFGVLLLETISGKKNRTLTYHEHDHNLI 677
GYMAPEY++HG S+K+DVFS+GVL+LE I+G++N + +HNL+
Sbjct: 885 GYMAPEYMMHGRLSVKADVFSYGVLVLELITGQRNSSFNLXVEEHNLL 1028
>TC80060 similar to PIR|D96574|D96574 hypothetical protein T3F20.24
[imported] - Arabidopsis thaliana, partial (32%)
Length = 1547
Score = 233 bits (594), Expect = 2e-61
Identities = 118/230 (51%), Positives = 158/230 (68%), Gaps = 1/230 (0%)
Frame = +1
Query: 436 KERDGGEHEDFDLPFFDLATIIKATDNFSTNNKLGEGGFGPVYKATLQDGHVIAVKRLSG 495
K++ E + ++ L I AT+NF NK+GEGGFGPVYK L DG VIAVK+LS
Sbjct: 112 KDQTDKELLELKTGYYSLRQIKVATNNFDPKNKIGEGGFGPVYKGVLSDGAVIAVKQLSS 291
Query: 496 NSEQGSKEFKNEVILCVKLQHRNLVKVLGCCIEGDEKLLIYEYMPNKSLDSFLFDPTQSK 555
S+QG++EF NE+ + LQH NLVK+ GCCIEG++ LL+YEYM N SL LF + +
Sbjct: 292 KSKQGNREFVNEIGMISALQHPNLVKLYGCCIEGNQLLLVYEYMENNSLARALFGKPEQR 471
Query: 556 L-LSWSMRLNILNAIARGIQYLHQDSRLRIIHRDLKASNILLDNEMDPKISDFGMARMCG 614
L L W R+ I IARG+ YLH++SRL+I+HRD+KA+N+LLD ++ KISDFG+A++
Sbjct: 472 LNLDWRTRMKICVGIARGLAYLHEESRLKIVHRDIKATNVLLDKNLNAKISDFGLAKLDE 651
Query: 615 GDQIEGKTRRIVGTYGYMAPEYVIHGLFSIKSDVFSFGVLLLETISGKKN 664
+ T RI GT GYMAPEY + G + K+DV+SFGV+ LE +SG N
Sbjct: 652 EENTHIST-RIAGTIGYMAPEYAMRGYLTDKADVYSFGVVALEIVSGMSN 798
>TC88193 similar to PIR|G96602|G96602 probable receptor protein kinase
F14G9.24 [imported] - Arabidopsis thaliana, partial
(20%)
Length = 1815
Score = 232 bits (592), Expect = 3e-61
Identities = 115/221 (52%), Positives = 155/221 (70%)
Frame = +3
Query: 451 FDLATIIKATDNFSTNNKLGEGGFGPVYKATLQDGHVIAVKRLSGNSEQGSKEFKNEVIL 510
F + AT +F+ +NKLGEGGFGPVYK TL DG +AVK+LS S QG +F E+
Sbjct: 540 FSYYELKNATSDFNRDNKLGEGGFGPVYKGTLNDGRFVAVKQLSIGSHQGKSQFIAEIAT 719
Query: 511 CVKLQHRNLVKVLGCCIEGDEKLLIYEYMPNKSLDSFLFDPTQSKLLSWSMRLNILNAIA 570
+QHRNLVK+ GCCIEG+++LL+YEY+ NKSLD LF L+WS R ++ +A
Sbjct: 720 ISAVQHRNLVKLYGCCIEGNKRLLVYEYLENKSLDQALFG--NVLFLNWSTRYDVCMGVA 893
Query: 571 RGIQYLHQDSRLRIIHRDLKASNILLDNEMDPKISDFGMARMCGGDQIEGKTRRIVGTYG 630
RG+ YLH++SRLRI+HRD+KASNILLD+E+ PK+SDFG+A++ + T R+ GT G
Sbjct: 894 RGLTYLHEESRLRIVHRDVKASNILLDSELVPKLSDFGLAKLYDDKKTHIST-RVAGTIG 1070
Query: 631 YMAPEYVIHGLFSIKSDVFSFGVLLLETISGKKNRTLTYHE 671
Y+APEY + G + K+DVFSFGV+ LE +SG+ N + E
Sbjct: 1071YLAPEYAMRGRLTEKADVFSFGVVALELVSGRPNSDSSLEE 1193
>TC92541 similar to PIR|T02153|T02153 protein kinase homolog T1F15.1 -
Arabidopsis thaliana, partial (22%)
Length = 791
Score = 145 bits (367), Expect(3) = 4e-58
Identities = 69/117 (58%), Positives = 88/117 (74%)
Frame = +2
Query: 457 IKATDNFSTNNKLGEGGFGPVYKATLQDGHVIAVKRLSGNSEQGSKEFKNEVILCVKLQH 516
++AT +FS NKLG+GG+GPVYK L G IAVKRLS S QG EFKNE++L +LQH
Sbjct: 2 LEATIDFSPENKLGQGGYGPVYKGILPTGQEIAVKRLSKTSRQGIVEFKNELVLICELQH 181
Query: 517 RNLVKVLGCCIEGDEKLLIYEYMPNKSLDSFLFDPTQSKLLSWSMRLNILNAIARGI 573
NLV++LGCCI +E++LIYEYM NKSLD +LFD T+ K L W RLNI+ I++G+
Sbjct: 182 TNLVQLLGCCIHEEERILIYEYMSNKSLDFYLFDSTRRKCLDWKKRLNIIEGISQGL 352
Score = 61.2 bits (147), Expect(3) = 4e-58
Identities = 39/98 (39%), Positives = 48/98 (48%), Gaps = 25/98 (25%)
Frame = +1
Query: 607 FGMARMCGGDQIEGKTRRIVGT-------------------------YGYMAPEYVIHGL 641
FGMARM + T RIVGT GYM+PEY + G+
Sbjct: 454 FGMARMFTQQESVVNTNRIVGT**VL*TFLMIT*RKRF*FFK*FFLSSGYMSPEYAMEGI 633
Query: 642 FSIKSDVFSFGVLLLETISGKKNRTLTYHEHDHNLIWH 679
S KSDV+SFGVLLLE I G++N + + NLI H
Sbjct: 634 CSTKSDVYSFGVLLLEIICGRRNNSFYDVDRPLNLIGH 747
Score = 57.8 bits (138), Expect(3) = 4e-58
Identities = 27/33 (81%), Positives = 30/33 (90%)
Frame = +3
Query: 575 YLHQDSRLRIIHRDLKASNILLDNEMDPKISDF 607
YLH+ SRL+IIHRDLKASNILLD M+PKISDF
Sbjct: 357 YLHKYSRLKIIHRDLKASNILLDENMNPKISDF 455
>TC89703 similar to GP|11994595|dbj|BAB02650. receptor-like serine/threonine
kinase {Arabidopsis thaliana}, partial (26%)
Length = 1212
Score = 172 bits (437), Expect(2) = 7e-58
Identities = 80/146 (54%), Positives = 109/146 (73%), Gaps = 1/146 (0%)
Frame = +2
Query: 459 ATDNFSTNNKLGEGGFGPVYKATLQDGHVIAVKRLSGNSEQGSKEFKNEVILCVKLQHRN 518
AT+NF +NK+GEGGFGPVYK L DG +IAVK LS S+QG++EF NE+ + LQH
Sbjct: 20 ATNNFDISNKIGEGGFGPVYKGRLSDGTLIAVKLLSSKSKQGNREFLNEIGMISALQHPQ 199
Query: 519 LVKVLGCCIEGDEKLLIYEYMPNKSLDSFLFDPTQSKL-LSWSMRLNILNAIARGIQYLH 577
LVK+ GCC+EGD+ +LIYEY+ N SL LF P + ++ L W R I IARG+ YLH
Sbjct: 200 LVKLYGCCVEGDQLMLIYEYLENNSLARALFGPAEHQIRLDWPTRYKICVGIARGLAYLH 379
Query: 578 QDSRLRIIHRDLKASNILLDNEMDPK 603
++SRL+++HRD+KA+N+LLD +++PK
Sbjct: 380 EESRLKVVHRDIKATNVLLDKDLNPK 457
Score = 70.5 bits (171), Expect(2) = 7e-58
Identities = 35/75 (46%), Positives = 48/75 (63%)
Frame = +3
Query: 603 KISDFGMARMCGGDQIEGKTRRIVGTYGYMAPEYVIHGLFSIKSDVFSFGVLLLETISGK 662
KISDFG+A++ + TR I GTYGYMAPEY +HG + K+DV+SFG++ LE + G
Sbjct: 456 KISDFGLAKLDEEENTHISTR-IAGTYGYMAPEYAMHGYLTDKADVYSFGIVALEILHGS 632
Query: 663 KNRTLTYHEHDHNLI 677
N L E +L+
Sbjct: 633 NNTILRQKEEAFHLL 677
>TC83416 weakly similar to GP|3056594|gb|AAC13905.1| T1F9.15 {Arabidopsis
thaliana}, partial (13%)
Length = 1142
Score = 218 bits (554), Expect = 8e-57
Identities = 132/370 (35%), Positives = 187/370 (49%), Gaps = 14/370 (3%)
Frame = +3
Query: 38 LSNGSTLVSKDGTFEMGFFRPGKSLNRYVGIWYKNIPVRRVVWVANRNNPTKDDSSKLI- 96
+ + TL S G F +GFF P S NRYVGIW K VVWVANRN P +DSS ++
Sbjct: 12 IKDSETLSSNSGNFTLGFFTPQNSTNRYVGIWCKTQLF--VVWVANRNQPLINDSSGVLE 185
Query: 97 ISQDGNLVLLNHNDSLVWSTNASR-KASSPVVQLLNNGNLVLRDEKDNNEESFLWQGFDH 155
IS D N+VLLN ++VWSTN + S+ V L + GNL+L + E +WQ DH
Sbjct: 186 ISNDNNIVLLNGKKNVVWSTNLNNVSTSNSSVTLSDYGNLILFE---TTTEKTIWQSSDH 356
Query: 156 PCDTLLPGMTFGYNRKLDFYWNLTAWKNEDDPSSGDLYASVVFTSNPESMIWKGSTKICR 215
P +++LP +TF N LT+WK +DPS+G + + PE I K + R
Sbjct: 357 PTNSILPSLTFTSNM------TLTSWKTSNDPSNGSFTLGIERLNMPEVFIRKENRPYWR 518
Query: 216 SGPWN-PLSSGVVGMKPNPLYDYKVVNNEDEVYYQFVLRNSSVTSIA-VLNQTLLIRQRL 273
SGPWN + G+ M L + + V R + V+N T + ++
Sbjct: 519 SGPWNNQIFIGIEDMSALYLNGFHFQKDRTGGTLDLVFRADDYGLVMYVVNSTGQMNEKS 698
Query: 274 VYVPESKIWSVYQIMPSDTCEYYNVCGANAQCTIDGSPMCQCLPGFKPKSPQQWNSMDWT 333
+ + + SD C+ Y CG+ C GSP+C+CL GF+P++ Q+WN +WT
Sbjct: 699 WSIENEEWMDTWTNQRSD-CDLYGFCGSFGICNSKGSPICRCLEGFEPRNNQEWNRQNWT 875
Query: 334 QGCVRGGNWSCGIKNR----------DGFQKFVRMKLPDTTNSWINLNMTLQDCKTKCLQ 383
GCVR C N D F K +K+PD L++ +C+ +CL
Sbjct: 876 NGCVRKTPLQCESANNQNKSANGNDADSFMKLTLVKVPDFAEL---LSVEQDECENQCLM 1046
Query: 384 NCSCTAYTYL 393
NCSCTAY+Y+
Sbjct: 1047NCSCTAYSYV 1076
>AW736407 similar to GP|4008008|gb| receptor-like protein kinase {Arabidopsis
thaliana}, partial (17%)
Length = 620
Score = 215 bits (547), Expect = 5e-56
Identities = 109/166 (65%), Positives = 128/166 (76%)
Frame = +2
Query: 435 KKERDGGEHEDFDLPFFDLATIIKATDNFSTNNKLGEGGFGPVYKATLQDGHVIAVKRLS 494
KKE++ GE FD +TI AT+NFS NKLGEGGFGPVYK + DG IAVKRLS
Sbjct: 131 KKEKEDGELATI----FDFSTITNATNNFSVRNKLGEGGFGPVYKGVMVDGQEIAVKRLS 298
Query: 495 GNSEQGSKEFKNEVILCVKLQHRNLVKVLGCCIEGDEKLLIYEYMPNKSLDSFLFDPTQS 554
S QG++EFKNEV L LQHRNLVK+LGC I+ DEK+LIYE+MPN+SLD F+FD T+S
Sbjct: 299 KTSGQGTEEFKNEVKLMATLQHRNLVKLLGCSIQQDEKMLIYEFMPNRSLDFFIFDTTRS 478
Query: 555 KLLSWSMRLNILNAIARGIQYLHQDSRLRIIHRDLKASNILLDNEM 600
KLL W+ RL I++ IARG+ YLHQDS LRIIHRDLK SNILLD +M
Sbjct: 479 KLLDWTKRLEIIDGIARGLLYLHQDSTLRIIHRDLKTSNILLDIDM 616
>TC90749 similar to PIR|T14450|T14450 serine/threonine kinase (EC 2.7.1.-)
BRLK - wild cabbage, partial (6%)
Length = 495
Score = 191 bits (485), Expect(2) = 3e-55
Identities = 95/133 (71%), Positives = 111/133 (83%), Gaps = 2/133 (1%)
Frame = +3
Query: 22 LSHVSYATDTITKSASLSNGSTLVSKDGTFEMGFFRPGKSLNRYVGIWYKNIPVRRVVWV 81
L +SYATDTIT+ S+ +GS+L+SKDG+FE+GFF PG S NRYVG+WYKNIPVRRVVWV
Sbjct: 9 LPQISYATDTITQPTSIRDGSSLISKDGSFELGFFSPGSSSNRYVGLWYKNIPVRRVVWV 188
Query: 82 ANRNNPTKDDSSKLIISQDGNLVLLNHNDSLV-WSTNASRKASSPVVQLLNNGNLVLRDE 140
NR+NP KDDSSKL ISQDGNL+LLN N+SLV WSTN S AS+ VVQLL+NGNLVL+D
Sbjct: 189 LNRDNPIKDDSSKLTISQDGNLMLLNQNESLVWWSTNISTNASNRVVQLLDNGNLVLKDV 368
Query: 141 -KDNNEESFLWQG 152
+N ESFLWQG
Sbjct: 369 INSDNGESFLWQG 407
Score = 43.1 bits (100), Expect(2) = 3e-55
Identities = 18/37 (48%), Positives = 25/37 (66%)
Frame = +1
Query: 145 EESFLWQGFDHPCDTLLPGMTFGYNRKLDFYWNLTAW 181
E+ F +GFD+PCDTLLP M G +++ +LTAW
Sbjct: 385 EKVFCGKGFDYPCDTLLPRMKIGIDKRTGLNRHLTAW 495
>TC87048 similar to GP|8778594|gb|AAF79602.1| F5M15.3 {Arabidopsis
thaliana}, partial (77%)
Length = 1571
Score = 208 bits (529), Expect = 6e-54
Identities = 109/230 (47%), Positives = 155/230 (67%), Gaps = 2/230 (0%)
Frame = +1
Query: 450 FFDLATIIKATDNFSTNNKLGEGGFGPVYKATLQDGHVIAVKRLSGNSEQGSKEFKNEVI 509
F +LAT AT F N +GEGGFG V+K L G ++AVK+LS + QG +EF EV+
Sbjct: 436 FRELAT---ATRGFKEANLIGEGGFGKVFKGRLSTGELVAVKQLSHDGRQGFQEFVTEVL 606
Query: 510 LCVKLQHRNLVKVLGCCIEGDEKLLIYEYMPNKSLDSFLFD-PTQSKLLSWSMRLNILNA 568
+ L H NLVK++G C +GD++LL+YEYMP SL+ LFD P + LSWS R+ I
Sbjct: 607 MLSLLHHSNLVKLIGYCTDGDQRLLVYEYMPMGSLEDHLFDLPQDKEPLSWSSRMKIAVG 786
Query: 569 IARGIQYLHQDSRLRIIHRDLKASNILLDNEMDPKISDFGMARMCG-GDQIEGKTRRIVG 627
ARG++YLH + +I+RDLK++NILLD++ PK+SDFG+A++ GD T R++G
Sbjct: 787 AARGLEYLHCKADPPVIYRDLKSANILLDSDFSPKLSDFGLAKLGPVGDNTHVST-RVMG 963
Query: 628 TYGYMAPEYVIHGLFSIKSDVFSFGVLLLETISGKKNRTLTYHEHDHNLI 677
TYGY APEY + G ++KSD++SFGV+LLE I+G++ + + NL+
Sbjct: 964 TYGYCAPEYAMSGKLTLKSDIYSFGVVLLELITGRRAIDASKKPGEQNLV 1113
>BE320157 weakly similar to PIR|T04835|T04 probable serine/threonine-specific
protein kinase (EC 2.7.1.-) F21P8.70 - Arabidopsis
thaliana, partial (27%)
Length = 529
Score = 206 bits (523), Expect = 3e-53
Identities = 101/167 (60%), Positives = 129/167 (76%)
Frame = +1
Query: 451 FDLATIIKATDNFSTNNKLGEGGFGPVYKATLQDGHVIAVKRLSGNSEQGSKEFKNEVIL 510
F +TI AT+ FS+ N++G+GGFG VYK L DG IAVK+LS +S QGS EF+NE++L
Sbjct: 22 FKFSTIEAATNKFSSENEIGKGGFGIVYKGVLSDGQQIAVKKLSRSSGQGSIEFQNEILL 201
Query: 511 CVKLQHRNLVKVLGCCIEGDEKLLIYEYMPNKSLDSFLFDPTQSKLLSWSMRLNILNAIA 570
KLQHRNLV +LG C+E EK+LIYEY+PNKSLD FLFD + ++L W R I+ IA
Sbjct: 202 IAKLQHRNLVTLLGFCLEEREKMLIYEYVPNKSLDYFLFDSKKHRVLHWFERYKIIGGIA 381
Query: 571 RGIQYLHQDSRLRIIHRDLKASNILLDNEMDPKISDFGMARMCGGDQ 617
RGI YLH+ SRL++IHRDLK SN+LLD++M+PKISDFG+AR+ DQ
Sbjct: 382 RGILYLHEYSRLKVIHRDLKPSNVLLDDKMNPKISDFGLARIVAIDQ 522
>AW560447 similar to PIR|T14520|T14 probable S-receptor kinase (EC 2.7.1.-)
SFR3 precursor - wild cabbage, partial (12%)
Length = 641
Score = 179 bits (455), Expect(2) = 5e-53
Identities = 91/162 (56%), Positives = 117/162 (72%), Gaps = 10/162 (6%)
Frame = +1
Query: 32 ITKSASLSNGSTLVSKDGTFEMGFFRPGKSLNRYVGIWYKNIPVRRVVWVANRNNPTKDD 91
I ++ +G+TLVS DGTFE+GFF PG S NRYVGIWYKNIP RR+VWVANR+NP KD+
Sbjct: 40 IYPNSIFDDGNTLVSNDGTFELGFFTPGSSTNRYVGIWYKNIPKRRIVWVANRDNPIKDN 219
Query: 92 SSK---LIISQDGNL-VLLNHNDSLVWSTNASRKA----SSPVVQLLNNGNLVLRDEKDN 143
+S LI+S DGNL +L N+N +LVWSTN + ++ SS V QLL+NGN V++ +
Sbjct: 220 TSNSTMLIMSNDGNLEILTNNNQTLVWSTNITTQSLSTTSSHVAQLLDNGNFVIKANNNT 399
Query: 144 NEES--FLWQGFDHPCDTLLPGMTFGYNRKLDFYWNLTAWKN 183
+++S FLWQGFD PCDTLLP M G++ K LT+WKN
Sbjct: 400 DQQSNNFLWQGFDFPCDTLLPDMKLGWDLKTGLNRQLTSWKN 525
Score = 47.4 bits (111), Expect(2) = 5e-53
Identities = 21/36 (58%), Positives = 26/36 (71%)
Frame = +3
Query: 185 DDPSSGDLYASVVFTSNPESMIWKGSTKICRSGPWN 220
DDPSSGD ++V SNPE ++ KGS +I SGPWN
Sbjct: 528 DDPSSGDFTWAIVLRSNPEIVLKKGSVEIHMSGPWN 635
>BE942940 weakly similar to GP|13506749|gb receptor-like protein kinase 6
{Arabidopsis thaliana}, partial (23%)
Length = 466
Score = 204 bits (520), Expect = 7e-53
Identities = 101/151 (66%), Positives = 118/151 (77%)
Frame = +1
Query: 514 LQHRNLVKVLGCCIEGDEKLLIYEYMPNKSLDSFLFDPTQSKLLSWSMRLNILNAIARGI 573
LQHRNLV +G +EG EK+LIYEY+ NKSLD FLFD Q KLL+W R NI+ IARGI
Sbjct: 1 LQHRNLVTFIGFWVEGHEKILIYEYVSNKSLDYFLFDSHQQKLLTWVERFNIIGGIARGI 180
Query: 574 QYLHQDSRLRIIHRDLKASNILLDNEMDPKISDFGMARMCGGDQIEGKTRRIVGTYGYMA 633
YLH+ SRL++IHRDLK SNILLD M PKISDFG+ARM Q EG T RIVGTYGYM+
Sbjct: 181 LYLHEHSRLKVIHRDLKPSNILLDENMIPKISDFGLARMVEISQEEGSTNRIVGTYGYMS 360
Query: 634 PEYVIHGLFSIKSDVFSFGVLLLETISGKKN 664
PEY + G FS KSDV+SFGV++LE ++GKKN
Sbjct: 361 PEYAMFGQFSEKSDVYSFGVMILEIVAGKKN 453
>TC78270 similar to GP|16649103|gb|AAL24403.1 Unknown protein {Arabidopsis
thaliana}, partial (80%)
Length = 1463
Score = 203 bits (516), Expect = 2e-52
Identities = 99/207 (47%), Positives = 138/207 (65%), Gaps = 1/207 (0%)
Frame = +3
Query: 459 ATDNFSTNNKLGEGGFGPVYKATLQDGHVIAVKRLSGNSEQGSKEFKNEVILCVKLQHRN 518
A+DNFS NK+GEGGFG VYK L+ G + A+K LS S+QG KEF E+ + +++H N
Sbjct: 297 ASDNFSPANKIGEGGFGSVYKGVLKGGKLAAIKVLSTESKQGVKEFLTEINVISEIKHEN 476
Query: 519 LVKVLGCCIEGDEKLLIYEYMPNKSLDSFLFDPTQSKL-LSWSMRLNILNAIARGIQYLH 577
LV + GCC+EGD ++L+Y Y+ N SL L S + W R I +ARG+ +LH
Sbjct: 477 LVILYGCCVEGDHRILVYNYLENNSLSQTLLAGGHSNIYFDWQTRRRICLGVARGLAFLH 656
Query: 578 QDSRLRIIHRDLKASNILLDNEMDPKISDFGMARMCGGDQIEGKTRRIVGTYGYMAPEYV 637
++ I+HRD+KASNILLD ++ PKISDFG+A++ T R+ GT GY+APEY
Sbjct: 657 EEVLPHIVHRDIKASNILLDKDLTPKISDFGLAKLIPSYMTHVST-RVAGTIGYLAPEYA 833
Query: 638 IHGLFSIKSDVFSFGVLLLETISGKKN 664
I G + K+D++SFGVLL+E +SG+ N
Sbjct: 834 IRGQLTRKADIYSFGVLLVEIVSGRSN 914
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.319 0.137 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 24,457,818
Number of Sequences: 36976
Number of extensions: 393775
Number of successful extensions: 3338
Number of sequences better than 10.0: 696
Number of HSP's better than 10.0 without gapping: 2685
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2746
length of query: 685
length of database: 9,014,727
effective HSP length: 103
effective length of query: 582
effective length of database: 5,206,199
effective search space: 3030007818
effective search space used: 3030007818
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 62 (28.5 bits)
Medicago: description of AC146866.11


