

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146864.6 - phase: 0 /pseudo
(180 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
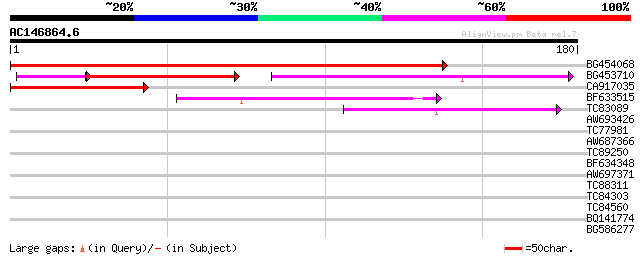
Score E
Sequences producing significant alignments: (bits) Value
BG454068 weakly similar to GP|22202804|dbj hypothetical protein~... 272 6e-74
BG453710 weakly similar to GP|21537057|gb| unknown {Arabidopsis ... 70 1e-25
CA917035 similar to GP|7303305|gb|A CG6061-PA {Drosophila melano... 87 3e-18
BF633515 52 1e-07
TC83089 46 9e-06
AW693426 homologue to GP|23307580|dbj similar to serine/threonin... 39 0.001
TC77981 similar to PIR|T12970|T12970 hypothetical protein T6H20.... 29 0.88
AW687366 similar to GP|6045135|dbj| oxidosqualene cyclase {Luffa... 28 1.5
TC89250 weakly similar to GP|21592651|gb|AAM64600.1 rhodanese-li... 28 2.6
BF634348 27 4.4
AW697371 27 4.4
TC88311 similar to GP|11994408|dbj|BAB02410. CTP synthase {Arabi... 27 5.7
TC84303 weakly similar to GP|3540182|gb|AAC34332.1| Unknown prot... 27 5.7
TC84560 similar to GP|17978924|gb|AAL47429.1 AT5g45290/K9E15_7 {... 26 7.5
BQ141774 similar to GP|19915999|gb| transposase {Methanosarcina ... 26 9.8
BG586277 26 9.8
>BG454068 weakly similar to GP|22202804|dbj hypothetical protein~predicted by
GeneMark.hmm etc. {Oryza sativa (japonica
cultivar-group)}, partial (5%)
Length = 673
Score = 272 bits (695), Expect = 6e-74
Identities = 130/139 (93%), Positives = 134/139 (95%)
Frame = +3
Query: 1 MGWFWNVIRASGLYLLLETNYGQVDHGLLIAFSERWHSETSSFHLPVGEMTITLDDVSCL 60
MGWFWNVIRASGLY LLETNYGQVDHGLLIAFS RWHSETSSFHLPVGEMTI LDD+SCL
Sbjct: 255 MGWFWNVIRASGLYPLLETNYGQVDHGLLIAFSLRWHSETSSFHLPVGEMTIALDDISCL 434
Query: 61 LHIPVGGNLLFHESLSIHQGTQYLVNYLGLEFEESAAQTKRLRSAHITYDTLLSIYTSYL 120
LHIPVGGNLLFHESLSIHQGT+YLVNYLGLEFEE AA+TKRL+SAHITYDTLLSIYTSYL
Sbjct: 435 LHIPVGGNLLFHESLSIHQGTEYLVNYLGLEFEEIAAETKRLKSAHITYDTLLSIYTSYL 614
Query: 121 TEAKSYANQPEEEDSMEWY 139
TEAKSYANQP EEDSMEWY
Sbjct: 615 TEAKSYANQPGEEDSMEWY 671
>BG453710 weakly similar to GP|21537057|gb| unknown {Arabidopsis thaliana},
partial (6%)
Length = 651
Score = 70.5 bits (171), Expect(3) = 1e-25
Identities = 32/49 (65%), Positives = 39/49 (79%)
Frame = -1
Query: 25 DHGLLIAFSERWHSETSSFHLPVGEMTITLDDVSCLLHIPVGGNLLFHE 73
D ++ A ERWH+ETSSFHLP+GEMTITLDDV LLHIP+ G +L H+
Sbjct: 474 DPSMIRAMCERWHTETSSFHLPMGEMTITLDDVYNLLHIPIHGCMLDHD 328
Score = 56.6 bits (135), Expect(3) = 1e-25
Identities = 31/98 (31%), Positives = 49/98 (49%), Gaps = 2/98 (2%)
Frame = -3
Query: 84 LVNYLGLEFEESAAQTKRLRSAHITYDTLLSIYTSYLTEAKSYANQPEEEDSMEWYRTR- 142
+ LG+ + A+ + + HI+Y TL +Y +L EA+ + E+ E R R
Sbjct: 295 MTRLLGMSDANAMAEIRTESAGHISYPTLKRVYEDHLIEARRLEDPLTREELQERARRRQ 116
Query: 143 -CIRAFLLYLVGCTLFSDKAGNSCCVVYLKYFDDLTTV 179
C+R+ LLYLV C LF+ K ++YL DL +
Sbjct: 115 WCVRSLLLYLVDCVLFTYKTNRHIDLIYLDCMADLQAI 2
Score = 25.4 bits (54), Expect(3) = 1e-25
Identities = 10/24 (41%), Positives = 14/24 (57%)
Frame = -2
Query: 3 WFWNVIRASGLYLLLETNYGQVDH 26
WF ++ SGL+ L+ T Y V H
Sbjct: 539 WFLELVEGSGLHDLIYTGYFTVTH 468
>CA917035 similar to GP|7303305|gb|A CG6061-PA {Drosophila melanogaster},
partial (2%)
Length = 365
Score = 87.4 bits (215), Expect = 3e-18
Identities = 40/44 (90%), Positives = 41/44 (92%)
Frame = +2
Query: 1 MGWFWNVIRASGLYLLLETNYGQVDHGLLIAFSERWHSETSSFH 44
MG FWNVI ASGLYLLLETNYGQVDH LLIAFS+RWHSETSSFH
Sbjct: 233 MGRFWNVIMASGLYLLLETNYGQVDHSLLIAFSDRWHSETSSFH 364
>BF633515
Length = 249
Score = 52.4 bits (124), Expect = 1e-07
Identities = 29/85 (34%), Positives = 47/85 (55%), Gaps = 1/85 (1%)
Frame = +2
Query: 54 LDDVSCLLHIPVGGNLLFH-ESLSIHQGTQYLVNYLGLEFEESAAQTKRLRSAHITYDTL 112
LDDV+CL H+P+ G +L H + + H+G L+ +LG+ +E+ + +I+Y L
Sbjct: 2 LDDVACLTHLPIEGRMLSHGKKMPKHEGAALLMRHLGVSQQEAEKICGQEYGGYISYPRL 181
Query: 113 LSIYTSYLTEAKSYANQPEEEDSME 137
YT+YL A A+ ED +E
Sbjct: 182 RDFYTTYLGRANLLAS---TEDPVE 247
>TC83089
Length = 887
Score = 45.8 bits (107), Expect = 9e-06
Identities = 26/71 (36%), Positives = 35/71 (48%), Gaps = 2/71 (2%)
Frame = +3
Query: 107 ITYDTLLSIYTSYLTEAKSYANQPEEED--SMEWYRTRCIRAFLLYLVGCTLFSDKAGNS 164
I L Y YL A ++ D +E RT C++ +LLYLVGC LF DK+
Sbjct: 249 ILSQALREYYEGYLDSATRLSDPQNLGDPQELEKIRTTCVKWYLLYLVGCLLFGDKSNKR 428
Query: 165 CCVVYLKYFDD 175
+VYL +D
Sbjct: 429 IELVYLTTMED 461
>AW693426 homologue to GP|23307580|dbj similar to serine/threonine
phosphatase PP7 {Oryza sativa (japonica
cultivar-group)}, partial (2%)
Length = 209
Score = 38.5 bits (88), Expect = 0.001
Identities = 19/30 (63%), Positives = 22/30 (73%)
Frame = +1
Query: 147 FLLYLVGCTLFSDKAGNSCCVVYLKYFDDL 176
FLL LVGCTLFSDK+ V YL++F DL
Sbjct: 1 FLLCLVGCTLFSDKSTFVVNVAYLEFFRDL 90
>TC77981 similar to PIR|T12970|T12970 hypothetical protein T6H20.190 -
Arabidopsis thaliana, partial (39%)
Length = 1191
Score = 29.3 bits (64), Expect = 0.88
Identities = 14/36 (38%), Positives = 22/36 (60%)
Frame = -1
Query: 57 VSCLLHIPVGGNLLFHESLSIHQGTQYLVNYLGLEF 92
+SC +HIP+ ++ +SL + Q T+YL N L F
Sbjct: 1140 LSCAVHIPLXLVIINIDSLLLEQATKYLPNLLRFLF 1033
>AW687366 similar to GP|6045135|dbj| oxidosqualene cyclase {Luffa
cylindrica}, partial (3%)
Length = 345
Score = 28.5 bits (62), Expect = 1.5
Identities = 14/36 (38%), Positives = 19/36 (51%), Gaps = 5/36 (13%)
Frame = -2
Query: 43 FHLPVGEMTI-----TLDDVSCLLHIPVGGNLLFHE 73
FHLPV + + TL C HIP+ L+FH+
Sbjct: 344 FHLPVPRILLSE*CETLYRC*CAFHIPIDNGLIFHK 237
>TC89250 weakly similar to GP|21592651|gb|AAM64600.1 rhodanese-like family
protein {Arabidopsis thaliana}, partial (78%)
Length = 1039
Score = 27.7 bits (60), Expect = 2.6
Identities = 13/51 (25%), Positives = 25/51 (48%)
Frame = -3
Query: 28 LLIAFSERWHSETSSFHLPVGEMTITLDDVSCLLHIPVGGNLLFHESLSIH 78
+++ F + WH + + FH+ CLL P+G N+ +E+ +H
Sbjct: 326 IILIFHKHWHMQRT-FHM-------------CLLSFPIGSNIKKNETF*VH 216
>BF634348
Length = 337
Score = 26.9 bits (58), Expect = 4.4
Identities = 13/29 (44%), Positives = 19/29 (64%)
Frame = +3
Query: 110 DTLLSIYTSYLTEAKSYANQPEEEDSMEW 138
+T +S+ +YL SYAN+ E+ SMEW
Sbjct: 198 NT*MSLIVNYL----SYANKQEQXSSMEW 272
>AW697371
Length = 644
Score = 26.9 bits (58), Expect = 4.4
Identities = 12/30 (40%), Positives = 16/30 (53%)
Frame = -3
Query: 151 LVGCTLFSDKAGNSCCVVYLKYFDDLTTVN 180
L G L D + SCC +L +F D T+N
Sbjct: 591 LKGRRLSFDGSSMSCCFFFLNHFPDRMTIN 502
>TC88311 similar to GP|11994408|dbj|BAB02410. CTP synthase {Arabidopsis
thaliana}, partial (53%)
Length = 1190
Score = 26.6 bits (57), Expect = 5.7
Identities = 25/99 (25%), Positives = 43/99 (43%), Gaps = 2/99 (2%)
Frame = +3
Query: 30 IAFSERWHSETSSFHLPVGEMTITLDDVSCLLHIPVGGNLLFHESLSIHQGTQYLVNYLG 89
+ E ++ S F L GE ITL DV + HIP L H+ ++N G
Sbjct: 879 MVLDENAKAKLSQFCLLPGENIITLYDVPNIWHIP-----LLLRDQKAHEAIFKVLNIKG 1043
Query: 90 LEFEESAAQ-TKRLRSAHITYDTL-LSIYTSYLTEAKSY 126
+ E + + T R S + ++ + +++ Y + SY
Sbjct: 1044MTQEPNLEEWTCRAESCDLLHEPVRIALVGKYTCLSDSY 1160
>TC84303 weakly similar to GP|3540182|gb|AAC34332.1| Unknown protein
{Arabidopsis thaliana}, partial (14%)
Length = 684
Score = 26.6 bits (57), Expect = 5.7
Identities = 22/97 (22%), Positives = 42/97 (42%), Gaps = 6/97 (6%)
Frame = +2
Query: 53 TLDDVSCLLHIPVGGNLLFHESLSIHQGTQYLVNYLGLEFEESAAQTKRLRSAHITYDTL 112
T+DD+S +++ VGG + E I TQ + + F++S + + + T+
Sbjct: 50 TVDDISNKVYVNVGGQKIDMEEEMIRGLTQVHWERVDVSFQKS-------KQRYTAHSTI 208
Query: 113 LSIYTSYLTEAK------SYANQPEEEDSMEWYRTRC 143
+ SY AK +Y P ++ Y+ +C
Sbjct: 209 QVKFLSYSFPAKLVDIILNYFITPALDEGHLLYKNKC 319
>TC84560 similar to GP|17978924|gb|AAL47429.1 AT5g45290/K9E15_7 {Arabidopsis
thaliana}, partial (12%)
Length = 1044
Score = 26.2 bits (56), Expect = 7.5
Identities = 13/33 (39%), Positives = 20/33 (60%)
Frame = -2
Query: 41 SSFHLPVGEMTITLDDVSCLLHIPVGGNLLFHE 73
S++HLP G M ++L + LHI NL+F +
Sbjct: 989 SNYHLPKGRMDLSLMVIQHTLHI----NLIFQD 903
>BQ141774 similar to GP|19915999|gb| transposase {Methanosarcina acetivorans
str. C2A} [Methanosarcina acetivorans C2A], partial (3%)
Length = 986
Score = 25.8 bits (55), Expect = 9.8
Identities = 10/24 (41%), Positives = 14/24 (57%)
Frame = -1
Query: 133 EDSMEWYRTRCIRAFLLYLVGCTL 156
E+ + W R RCI Y+VGC +
Sbjct: 842 ENGIVWERGRCIMGRRRYVVGCAV 771
>BG586277
Length = 748
Score = 25.8 bits (55), Expect = 9.8
Identities = 10/29 (34%), Positives = 17/29 (58%)
Frame = -2
Query: 145 RAFLLYLVGCTLFSDKAGNSCCVVYLKYF 173
+ F LY++ L S + +SC ++ KYF
Sbjct: 663 KLFALYVICSLLESKRVSSSCSTLHYKYF 577
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.322 0.137 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,514,371
Number of Sequences: 36976
Number of extensions: 91212
Number of successful extensions: 438
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 437
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 437
length of query: 180
length of database: 9,014,727
effective HSP length: 90
effective length of query: 90
effective length of database: 5,686,887
effective search space: 511819830
effective search space used: 511819830
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 55 (25.8 bits)
Medicago: description of AC146864.6


