

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146863.4 + phase: 0
(309 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
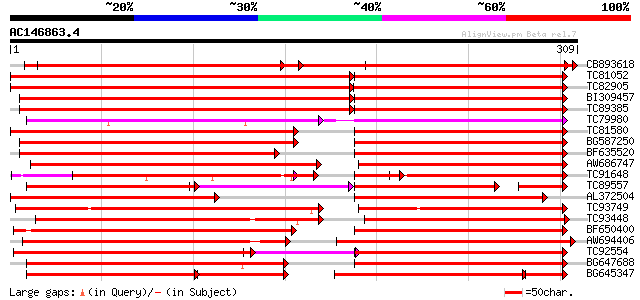
Score E
Sequences producing significant alignments: (bits) Value
CB893618 weakly similar to GP|3947733|emb| NL25 {Solanum tuberos... 309 e-168
TC81052 weakly similar to GP|10178211|dbj|BAB11635. TMV resistan... 247 5e-66
TC82905 weakly similar to GP|3947733|emb|CAA08797.1 NL25 {Solanu... 207 5e-54
BI309457 weakly similar to GP|15787901|gb resistance gene analog... 204 3e-53
TC89385 similar to GP|19774173|gb|AAL99063.1 NBS-LRR-Toll resist... 201 2e-52
TC79980 weakly similar to GP|15787901|gb|AAL07542.1 resistance g... 190 7e-49
TC81580 weakly similar to GP|7341113|gb|AAF61210.1| unknown {Gly... 187 6e-48
BG587250 weakly similar to GP|3947735|emb| NL27 {Solanum tuberos... 181 4e-46
BF635520 weakly similar to GP|3947735|emb| NL27 {Solanum tuberos... 159 1e-39
AW686747 weakly similar to PIR|A54810|A5 TMV resistance protein ... 156 1e-38
TC91648 weakly similar to GP|23477201|emb|CAD36199. NLS-TIR-NBS ... 134 2e-38
TC89557 weakly similar to GP|10178211|dbj|BAB11635. TMV resistan... 104 3e-38
AL372504 weakly similar to GP|7341113|gb| unknown {Glycine max},... 151 3e-37
TC93749 similar to GP|3947733|emb|CAA08797.1 NL25 {Solanum tuber... 147 4e-36
TC93448 similar to GP|9858478|gb|AAG01052.1| resistance protein ... 144 4e-35
BF650400 weakly similar to GP|9965109|gb| resistance protein MG1... 142 1e-34
AW694406 weakly similar to GP|3947735|emb NL27 {Solanum tuberosu... 139 2e-33
TC92554 similar to GP|9758876|dbj|BAB09430.1 disease resistance ... 134 3e-33
BG647688 similar to GP|9965109|gb| resistance protein MG13 {Glyc... 136 1e-32
BG645347 weakly similar to GP|18033111|g functional candidate re... 103 2e-32
>CB893618 weakly similar to GP|3947733|emb| NL25 {Solanum tuberosum}, partial
(18%)
Length = 927
Score = 309 bits (792), Expect(2) = e-168
Identities = 149/151 (98%), Positives = 149/151 (98%)
Frame = +3
Query: 159 HHAELENIIEHINILGCKFPSRTKDLVEINYDVFVSFRGPDTRFNFTDHFCAALQRRGIN 218
H ELENIIEHINILGCKFPSRTKDLVEINYDVFVSFRGPDTRFNFTDHFCAALQRRGIN
Sbjct: 429 HIMELENIIEHINILGCKFPSRTKDLVEINYDVFVSFRGPDTRFNFTDHFCAALQRRGIN 608
Query: 219 AFRDDTKLKKGEFIAPGLFRAIEASQVYIVVFSKNYASSTWCLRELEYILHCSKKHGKHV 278
AFRDDTKLKKGEFIAPGLFRAIEASQVYIVVFSKNYASSTWCLRELEYILHCSKKHGKHV
Sbjct: 609 AFRDDTKLKKGEFIAPGLFRAIEASQVYIVVFSKNYASSTWCLRELEYILHCSKKHGKHV 788
Query: 279 LPVFYDVDPSEVQKQSGGYGDALSKHGLNMV 309
LPVFYDVDPSEVQKQSGGYGDALSKHGLNMV
Sbjct: 789 LPVFYDVDPSEVQKQSGGYGDALSKHGLNMV 881
Score = 300 bits (769), Expect(2) = e-168
Identities = 145/145 (100%), Positives = 145/145 (100%)
Frame = +2
Query: 16 FRGTDTRFNFTDHLFGALQRKRIFTFRDDTNLQKGNSIASDLIQAIEGSQVFIVVFSKNY 75
FRGTDTRFNFTDHLFGALQRKRIFTFRDDTNLQKGNSIASDLIQAIEGSQVFIVVFSKNY
Sbjct: 2 FRGTDTRFNFTDHLFGALQRKRIFTFRDDTNLQKGNSIASDLIQAIEGSQVFIVVFSKNY 181
Query: 76 ASSTWCLRELAYILNCSVLYGKRVLPVFYDVDLSEVRKQSGGYGESFNYHGKRFQDHSNM 135
ASSTWCLRELAYILNCSVLYGKRVLPVFYDVDLSEVRKQSGGYGESFNYHGKRFQDHSNM
Sbjct: 182 ASSTWCLRELAYILNCSVLYGKRVLPVFYDVDLSEVRKQSGGYGESFNYHGKRFQDHSNM 361
Query: 136 VQRRRETLQLVGNISGWDLRDKPHH 160
VQRRRETLQLVGNISGWDLRDKPHH
Sbjct: 362 VQRRRETLQLVGNISGWDLRDKPHH 436
Score = 184 bits (468), Expect = 3e-47
Identities = 97/142 (68%), Positives = 109/142 (76%)
Frame = +3
Query: 9 DHTVFVSFRGTDTRFNFTDHLFGALQRKRIFTFRDDTNLQKGNSIASDLIQAIEGSQVFI 68
++ VFVSFRG DTRFNFTDH ALQR+ I FRDDT L+KG IA L +AIE SQV+I
Sbjct: 516 NYDVFVSFRGPDTRFNFTDHFCAALQRRGINAFRDDTKLKKGEFIAPGLFRAIEASQVYI 695
Query: 69 VVFSKNYASSTWCLRELAYILNCSVLYGKRVLPVFYDVDLSEVRKQSGGYGESFNYHGKR 128
VVFSKNYASSTWCLREL YIL+CS +GK VLPVFYDVD SEV+KQSGGYG++ + HG
Sbjct: 696 VVFSKNYASSTWCLRELEYILHCSKKHGKHVLPVFYDVDPSEVQKQSGGYGDALSKHG-- 869
Query: 129 FQDHSNMVQRRRETLQLVGNIS 150
NMV R RETL VGNIS
Sbjct: 870 ----LNMV*RWRETLTQVGNIS 923
Score = 164 bits (414), Expect = 5e-41
Identities = 80/111 (72%), Positives = 90/111 (81%)
Frame = +2
Query: 195 FRGPDTRFNFTDHFCAALQRRGINAFRDDTKLKKGEFIAPGLFRAIEASQVYIVVFSKNY 254
FRG DTRFNFTDH ALQR+ I FRDDT L+KG IA L +AIE SQV+IVVFSKNY
Sbjct: 2 FRGTDTRFNFTDHLFGALQRKRIFTFRDDTNLQKGNSIASDLIQAIEGSQVFIVVFSKNY 181
Query: 255 ASSTWCLRELEYILHCSKKHGKHVLPVFYDVDPSEVQKQSGGYGDALSKHG 305
ASSTWCLREL YIL+CS +GK VLPVFYDVD SEV+KQSGGYG++ + HG
Sbjct: 182 ASSTWCLRELAYILNCSVLYGKRVLPVFYDVDLSEVRKQSGGYGESFNYHG 334
>TC81052 weakly similar to GP|10178211|dbj|BAB11635. TMV resistance protein
N {Arabidopsis thaliana}, partial (8%)
Length = 656
Score = 247 bits (630), Expect = 5e-66
Identities = 126/189 (66%), Positives = 147/189 (77%), Gaps = 1/189 (0%)
Frame = +2
Query: 1 MTSPRKKNDHTVFVSFRGTDTRFNFTDHLFGALQRKRIFTFRDDTNLQKGNSIASDLIQA 60
M PR+KN + VFV+FRG DTRFNF DHLF ALQRK IF FRDD NLQKG SI +LI+A
Sbjct: 83 MALPRRKNYYDVFVTFRGEDTRFNFIDHLFAALQRKGIFAFRDDANLQKGESIPPELIRA 262
Query: 61 IEGSQVFIVVFSKNYASSTWCLRELAYILNCSVLYGKRVLPVFYDVDLSEVRKQSGGYGE 120
IEGSQVFI V SKNY+SSTWCLREL +IL+CS + G+RVLPVFYDVD SEVR Q G YGE
Sbjct: 263 IEGSQVFIAVLSKNYSSSTWCLRELVHILDCSQVSGRRVLPVFYDVDPSEVRHQKGIYGE 442
Query: 121 SFNYHGKRFQDHSNMVQRRRETLQLVGNISGWDLRDKPHHAELENIIEHI-NILGCKFPS 179
+F+ H + FQ S++VQ RE L VGNISGWDLRDKP +AE++ I+E I NILG F S
Sbjct: 443 AFSKHEQTFQHDSHVVQSWREALTQVGNISGWDLRDKPQYAEIKKIVEEILNILGHNFSS 622
Query: 180 RTKDLVEIN 188
K+LV +N
Sbjct: 623 LPKELVGMN 649
Score = 170 bits (431), Expect = 6e-43
Identities = 83/116 (71%), Positives = 93/116 (79%)
Frame = +2
Query: 189 YDVFVSFRGPDTRFNFTDHFCAALQRRGINAFRDDTKLKKGEFIAPGLFRAIEASQVYIV 248
YDVFV+FRG DTRFNF DH AALQR+GI AFRDD L+KGE I P L RAIE SQV+I
Sbjct: 110 YDVFVTFRGEDTRFNFIDHLFAALQRKGIFAFRDDANLQKGESIPPELIRAIEGSQVFIA 289
Query: 249 VFSKNYASSTWCLRELEYILHCSKKHGKHVLPVFYDVDPSEVQKQSGGYGDALSKH 304
V SKNY+SSTWCLREL +IL CS+ G+ VLPVFYDVDPSEV+ Q G YG+A SKH
Sbjct: 290 VLSKNYSSSTWCLRELVHILDCSQVSGRRVLPVFYDVDPSEVRHQKGIYGEAFSKH 457
>TC82905 weakly similar to GP|3947733|emb|CAA08797.1 NL25 {Solanum
tuberosum}, partial (22%)
Length = 832
Score = 207 bits (526), Expect = 5e-54
Identities = 110/189 (58%), Positives = 133/189 (70%), Gaps = 1/189 (0%)
Frame = +1
Query: 1 MTSPRKKNDHTVFVSFRGTDTRFNFTDHLFGALQRKRIFTFRDDTNLQKGNSIASDLIQA 60
+ + KKN + VFV+FRG DTR NFTD LF AL+ K I FRD NLQKG I +L +A
Sbjct: 61 LVTSSKKNHYDVFVTFRGEDTRNNFTDFLFDALETKGIMVFRDVINLQKGECIGPELFRA 240
Query: 61 IEGSQVFIVVFSKNYASSTWCLRELAYILNCSVLYGKRVLPVFYDVDLSEVRKQSGGYGE 120
IE SQV++ +FSKNYASSTWCL+EL I C GK VLPVFYDVD SEVRKQSG Y E
Sbjct: 241 IEISQVYVAIFSKNYASSTWCLQELEKICECIKGSGKHVLPVFYDVDPSEVRKQSGIYSE 420
Query: 121 SFNYHGKRFQDHSNMVQRRRETLQLVGNISGWDLRDKPHHAELENIIEH-INILGCKFPS 179
+F H +RFQ S V R RE L+ VG+ISGWDLRD+P E++ I++ INIL CK+
Sbjct: 421 AFVKHEQRFQQDSMKVSRWREALEQVGSISGWDLRDEPLAREIKEIVQKIINILECKYSC 600
Query: 180 RTKDLVEIN 188
+KDLV I+
Sbjct: 601 VSKDLVGID 627
Score = 162 bits (411), Expect = 1e-40
Identities = 80/117 (68%), Positives = 90/117 (76%)
Frame = +1
Query: 188 NYDVFVSFRGPDTRFNFTDHFCAALQRRGINAFRDDTKLKKGEFIAPGLFRAIEASQVYI 247
+YDVFV+FRG DTR NFTD AL+ +GI FRD L+KGE I P LFRAIE SQVY+
Sbjct: 85 HYDVFVTFRGEDTRNNFTDFLFDALETKGIMVFRDVINLQKGECIGPELFRAIEISQVYV 264
Query: 248 VVFSKNYASSTWCLRELEYILHCSKKHGKHVLPVFYDVDPSEVQKQSGGYGDALSKH 304
+FSKNYASSTWCL+ELE I C K GKHVLPVFYDVDPSEV+KQSG Y +A KH
Sbjct: 265 AIFSKNYASSTWCLQELEKICECIKGSGKHVLPVFYDVDPSEVRKQSGIYSEAFVKH 435
>BI309457 weakly similar to GP|15787901|gb resistance gene analog NBS7
{Helianthus annuus}, partial (45%)
Length = 806
Score = 204 bits (520), Expect = 3e-53
Identities = 109/184 (59%), Positives = 131/184 (70%), Gaps = 1/184 (0%)
Frame = +3
Query: 6 KKNDHTVFVSFRGTDTRFNFTDHLFGALQRKRIFTFRDDTNLQKGNSIASDLIQAIEGSQ 65
++N + VFV+FRG DTR NFTD LF ALQ K I F DDTNL KG SI +L++AIEGSQ
Sbjct: 60 RRNYYDVFVTFRGEDTRNNFTDFLFDALQTKGIIVFSDDTNLPKGESIGPELLRAIEGSQ 239
Query: 66 VFIVVFSKNYASSTWCLRELAYILNCSVLYGKRVLPVFYDVDLSEVRKQSGGYGESFNYH 125
VF+ VFS NYASSTWCL+EL I C GK VLPVFYDVD S+VRKQSG YGE+F H
Sbjct: 240 VFVAVFSINYASSTWCLQELEKICECVKGSGKHVLPVFYDVDPSDVRKQSGIYGEAFIKH 419
Query: 126 GKRFQDHSNMVQRRRETLQLVGNISGWDLRDKPHHAELENIIEHI-NILGCKFPSRTKDL 184
+RFQ V + R+ L+ VG+ISGWDLRDKP E++ I++ I NIL K +KDL
Sbjct: 420 EQRFQQEFQKVSKWRDALKQVGSISGWDLRDKPQAGEIKKIVQTILNILKYKSSCFSKDL 599
Query: 185 VEIN 188
V I+
Sbjct: 600 VGID 611
Score = 160 bits (406), Expect = 4e-40
Identities = 80/116 (68%), Positives = 88/116 (74%)
Frame = +3
Query: 189 YDVFVSFRGPDTRFNFTDHFCAALQRRGINAFRDDTKLKKGEFIAPGLFRAIEASQVYIV 248
YDVFV+FRG DTR NFTD ALQ +GI F DDT L KGE I P L RAIE SQV++
Sbjct: 72 YDVFVTFRGEDTRNNFTDFLFDALQTKGIIVFSDDTNLPKGESIGPELLRAIEGSQVFVA 251
Query: 249 VFSKNYASSTWCLRELEYILHCSKKHGKHVLPVFYDVDPSEVQKQSGGYGDALSKH 304
VFS NYASSTWCL+ELE I C K GKHVLPVFYDVDPS+V+KQSG YG+A KH
Sbjct: 252 VFSINYASSTWCLQELEKICECVKGSGKHVLPVFYDVDPSDVRKQSGIYGEAFIKH 419
>TC89385 similar to GP|19774173|gb|AAL99063.1 NBS-LRR-Toll resistance gene
analog protein {Medicago ruthenica}, partial (98%)
Length = 1285
Score = 201 bits (512), Expect = 2e-52
Identities = 105/184 (57%), Positives = 130/184 (70%), Gaps = 1/184 (0%)
Frame = +3
Query: 6 KKNDHTVFVSFRGTDTRFNFTDHLFGALQRKRIFTFRDDTNLQKGNSIASDLIQAIEGSQ 65
++N + VFV+FRG DTR NFT+ LF AL+RK I+ FRDDTNL KG SI +L++ IEGSQ
Sbjct: 57 RRNCYDVFVTFRGEDTRNNFTNFLFAALERKGIYAFRDDTNLPKGESIGPELLRTIEGSQ 236
Query: 66 VFIVVFSKNYASSTWCLRELAYILNCSVLYGKRVLPVFYDVDLSEVRKQSGGYGESFNYH 125
VF+ V S+NYASSTWCL+EL I C GK VLP+FY VD SEV+KQSG Y + F H
Sbjct: 237 VFVAVLSRNYASSTWCLQELEKICECIKGSGKYVLPIFYGVDPSEVKKQSGIYWDDFAKH 416
Query: 126 GKRFQDHSNMVQRRRETLQLVGNISGWDLRDKPHHAELENIIEHI-NILGCKFPSRTKDL 184
+RF+ + V R RE L VG+I+GWDLRDK E+E I++ I NIL CK +KDL
Sbjct: 417 EQRFKQDPHKVSRWREALNQVGSIAGWDLRDKQQSVEVEKIVQTILNILKCKSSFVSKDL 596
Query: 185 VEIN 188
V IN
Sbjct: 597 VGIN 608
Score = 157 bits (398), Expect = 4e-39
Identities = 77/116 (66%), Positives = 89/116 (76%)
Frame = +3
Query: 189 YDVFVSFRGPDTRFNFTDHFCAALQRRGINAFRDDTKLKKGEFIAPGLFRAIEASQVYIV 248
YDVFV+FRG DTR NFT+ AAL+R+GI AFRDDT L KGE I P L R IE SQV++
Sbjct: 69 YDVFVTFRGEDTRNNFTNFLFAALERKGIYAFRDDTNLPKGESIGPELLRTIEGSQVFVA 248
Query: 249 VFSKNYASSTWCLRELEYILHCSKKHGKHVLPVFYDVDPSEVQKQSGGYGDALSKH 304
V S+NYASSTWCL+ELE I C K GK+VLP+FY VDPSEV+KQSG Y D +KH
Sbjct: 249 VLSRNYASSTWCLQELEKICECIKGSGKYVLPIFYGVDPSEVKKQSGIYWDDFAKH 416
>TC79980 weakly similar to GP|15787901|gb|AAL07542.1 resistance gene analog
NBS7 {Helianthus annuus}, partial (33%)
Length = 1258
Score = 190 bits (482), Expect = 7e-49
Identities = 113/301 (37%), Positives = 169/301 (55%), Gaps = 6/301 (1%)
Frame = +3
Query: 10 HTVFVSFRGTDTRFNFTDHLFGALQRKRIFTFRDDTNLQKGNS---IASDLIQAIEGSQV 66
+ VF+SFRG DT F +L+ AL+ K+I TF +Q + ++ +++AI+ S++
Sbjct: 60 YDVFLSFRGEDTYCTFAGNLYHALRNKKIKTFFPHDQIQNDDEELQLSPSILKAIQESRI 239
Query: 67 FIVVFSKNYASSTWCLRELAYILNCSVLYGKRVLPVFYDVDLSEVRKQSGGYGESFNYHG 126
IVV SKNYA+ST CL EL IL C + + V P+FY+V S+V+ Q YG S
Sbjct: 240 SIVVLSKNYATSTRCLNELVIILQCMKMKNQLVWPIFYEVHSSDVKLQRCKYGSSSKAIL 419
Query: 127 K---RFQDHSNMVQRRRETLQLVGNISGWDLRDKPHHAELENIIEHINILGCKFPSRTKD 183
K RF+D+ + ++ L V +I+GW+ K + ++ I+E + P
Sbjct: 420 KFRERFKDYPRRMWEWQQALSQVTSIAGWNYGIKFEYELIQKIVE---LTVQSLP----- 575
Query: 184 LVEINYDVFVSFRGPDTRFNFTDHFCAALQRRGINAFRDDTKLKKGEFIAPGLFRAIEAS 243
YDVF+SF G DTR++FT AL+ G F DD L+ G I+ L +AIE S
Sbjct: 576 ----RYDVFLSFCGEDTRYSFTGFLYHALRLEGFKIFMDDEGLEGGNQISQTLLKAIEKS 743
Query: 244 QVYIVVFSKNYASSTWCLRELEYILHCSKKHGKHVLPVFYDVDPSEVQKQSGGYGDALSK 303
++ IVV S+NY STWCL EL I+ C K + K V P+FY ++ S++ + YG A++
Sbjct: 744 RLSIVVLSENYGYSTWCLDELVKIMECKKTNNKLVWPLFYKIEQSDLSYKKSSYGKAMAA 923
Query: 304 H 304
H
Sbjct: 924 H 926
Score = 115 bits (289), Expect = 2e-26
Identities = 62/163 (38%), Positives = 93/163 (57%), Gaps = 1/163 (0%)
Frame = +3
Query: 10 HTVFVSFRGTDTRFNFTDHLFGALQRKRIFTFRDDTNLQKGNSIASDLIQAIEGSQVFIV 69
+ VF+SF G DTR++FT L+ AL+ + F DD L+ GN I+ L++AIE S++ IV
Sbjct: 579 YDVFLSFCGEDTRYSFTGFLYHALRLEGFKIFMDDEGLEGGNQISQTLLKAIEKSRLSIV 758
Query: 70 VFSKNYASSTWCLRELAYILNCSVLYGKRVLPVFYDVDLSEVRKQSGGYGESFNYHGKRF 129
V S+NY STWCL EL I+ C K V P+FY ++ S++ + YG++ H RF
Sbjct: 759 VLSENYGYSTWCLDELVKIMECKKTNNKLVWPLFYKIEQSDLSYKKSSYGKAMAAHEDRF 938
Query: 130 QDHSNMVQRRRETLQLVGNISGWDLRDKPHHAE-LENIIEHIN 171
S VQ+ R L V + +++ H E ++ I+E N
Sbjct: 939 GKESENVQKWRSALSEVALLKADHIKENEHEYEFIKKIVERAN 1067
>TC81580 weakly similar to GP|7341113|gb|AAF61210.1| unknown {Glycine max},
partial (84%)
Length = 827
Score = 187 bits (474), Expect = 6e-48
Identities = 96/157 (61%), Positives = 114/157 (72%)
Frame = +2
Query: 1 MTSPRKKNDHTVFVSFRGTDTRFNFTDHLFGALQRKRIFTFRDDTNLQKGNSIASDLIQA 60
+ + + N + VF +FRG DTR NFTD LF AL+ K IF FRDDTNLQKG SI +L++A
Sbjct: 86 LVTSSRNNYYDVFGTFRGEDTRNNFTDFLFDALETKGIFAFRDDTNLQKGESIEPELLRA 265
Query: 61 IEGSQVFIVVFSKNYASSTWCLRELAYILNCSVLYGKRVLPVFYDVDLSEVRKQSGGYGE 120
IEGS+VF+ VFS+NYASSTWCL+EL I C K +LPVFYDVD S VRKQSG Y E
Sbjct: 266 IEGSRVFVAVFSRNYASSTWCLQELEKICKCVQRSRKHILPVFYDVDPSVVRKQSGIYCE 445
Query: 121 SFNYHGKRFQDHSNMVQRRRETLQLVGNISGWDLRDK 157
+F H +RFQ MV R RE L+ VG+ISGWDLRDK
Sbjct: 446 AFVKHEQRFQQDFEMVSRWREALKHVGSISGWDLRDK 556
Score = 152 bits (385), Expect = 1e-37
Identities = 75/116 (64%), Positives = 89/116 (76%)
Frame = +2
Query: 189 YDVFVSFRGPDTRFNFTDHFCAALQRRGINAFRDDTKLKKGEFIAPGLFRAIEASQVYIV 248
YDVF +FRG DTR NFTD AL+ +GI AFRDDT L+KGE I P L RAIE S+V++
Sbjct: 113 YDVFGTFRGEDTRNNFTDFLFDALETKGIFAFRDDTNLQKGESIEPELLRAIEGSRVFVA 292
Query: 249 VFSKNYASSTWCLRELEYILHCSKKHGKHVLPVFYDVDPSEVQKQSGGYGDALSKH 304
VFS+NYASSTWCL+ELE I C ++ KH+LPVFYDVDPS V+KQSG Y +A KH
Sbjct: 293 VFSRNYASSTWCLQELEKICKCVQRSRKHILPVFYDVDPSVVRKQSGIYCEAFVKH 460
>BG587250 weakly similar to GP|3947735|emb| NL27 {Solanum tuberosum}, partial
(9%)
Length = 655
Score = 181 bits (458), Expect = 4e-46
Identities = 90/152 (59%), Positives = 111/152 (72%)
Frame = +1
Query: 6 KKNDHTVFVSFRGTDTRFNFTDHLFGALQRKRIFTFRDDTNLQKGNSIASDLIQAIEGSQ 65
++N + VFV+FRG DTR NFT+ LF AL+RK I+ FRDDTNL KG SI +L++ IEGSQ
Sbjct: 58 RRNCYDVFVTFRGEDTRNNFTNFLFAALERKGIYAFRDDTNLPKGESIGPELLRTIEGSQ 237
Query: 66 VFIVVFSKNYASSTWCLRELAYILNCSVLYGKRVLPVFYDVDLSEVRKQSGGYGESFNYH 125
VF+ V S+NYASSTWCL+EL I C GK VLP+FY VD SEV+KQSG Y + F H
Sbjct: 238 VFVAVLSRNYASSTWCLQELEKICECIKGSGKYVLPIFYGVDPSEVKKQSGIYWDDFAKH 417
Query: 126 GKRFQDHSNMVQRRRETLQLVGNISGWDLRDK 157
+RF+ + V R RE L VG+I+GWDLRDK
Sbjct: 418 EQRFKQDPHKVSRWREALNQVGSIAGWDLRDK 513
Score = 157 bits (398), Expect = 4e-39
Identities = 77/116 (66%), Positives = 89/116 (76%)
Frame = +1
Query: 189 YDVFVSFRGPDTRFNFTDHFCAALQRRGINAFRDDTKLKKGEFIAPGLFRAIEASQVYIV 248
YDVFV+FRG DTR NFT+ AAL+R+GI AFRDDT L KGE I P L R IE SQV++
Sbjct: 70 YDVFVTFRGEDTRNNFTNFLFAALERKGIYAFRDDTNLPKGESIGPELLRTIEGSQVFVA 249
Query: 249 VFSKNYASSTWCLRELEYILHCSKKHGKHVLPVFYDVDPSEVQKQSGGYGDALSKH 304
V S+NYASSTWCL+ELE I C K GK+VLP+FY VDPSEV+KQSG Y D +KH
Sbjct: 250 VLSRNYASSTWCLQELEKICECIKGSGKYVLPIFYGVDPSEVKKQSGIYWDDFAKH 417
>BF635520 weakly similar to GP|3947735|emb| NL27 {Solanum tuberosum}, partial
(8%)
Length = 577
Score = 159 bits (403), Expect = 1e-39
Identities = 84/142 (59%), Positives = 98/142 (68%)
Frame = +1
Query: 6 KKNDHTVFVSFRGTDTRFNFTDHLFGALQRKRIFTFRDDTNLQKGNSIASDLIQAIEGSQ 65
KKN + VFV++RG DTR N TD LF AL+ K I FRDD NLQKG I S+L++AIE S
Sbjct: 76 KKNYYDVFVTYRGEDTRNNLTDFLFDALESKGIMVFRDDINLQKGEYIGSELLRAIERSH 255
Query: 66 VFIVVFSKNYASSTWCLRELAYILNCSVLYGKRVLPVFYDVDLSEVRKQSGGYGESFNYH 125
V++ VFS+NYASSTWCL+EL I C K VLP+FYDVD SEVRKQSG Y E+F H
Sbjct: 256 VYVAVFSRNYASSTWCLQELEKICECIEGLEKHVLPIFYDVDPSEVRKQSGIYWEAFVKH 435
Query: 126 GKRFQDHSNMVQRRRETLQLVG 147
+RFQ MV R RE L G
Sbjct: 436 EQRFQQDFQMVSRWREALNKWG 501
Score = 149 bits (377), Expect = 1e-36
Identities = 73/116 (62%), Positives = 86/116 (73%)
Frame = +1
Query: 189 YDVFVSFRGPDTRFNFTDHFCAALQRRGINAFRDDTKLKKGEFIAPGLFRAIEASQVYIV 248
YDVFV++RG DTR N TD AL+ +GI FRDD L+KGE+I L RAIE S VY+
Sbjct: 88 YDVFVTYRGEDTRNNLTDFLFDALESKGIMVFRDDINLQKGEYIGSELLRAIERSHVYVA 267
Query: 249 VFSKNYASSTWCLRELEYILHCSKKHGKHVLPVFYDVDPSEVQKQSGGYGDALSKH 304
VFS+NYASSTWCL+ELE I C + KHVLP+FYDVDPSEV+KQSG Y +A KH
Sbjct: 268 VFSRNYASSTWCLQELEKICECIEGLEKHVLPIFYDVDPSEVRKQSGIYWEAFVKH 435
Score = 37.7 bits (86), Expect = 0.006
Identities = 19/38 (50%), Positives = 26/38 (68%)
Frame = +3
Query: 133 SNMVQRRRETLQLVGNISGWDLRDKPHHAELENIIEHI 170
SN V+ + Q VG+ISGWDLRDKP A ++ I++ I
Sbjct: 459 SNGVKMEGSSKQ-VGSISGWDLRDKPQAAXIKKIVQKI 569
>AW686747 weakly similar to PIR|A54810|A5 TMV resistance protein N - tobacco
(Nicotiana glutinosa), partial (10%)
Length = 644
Score = 156 bits (394), Expect = 1e-38
Identities = 76/159 (47%), Positives = 107/159 (66%)
Frame = +3
Query: 12 VFVSFRGTDTRFNFTDHLFGALQRKRIFTFRDDTNLQKGNSIASDLIQAIEGSQVFIVVF 71
VF+SFRG DTR FTDHLF +L+R+ I TF+DD +L++G I+ +L +AIE S I++
Sbjct: 126 VFLSFRGEDTRQGFTDHLFASLERRGIKTFKDDHDLERGEVISYELNKAIEESMFAIIIL 305
Query: 72 SKNYASSTWCLRELAYILNCSVLYGKRVLPVFYDVDLSEVRKQSGGYGESFNYHGKRFQD 131
S NYASSTWCL EL I+ CS +G+ V P+FY VD S+VR Q G + E+F H ++F+
Sbjct: 306 SPNYASSTWCLDELKKIVECSKSFGQAVFPIFYGVDPSDVRHQRGSFDEAFRKHEEKFRK 485
Query: 132 HSNMVQRRRETLQLVGNISGWDLRDKPHHAELENIIEHI 170
V+R R+ L+ V SGWD + + + +E I+EHI
Sbjct: 486 DRTKVERWRDALREVAGYSGWDSKGRHEASLVETIVEHI 602
Score = 132 bits (331), Expect = 2e-31
Identities = 63/114 (55%), Positives = 81/114 (70%)
Frame = +3
Query: 191 VFVSFRGPDTRFNFTDHFCAALQRRGINAFRDDTKLKKGEFIAPGLFRAIEASQVYIVVF 250
VF+SFRG DTR FTDH A+L+RRGI F+DD L++GE I+ L +AIE S I++
Sbjct: 126 VFLSFRGEDTRQGFTDHLFASLERRGIKTFKDDHDLERGEVISYELNKAIEESMFAIIIL 305
Query: 251 SKNYASSTWCLRELEYILHCSKKHGKHVLPVFYDVDPSEVQKQSGGYGDALSKH 304
S NYASSTWCL EL+ I+ CSK G+ V P+FY VDPS+V+ Q G + +A KH
Sbjct: 306 SPNYASSTWCLDELKKIVECSKSFGQAVFPIFYGVDPSDVRHQRGSFDEAFRKH 467
>TC91648 weakly similar to GP|23477201|emb|CAD36199. NLS-TIR-NBS disease
resistance protein {Populus tremula}, partial (17%)
Length = 666
Score = 134 bits (336), Expect = 6e-32
Identities = 74/140 (52%), Positives = 91/140 (64%), Gaps = 6/140 (4%)
Frame = +2
Query: 35 RKRIFTFRDDTNLQKGNSIASDLIQAIEGSQVFIVVFSKNYASSTWCLRELAYILNCSVL 94
+K IF+FRDDT L+KG SIA +L++AIE SQ+F+VVFSKNYASS WCLREL IL L
Sbjct: 245 KKIIFSFRDDTKLKKGESIAPELLRAIEDSQIFVVVFSKNYASSVWCLRELECILQSFQL 424
Query: 95 YGKRVLPVFYDVDLSEVRKQSGGYGESFNYHGKRFQDHSNMVQRRRETLQLVGNISGW-- 152
GKRVLPVFYDVD SEVR Q G Y E+ H +RFQ + + + G S W
Sbjct: 425 SGKRVLPVFYDVDPSEVRYQKGCYAEALAKHEERFQQNFENXAKMEGSTDTSGQ-SLWMG 601
Query: 153 ----DLRDKPHHAELENIIE 168
+ +P HAE+E I+E
Sbjct: 602 CTX*NKTCRPQHAEIEKIVE 661
Score = 127 bits (319), Expect(2) = 2e-38
Identities = 65/97 (67%), Positives = 75/97 (77%)
Frame = +2
Query: 208 FCAALQRRGINAFRDDTKLKKGEFIAPGLFRAIEASQVYIVVFSKNYASSTWCLRELEYI 267
FC ++ I +FRDDTKLKKGE IAP L RAIE SQ+++VVFSKNYASS WCLRELE I
Sbjct: 230 FCCPAKKI-IFSFRDDTKLKKGESIAPELLRAIEDSQIFVVVFSKNYASSVWCLRELECI 406
Query: 268 LHCSKKHGKHVLPVFYDVDPSEVQKQSGGYGDALSKH 304
L + GK VLPVFYDVDPSEV+ Q G Y +AL+KH
Sbjct: 407 LQSFQLSGKRVLPVFYDVDPSEVRYQKGCYAEALAKH 517
Score = 59.7 bits (143), Expect = 1e-09
Identities = 56/179 (31%), Positives = 79/179 (43%), Gaps = 23/179 (12%)
Frame = +3
Query: 2 TSPRKKNDHTVFVSFRGTDTRFNFTDHLFGALQRKRIFTFRDDTNLQKGNSIASDLIQAI 61
+SPR+ N + VFVSFRG DTR NFTDHLF ALQRK F + +K N + + +
Sbjct: 150 SSPRR-NYYDVFVSFRGKDTRLNFTDHLFAALQRKSFFHSGMILS*RKVNP*HLNSFEQL 326
Query: 62 EGSQVFIVVFSK--------------------NYASSTWCLRELAYILNCSVLYGKRVLP 101
+ + F++ FS NY + +CL CS++ +
Sbjct: 327 KTLR-FLLWFSLRTMLPQFGVCENWSAYFKAFNYLENVFCL--------CSMMLIHQRCD 479
Query: 102 VFYDVDLS---EVRKQSGGYGESFNYHGKRFQDHSNMVQRRRETLQLVGNISGWDLRDK 157
+ DV L ++K S + QR RE L V N+SGWD+R K
Sbjct: 480 IKKDVMLKP*LNMKKDS--------------NKTLKIXQRWREALTQVANLSGWDVRXK 614
Score = 49.3 bits (116), Expect(2) = 2e-38
Identities = 22/27 (81%), Positives = 23/27 (84%)
Frame = +3
Query: 189 YDVFVSFRGPDTRFNFTDHFCAALQRR 215
YDVFVSFRG DTR NFTDH AALQR+
Sbjct: 171 YDVFVSFRGKDTRLNFTDHLFAALQRK 251
>TC89557 weakly similar to GP|10178211|dbj|BAB11635. TMV resistance protein
N {Arabidopsis thaliana}, partial (8%)
Length = 1567
Score = 104 bits (259), Expect(2) = 3e-38
Identities = 51/94 (54%), Positives = 66/94 (69%)
Frame = +3
Query: 10 HTVFVSFRGTDTRFNFTDHLFGALQRKRIFTFRDDTNLQKGNSIASDLIQAIEGSQVFIV 69
+ VF+SFRG+DTR +F DHL+ L RK IFTF+DD LQKG +I+ L+QAI+ S++ IV
Sbjct: 264 YDVFISFRGSDTRNSFVDHLYAHLNRKGIFTFKDDKQLQKGEAISPQLLQAIQQSRISIV 443
Query: 70 VFSKNYASSTWCLRELAYILNCSVLYGKRVLPVF 103
VFSK+YASSTWCL E+A I + P F
Sbjct: 444 VFSKDYASSTWCLDEMAAINESRINLKTSCFPCF 545
Score = 100 bits (249), Expect(2) = 3e-27
Identities = 48/79 (60%), Positives = 61/79 (76%)
Frame = +3
Query: 189 YDVFVSFRGPDTRFNFTDHFCAALQRRGINAFRDDTKLKKGEFIAPGLFRAIEASQVYIV 248
YDVF+SFRG DTR +F DH A L R+GI F+DD +L+KGE I+P L +AI+ S++ IV
Sbjct: 264 YDVFISFRGSDTRNSFVDHLYAHLNRKGIFTFKDDKQLQKGEAISPQLLQAIQQSRISIV 443
Query: 249 VFSKNYASSTWCLRELEYI 267
VFSK+YASSTWCL E+ I
Sbjct: 444 VFSKDYASSTWCLDEMAAI 500
Score = 72.0 bits (175), Expect(2) = 3e-38
Identities = 39/90 (43%), Positives = 53/90 (58%), Gaps = 1/90 (1%)
Frame = +1
Query: 99 VLPVFYDVDLSEVRKQSGGYGESFNYHGKRFQDHSNMVQRRRETLQLVGNISGWDLRDKP 158
V PVFYDVD S V+KQ+G Y +F H + F+ +S V R R T+ +G +GWD+R +P
Sbjct: 532 VFPVFYDVDPSHVKKQNGVYENAFVLHTEAFKHNSGKVARWRTTMTYLGGTAGWDVRVEP 711
Query: 159 HHAELENIIEH-INILGCKFPSRTKDLVEI 187
+E I+E I LG F T DL+ I
Sbjct: 712 EFEMIEKIVEAVIKKLGRTFSGSTDDLIGI 801
Score = 38.5 bits (88), Expect(2) = 3e-27
Identities = 17/27 (62%), Positives = 20/27 (73%)
Frame = +1
Query: 278 VLPVFYDVDPSEVQKQSGGYGDALSKH 304
V PVFYDVDPS V+KQ+G Y +A H
Sbjct: 532 VFPVFYDVDPSHVKKQNGVYENAFVLH 612
>AL372504 weakly similar to GP|7341113|gb| unknown {Glycine max}, partial
(70%)
Length = 478
Score = 151 bits (382), Expect = 3e-37
Identities = 73/105 (69%), Positives = 84/105 (79%)
Frame = +2
Query: 189 YDVFVSFRGPDTRFNFTDHFCAALQRRGINAFRDDTKLKKGEFIAPGLFRAIEASQVYIV 248
YDVFV+FRG DTR NFTD+ AL+ +GI AFRDDT LKKGE I P L RAIE SQV++
Sbjct: 164 YDVFVTFRGEDTRNNFTDYLFDALETKGIYAFRDDTNLKKGEVIGPELLRAIEGSQVFVA 343
Query: 249 VFSKNYASSTWCLRELEYILHCSKKHGKHVLPVFYDVDPSEVQKQ 293
VFS+NYASSTWCL+ELE I C + KHVLPVFYD+DPSEV+KQ
Sbjct: 344 VFSRNYASSTWCLQELEKICECVQGPEKHVLPVFYDIDPSEVRKQ 478
Score = 143 bits (360), Expect = 1e-34
Identities = 70/114 (61%), Positives = 87/114 (75%)
Frame = +2
Query: 1 MTSPRKKNDHTVFVSFRGTDTRFNFTDHLFGALQRKRIFTFRDDTNLQKGNSIASDLIQA 60
+ + ++N + VFV+FRG DTR NFTD+LF AL+ K I+ FRDDTNL+KG I +L++A
Sbjct: 137 LVTSSRRNYYDVFVTFRGEDTRNNFTDYLFDALETKGIYAFRDDTNLKKGEVIGPELLRA 316
Query: 61 IEGSQVFIVVFSKNYASSTWCLRELAYILNCSVLYGKRVLPVFYDVDLSEVRKQ 114
IEGSQVF+ VFS+NYASSTWCL+EL I C K VLPVFYD+D SEVRKQ
Sbjct: 317 IEGSQVFVAVFSRNYASSTWCLQELEKICECVQGPEKHVLPVFYDIDPSEVRKQ 478
>TC93749 similar to GP|3947733|emb|CAA08797.1 NL25 {Solanum tuberosum},
partial (17%)
Length = 772
Score = 147 bits (372), Expect = 4e-36
Identities = 78/173 (45%), Positives = 107/173 (61%), Gaps = 5/173 (2%)
Frame = +2
Query: 4 PRKKNDHTVFVSFRGTDTRFNFTDHLFGALQRKRIFTFRDDTNLQKGNSIASDLIQAIEG 63
P + VF+SFRG DTR+ FT HL L R ++ T+ D NLQ+G+ I+S L+ AIE
Sbjct: 32 PHHQKHGQVFLSFRGEDTRYTFTSHLHATLTRLKVGTYID-YNLQRGDEISSTLLMAIEE 208
Query: 64 SQVFIVVFSKNYASSTWCLRELAYILNCSVLYGKRVLPVFYDVDLSEVRKQSGGYGESFN 123
++V IV+FSKNY +S WCL EL IL C + G+ +LP+FYD+D S VR Q+G Y E+F
Sbjct: 209 AKVSIVIFSKNYGNSKWCLDELVKILECKKMKGQILLPIFYDIDPSHVRNQTGSYAEAFV 388
Query: 124 YHGKRFQDHSNMVQRRRETLQLVGNISGWDLRDKPHHAEL-----ENIIEHIN 171
H K+FQ VQ R L+ NISGW+ +EL +++IE +N
Sbjct: 389 KHEKQFQGKLEKVQTWRHALREAANISGWECSVNRMESELLEKIAKDVIEKLN 547
Score = 116 bits (291), Expect = 1e-26
Identities = 57/114 (50%), Positives = 76/114 (66%)
Frame = +2
Query: 191 VFVSFRGPDTRFNFTDHFCAALQRRGINAFRDDTKLKKGEFIAPGLFRAIEASQVYIVVF 250
VF+SFRG DTR+ FT H A L R + + D L++G+ I+ L AIE ++V IV+F
Sbjct: 56 VFLSFRGEDTRYTFTSHLHATLTRLKVGTYID-YNLQRGDEISSTLLMAIEEAKVSIVIF 232
Query: 251 SKNYASSTWCLRELEYILHCSKKHGKHVLPVFYDVDPSEVQKQSGGYGDALSKH 304
SKNY +S WCL EL IL C K G+ +LP+FYD+DPS V+ Q+G Y +A KH
Sbjct: 233 SKNYGNSKWCLDELVKILECKKMKGQILLPIFYDIDPSHVRNQTGSYAEAFVKH 394
>TC93448 similar to GP|9858478|gb|AAG01052.1| resistance protein MG23
{Glycine max}, partial (28%)
Length = 673
Score = 144 bits (363), Expect = 4e-35
Identities = 77/159 (48%), Positives = 105/159 (65%), Gaps = 2/159 (1%)
Frame = +1
Query: 15 SFRGTDTRFNFTDHLFGALQRKRIFTFRDDTNLQKGNSIASDLIQAIEGSQVFIVVFSKN 74
SFRG DTR+ FT +L+ AL K I TF DD LQ+G+ I L++AIE S++ I+V SKN
Sbjct: 139 SFRGLDTRYGFTGNLYKALYDKGIHTFIDDEELQRGHEITPSLLEAIEESRIAIIVLSKN 318
Query: 75 YASSTWCLRELAYILNCSVLYGKRVLPVFYDVDLSEVRKQSGGYGESFNYHGKRFQDHSN 134
YASS++CL EL IL+C G+ V P+FYDVD S+VRKQ+G YGE+ G+RF D N
Sbjct: 319 YASSSFCLHELVKILDCIKGKGRLVWPIFYDVDPSDVRKQTGSYGEALAMLGERFND--N 492
Query: 135 MVQRRRETLQLVGNISGWDLR--DKPHHAELENIIEHIN 171
+Q + LQ V N+SGW + D + + I+EH++
Sbjct: 493 NLQIWKNALQQVANLSGWHFKIGDGYEYQFIGKIVEHVS 609
Score = 123 bits (309), Expect = 8e-29
Identities = 61/112 (54%), Positives = 80/112 (70%)
Frame = +1
Query: 194 SFRGPDTRFNFTDHFCAALQRRGINAFRDDTKLKKGEFIAPGLFRAIEASQVYIVVFSKN 253
SFRG DTR+ FT + AL +GI+ F DD +L++G I P L AIE S++ I+V SKN
Sbjct: 139 SFRGLDTRYGFTGNLYKALYDKGIHTFIDDEELQRGHEITPSLLEAIEESRIAIIVLSKN 318
Query: 254 YASSTWCLRELEYILHCSKKHGKHVLPVFYDVDPSEVQKQSGGYGDALSKHG 305
YASS++CL EL IL C K G+ V P+FYDVDPS+V+KQ+G YG+AL+ G
Sbjct: 319 YASSSFCLHELVKILDCIKGKGRLVWPIFYDVDPSDVRKQTGSYGEALAMLG 474
>BF650400 weakly similar to GP|9965109|gb| resistance protein MG13 {Glycine
max}, partial (36%)
Length = 644
Score = 142 bits (359), Expect = 1e-34
Identities = 71/154 (46%), Positives = 103/154 (66%)
Frame = +1
Query: 3 SPRKKNDHTVFVSFRGTDTRFNFTDHLFGALQRKRIFTFRDDTNLQKGNSIASDLIQAIE 62
+P+KK D VF+SFRG DTR NFT HL+ L RK++ TF D+ L+KG+ I+ LI+AIE
Sbjct: 136 APQKKYD--VFLSFRGDDTRRNFTSHLYDTLSRKKVETFIDNNELEKGDEISPALIKAIE 309
Query: 63 GSQVFIVVFSKNYASSTWCLRELAYILNCSVLYGKRVLPVFYDVDLSEVRKQSGGYGESF 122
S V I++FS+NYASS WCL EL IL C + V+PVFY++D S VRKQ+G Y ++F
Sbjct: 310 ESHVSIIIFSENYASSKWCLNELKKILQCKKYMQQIVIPVFYNIDPSHVRKQTGSYEQAF 489
Query: 123 NYHGKRFQDHSNMVQRRRETLQLVGNISGWDLRD 156
H + + +++ +Q + L ++ GWD ++
Sbjct: 490 TKHMRDLKLNNDKLQEWKAALAEAASLVGWDFQN 591
Score = 135 bits (339), Expect = 3e-32
Identities = 63/116 (54%), Positives = 83/116 (71%)
Frame = +1
Query: 189 YDVFVSFRGPDTRFNFTDHFCAALQRRGINAFRDDTKLKKGEFIAPGLFRAIEASQVYIV 248
YDVF+SFRG DTR NFT H L R+ + F D+ +L+KG+ I+P L +AIE S V I+
Sbjct: 151 YDVFLSFRGDDTRRNFTSHLYDTLSRKKVETFIDNNELEKGDEISPALIKAIEESHVSII 330
Query: 249 VFSKNYASSTWCLRELEYILHCSKKHGKHVLPVFYDVDPSEVQKQSGGYGDALSKH 304
+FS+NYASS WCL EL+ IL C K + V+PVFY++DPS V+KQ+G Y A +KH
Sbjct: 331 IFSENYASSKWCLNELKKILQCKKYMQQIVIPVFYNIDPSHVRKQTGSYEQAFTKH 498
>AW694406 weakly similar to GP|3947735|emb NL27 {Solanum tuberosum}, partial
(14%)
Length = 628
Score = 139 bits (349), Expect = 2e-33
Identities = 65/130 (50%), Positives = 90/130 (69%)
Frame = +3
Query: 179 SRTKDLVEINYDVFVSFRGPDTRFNFTDHFCAALQRRGINAFRDDTKLKKGEFIAPGLFR 238
S T ++ ++DVF+SFRG DTR FT H AL++ G+ F DD++LKKG+ I+ L +
Sbjct: 81 SSTLEVASNSFDVFISFRGDDTRRKFTSHLNEALKKSGVKTFIDDSELKKGDEISSALIK 260
Query: 239 AIEASQVYIVVFSKNYASSTWCLRELEYILHCSKKHGKHVLPVFYDVDPSEVQKQSGGYG 298
AIE S IV+FS++YASS WCL EL IL C K +G+ V+P+FY++DPS V+ Q G YG
Sbjct: 261 AIEESCASIVIFSEDYASSKWCLNELVKILECKKDNGQIVIPIFYEIDPSHVRNQIGSYG 440
Query: 299 DALSKHGLNM 308
A +KH N+
Sbjct: 441 QAFAKHEKNL 470
Score = 137 bits (346), Expect = 4e-33
Identities = 71/146 (48%), Positives = 95/146 (64%)
Frame = +3
Query: 8 NDHTVFVSFRGTDTRFNFTDHLFGALQRKRIFTFRDDTNLQKGNSIASDLIQAIEGSQVF 67
N VF+SFRG DTR FT HL AL++ + TF DD+ L+KG+ I+S LI+AIE S
Sbjct: 105 NSFDVFISFRGDDTRRKFTSHLNEALKKSGVKTFIDDSELKKGDEISSALIKAIEESCAS 284
Query: 68 IVVFSKNYASSTWCLRELAYILNCSVLYGKRVLPVFYDVDLSEVRKQSGGYGESFNYHGK 127
IV+FS++YASS WCL EL IL C G+ V+P+FY++D S VR Q G YG++F H K
Sbjct: 285 IVIFSEDYASSKWCLNELVKILECKKDNGQIVIPIFYEIDPSHVRNQIGSYGQAFAKHEK 464
Query: 128 RFQDHSNMVQRRRETLQLVGNISGWD 153
+ Q+ ++ L V N+SGWD
Sbjct: 465 NLKQ-----QKWKDALTEVSNLSGWD 527
>TC92554 similar to GP|9758876|dbj|BAB09430.1 disease resistance protein
{Arabidopsis thaliana}, partial (7%)
Length = 1043
Score = 134 bits (336), Expect(2) = 3e-33
Identities = 63/132 (47%), Positives = 88/132 (65%)
Frame = +1
Query: 3 SPRKKNDHTVFVSFRGTDTRFNFTDHLFGALQRKRIFTFRDDTNLQKGNSIASDLIQAIE 62
S H VF++FRG DTR+NFT HL+ AL K+I TF DD +++G++I+ L+ AIE
Sbjct: 214 SHSNSKQHHVFLNFRGEDTRYNFTSHLYAALCGKKIHTFMDDEEIERGDNISPTLLSAIE 393
Query: 63 GSQVFIVVFSKNYASSTWCLRELAYILNCSVLYGKRVLPVFYDVDLSEVRKQSGGYGESF 122
S++ +++FS++YASS+WCL EL I+ CS V+PVFY +D S VR Q G YG++F
Sbjct: 394 SSKICVIIFSQDYASSSWCLDELVKIIECSEKKKLVVIPVFYHIDASHVRHQRGTYGDAF 573
Query: 123 NYHGKRFQDHSN 134
H FQ SN
Sbjct: 574 AKHEDPFQGQSN 609
Score = 127 bits (318), Expect = 7e-30
Identities = 58/116 (50%), Positives = 85/116 (73%)
Frame = +1
Query: 189 YDVFVSFRGPDTRFNFTDHFCAALQRRGINAFRDDTKLKKGEFIAPGLFRAIEASQVYIV 248
+ VF++FRG DTR+NFT H AAL + I+ F DD ++++G+ I+P L AIE+S++ ++
Sbjct: 235 HHVFLNFRGEDTRYNFTSHLYAALCGKKIHTFMDDEEIERGDNISPTLLSAIESSKICVI 414
Query: 249 VFSKNYASSTWCLRELEYILHCSKKHGKHVLPVFYDVDPSEVQKQSGGYGDALSKH 304
+FS++YASS+WCL EL I+ CS+K V+PVFY +D S V+ Q G YGDA +KH
Sbjct: 415 IFSQDYASSSWCLDELVKIIECSEKKKLVVIPVFYHIDASHVRHQRGTYGDAFAKH 582
Score = 25.4 bits (54), Expect(2) = 3e-33
Identities = 20/67 (29%), Positives = 33/67 (48%), Gaps = 3/67 (4%)
Frame = +2
Query: 128 RFQDHSNMVQRRRETLQLVGNISGW-DLRDKPHHAELENIIEHI-NILGCKFPS-RTKDL 184
RF+D+ V R L N++GW +++ ++ I+E I + L C P + K L
Sbjct: 590 RFRDNLTKVHMWRTALHKAANLAGWVSEKNRSEAVLIKEIVEVILDKLKCMSPHVQNKGL 769
Query: 185 VEINYDV 191
V I+ V
Sbjct: 770 VGISVHV 790
>BG647688 similar to GP|9965109|gb| resistance protein MG13 {Glycine max},
partial (38%)
Length = 719
Score = 136 bits (342), Expect = 1e-32
Identities = 62/116 (53%), Positives = 91/116 (78%)
Frame = +3
Query: 189 YDVFVSFRGPDTRFNFTDHFCAALQRRGINAFRDDTKLKKGEFIAPGLFRAIEASQVYIV 248
YDVF+SFRG DTR+ FT + AL+ +GI+ F DD +L++G+ I P L +AI+ S++ I+
Sbjct: 60 YDVFLSFRGTDTRYGFTGNLYEALRVKGIHTFIDDRELQRGDQITPSLLKAIQESKIVII 239
Query: 249 VFSKNYASSTWCLRELEYILHCSKKHGKHVLPVFYDVDPSEVQKQSGGYGDALSKH 304
VFS +YASS++CL EL +I+HCSK++G VLP+FY V+PS V+ Q+G YG+AL++H
Sbjct: 240 VFSNHYASSSFCLDELVHIIHCSKENGCLVLPIFYGVEPSHVRYQTGSYGEALAEH 407
Score = 129 bits (325), Expect = 1e-30
Identities = 66/148 (44%), Positives = 100/148 (66%), Gaps = 5/148 (3%)
Frame = +3
Query: 10 HTVFVSFRGTDTRFNFTDHLFGALQRKRIFTFRDDTNLQKGNSIASDLIQAIEGSQVFIV 69
+ VF+SFRGTDTR+ FT +L+ AL+ K I TF DD LQ+G+ I L++AI+ S++ I+
Sbjct: 60 YDVFLSFRGTDTRYGFTGNLYEALRVKGIHTFIDDRELQRGDQITPSLLKAIQESKIVII 239
Query: 70 VFSKNYASSTWCLRELAYILNCSVLYGKRVLPVFYDVDLSEVRKQSGGYGESFNYH---- 125
VFS +YASS++CL EL +I++CS G VLP+FY V+ S VR Q+G YGE+ H
Sbjct: 240 VFSNHYASSSFCLDELVHIIHCSKENGCLVLPIFYGVEPSHVRYQTGSYGEALAEHEEAR 419
Query: 126 -GKRFQDHSNMVQRRRETLQLVGNISGW 152
++++D+ +Q+ L+ N+SG+
Sbjct: 420 KKEKYKDNMEKLQKWEMALKQAANLSGY 503
>BG645347 weakly similar to GP|18033111|g functional candidate resistance
protein KR1 {Glycine max}, partial (9%)
Length = 760
Score = 103 bits (256), Expect(2) = 2e-32
Identities = 52/94 (55%), Positives = 67/94 (70%)
Frame = +2
Query: 10 HTVFVSFRGTDTRFNFTDHLFGALQRKRIFTFRDDTNLQKGNSIASDLIQAIEGSQVFIV 69
+ VF+SFRG DTR NFT +L+ +L ++ I TF D+ +QKG I L+QAI+ S++FI
Sbjct: 38 YDVFLSFRGIDTRNNFTGNLYNSLNQRGIKTFFDEQEIQKGEEITHTLLQAIK*SRIFIA 217
Query: 70 VFSKNYASSTWCLRELAYILNCSVLYGKRVLPVF 103
VFS NYASST+CL EL IL CS L G+ LPVF
Sbjct: 218 VFSTNYASSTFCLTELVTILECSKLQGRLFLPVF 319
Score = 102 bits (255), Expect(2) = 3e-26
Identities = 54/105 (51%), Positives = 71/105 (67%)
Frame = +2
Query: 178 PSRTKDLVEINYDVFVSFRGPDTRFNFTDHFCAALQRRGINAFRDDTKLKKGEFIAPGLF 237
PS + + YDVF+SFRG DTR NFT + +L +RGI F D+ +++KGE I L
Sbjct: 5 PSLSSFTCDWTYDVFLSFRGIDTRNNFTGNLYNSLNQRGIKTFFDEQEIQKGEEITHTLL 184
Query: 238 RAIEASQVYIVVFSKNYASSTWCLRELEYILHCSKKHGKHVLPVF 282
+AI+ S+++I VFS NYASST+CL EL IL CSK G+ LPVF
Sbjct: 185 QAIK*SRIFIAVFSTNYASSTFCLTELVTILECSKLQGRLFLPVF 319
Score = 53.5 bits (127), Expect(2) = 2e-32
Identities = 25/50 (50%), Positives = 31/50 (62%)
Frame = +3
Query: 103 FYDVDLSEVRKQSGGYGESFNYHGKRFQDHSNMVQRRRETLQLVGNISGW 152
FYDVD S VRK + Y E+F H RF+D N VQ+ R+ L N+SGW
Sbjct: 318 FYDVDPSHVRKLNEAYAEAFAKHEVRFRDEKNKVQKWRDALYQAANMSGW 467
Score = 33.1 bits (74), Expect(2) = 3e-26
Identities = 13/23 (56%), Positives = 17/23 (73%)
Frame = +3
Query: 282 FYDVDPSEVQKQSGGYGDALSKH 304
FYDVDPS V+K + Y +A +KH
Sbjct: 318 FYDVDPSHVRKLNEAYAEAFAKH 386
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.323 0.139 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,794,480
Number of Sequences: 36976
Number of extensions: 134721
Number of successful extensions: 907
Number of sequences better than 10.0: 128
Number of HSP's better than 10.0 without gapping: 695
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 849
length of query: 309
length of database: 9,014,727
effective HSP length: 96
effective length of query: 213
effective length of database: 5,465,031
effective search space: 1164051603
effective search space used: 1164051603
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 58 (26.9 bits)
Medicago: description of AC146863.4


