

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146862.24 - phase: 0
(580 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
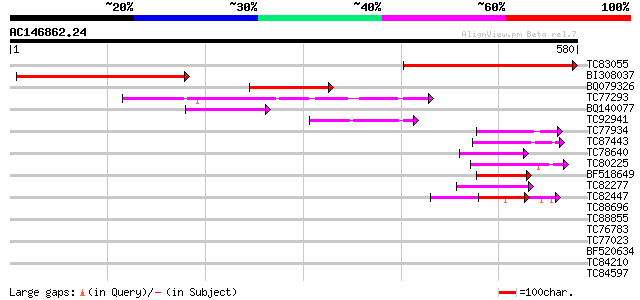
Score E
Sequences producing significant alignments: (bits) Value
TC83055 similar to GP|12583663|dbj|BAB21485. chloroplast RelA ho... 347 7e-96
BI308037 weakly similar to GP|12583663|dbj chloroplast RelA homo... 341 5e-94
BQ079326 weakly similar to GP|9294134|dbj| contains similarity t... 171 9e-43
TC77293 similar to GP|15081594|gb|AAK82651.1 RSH-like protein {C... 134 1e-31
BQ140077 similar to GP|7141304|gb|A RSH1 {Arabidopsis thaliana},... 61 1e-09
TC92941 weakly similar to GP|20516195|gb|AAM24425.1 Guanosine po... 57 1e-08
TC77934 similar to GP|9965747|gb|AAG10150.1| calmodulin-like MSS... 52 5e-07
TC87443 similar to GP|13397927|emb|CAC34625. putative calmodulin... 49 5e-06
TC78640 homologue to GP|13397927|emb|CAC34625. putative calmodul... 48 9e-06
TC80225 similar to GP|14589311|emb|CAC43238. calcium binding pro... 47 2e-05
BF518649 weakly similar to GP|12963415|gb| regulator of gene sil... 46 3e-05
TC82277 weakly similar to PIR|A84532|A84532 probable calmodulin-... 46 3e-05
TC82447 similar to GP|14589311|emb|CAC43238. calcium binding pro... 46 3e-05
TC88696 weakly similar to PIR|T10620|T10620 probable calcium-bin... 41 0.001
TC88855 PIR|A30900|MCBH calmodulin - barley, complete 40 0.002
TC76783 calmodulin 1 [Medicago truncatula] 40 0.002
TC77023 calmodulin 2 [Medicago truncatula] 40 0.002
BF520634 weakly similar to GP|17428592|emb PROBABLE GTP PYROPHOS... 39 0.006
TC84210 homologue to PIR|T07122|T07122 calmodulin CAM5 - soybean... 38 0.009
TC84597 38 0.012
>TC83055 similar to GP|12583663|dbj|BAB21485. chloroplast RelA homologue 2
{Oryza sativa (japonica cultivar-group)}, partial (26%)
Length = 837
Score = 347 bits (890), Expect = 7e-96
Identities = 177/177 (100%), Positives = 177/177 (100%)
Frame = +2
Query: 404 RTRPLMEIQIRTTEMDRLAVGGMASHSLYKAGLTNPEEAKRLKTIMLAAAELAALRLKDF 463
RTRPLMEIQIRTTEMDRLAVGGMASHSLYKAGLTNPEEAKRLKTIMLAAAELAALRLKDF
Sbjct: 2 RTRPLMEIQIRTTEMDRLAVGGMASHSLYKAGLTNPEEAKRLKTIMLAAAELAALRLKDF 181
Query: 464 PSANHKGIEFDQRDRVFRLLDKNGDGKISIEELTEVIEELGAPGEDAHDMMLLLDSNSDG 523
PSANHKGIEFDQRDRVFRLLDKNGDGKISIEELTEVIEELGAPGEDAHDMMLLLDSNSDG
Sbjct: 182 PSANHKGIEFDQRDRVFRLLDKNGDGKISIEELTEVIEELGAPGEDAHDMMLLLDSNSDG 361
Query: 524 SLSSDEFQMFQKQVEMVRNLEDRDDEYKKILDEKLHMADDSGLIQVYNKEFGNRLVS 580
SLSSDEFQMFQKQVEMVRNLEDRDDEYKKILDEKLHMADDSGLIQVYNKEFGNRLVS
Sbjct: 362 SLSSDEFQMFQKQVEMVRNLEDRDDEYKKILDEKLHMADDSGLIQVYNKEFGNRLVS 532
>BI308037 weakly similar to GP|12583663|dbj chloroplast RelA homologue 2
{Oryza sativa (japonica cultivar-group)}, partial (10%)
Length = 532
Score = 341 bits (874), Expect = 5e-94
Identities = 176/177 (99%), Positives = 176/177 (99%)
Frame = +2
Query: 8 ASDFPTRHHYLSKPTTFSTTTHRRLLSLFFPTRPTRSTTLRWSSTPRACSVEVPGGGKMV 67
ASDFPTRHHYLSKPTTFSTTTHRRLLSLFFPTRPTRSTTLRWSSTPRACSVEVPGGGKMV
Sbjct: 2 ASDFPTRHHYLSKPTTFSTTTHRRLLSLFFPTRPTRSTTLRWSSTPRACSVEVPGGGKMV 181
Query: 68 IELVGAFNDLTERMKVLSTSSSGLLFKSLKLSIPVLQTSPLTPDGRSPLSKALSIAMLLA 127
IELVGAFNDLTERMKVLSTSSSGLLFKSLKLSIPVLQTSPLTPDGRSPLSKALSIAMLLA
Sbjct: 182 IELVGAFNDLTERMKVLSTSSSGLLFKSLKLSIPVLQTSPLTPDGRSPLSKALSIAMLLA 361
Query: 128 DLQMDAEVISAGILREVLEVGELNLHEIRSQIGSATAHLLHESLRVKNFASRVDILD 184
DLQMDAEVISAGILREVLEVGELNLHEIRSQIGSATAHLLHESLR KNFASRVDILD
Sbjct: 362 DLQMDAEVISAGILREVLEVGELNLHEIRSQIGSATAHLLHESLRGKNFASRVDILD 532
>BQ079326 weakly similar to GP|9294134|dbj| contains similarity to (p)ppGpp
synthase (GTP pyrophosphokinase)~gene_id:MKP6.2, partial
(17%)
Length = 294
Score = 171 bits (432), Expect = 9e-43
Identities = 85/86 (98%), Positives = 85/86 (98%)
Frame = +3
Query: 246 TNYISLELEDLSFQYLFPYSYLYVDTWLRSQETGGISLIDVYKDELLESLKSDPILAELV 305
TNYISLELEDLSFQYLFPYSYLYVDTWLRSQETGGISLIDVYKDELLESLKSDPILAELV
Sbjct: 3 TNYISLELEDLSFQYLFPYSYLYVDTWLRSQETGGISLIDVYKDELLESLKSDPILAELV 182
Query: 306 DDISVKGRYKSRYSTMKKLLKDGRRP 331
DDISVKGRYKSRYSTMKKLLKDG RP
Sbjct: 183 DDISVKGRYKSRYSTMKKLLKDGPRP 260
>TC77293 similar to GP|15081594|gb|AAK82651.1 RSH-like protein {Capsicum
annuum}, partial (70%)
Length = 1931
Score = 134 bits (336), Expect = 1e-31
Identities = 96/325 (29%), Positives = 167/325 (50%), Gaps = 7/325 (2%)
Frame = +2
Query: 116 LSKALSIAMLLADLQMDAEVISAGILREVLEVGELNLHEIRSQIGSATAHLLHESLRVKN 175
L L A+LLA + ++ V++AG+L + ++ L I G+ A L+ ++ +
Sbjct: 311 LQHCLETAVLLALIGANSTVVAAGLLHDTVDDAFLTYDYIYGMFGAGVADLVEGVSKLSH 490
Query: 176 FASRVDILDDENAAA-------LRKFCLTYYDIRALILDLALKLDMMRHLGHLPRYQQQI 228
+ + D N A+ L L D RA+++ LA +L M L LP +QQ
Sbjct: 491 LSK---LARDNNTASKSVEADRLHTMFLAMADARAVLIKLADRLHNMMTLDALPVAKQQR 661
Query: 229 ISLQVMKIYAPLAHAVGTNYISLELEDLSFQYLFPYSYLYVDTWLRSQETGGISLIDVYK 288
+ + ++I+APLA+ +G +LE+L F++L P + + + L E+ ++I
Sbjct: 662 FAKETLEIFAPLANRLGIANWKDQLENLCFKHLNPVQHKELSSKL--VESYDDAMIASAI 835
Query: 289 DELLESLKSDPILAELVDDISVKGRYKSRYSTMKKLLKDGRRPEDVNDVLGLRVVLNPKS 348
+ L ++LK + I ++ GR+KS YS K+LK +D++D+ GLR++++
Sbjct: 836 ERLEQALKDEGISYHVIS-----GRHKSLYSVYCKILKKKLTIDDIHDIYGLRLIVDK-- 994
Query: 349 RENALEAGERACYRAHQIIQSMWKEIPSRTKDYISRPKGNGYRSLHMAVDVSEIGRTRPL 408
E CY+A +++ +W E+ + KDYI PK NGY+SLH V +G +
Sbjct: 995 --------EEDCYKALKVVHRLWSEVHGKLKDYIRFPKFNGYQSLHTVV----MGEGKVP 1138
Query: 409 MEIQIRTTEMDRLAVGGMASHSLYK 433
+E+Q+RT +M A G A+H YK
Sbjct: 1139LEVQVRTKDMHLQAEFGFAAHWRYK 1213
>BQ140077 similar to GP|7141304|gb|A RSH1 {Arabidopsis thaliana}, partial
(10%)
Length = 288
Score = 61.2 bits (147), Expect = 1e-09
Identities = 33/87 (37%), Positives = 52/87 (58%), Gaps = 1/87 (1%)
Frame = +1
Query: 181 DILDDENAAALRKFCLTYYD-IRALILDLALKLDMMRHLGHLPRYQQQIISLQVMKIYAP 239
D + D A LR+ L + +R +I+ LA +L MR L H+P ++Q I+++ ++++AP
Sbjct: 28 DSVQDVKAEDLRQMFLAMTEEVRVIIVKLADRLHNMRTLSHMPPHKQNSIAMETLQVFAP 207
Query: 240 LAHAVGTNYISLELEDLSFQYLFPYSY 266
LA +G I ELE+LSF Y P Y
Sbjct: 208 LAKLLGMYQIKSELENLSFMYTNPEDY 288
>TC92941 weakly similar to GP|20516195|gb|AAM24425.1 Guanosine polyphosphate
pyrophosphohydrolases/synthetases {Thermoanaerobacter
tengcongensis}, partial (11%)
Length = 834
Score = 57.4 bits (137), Expect = 1e-08
Identities = 37/113 (32%), Positives = 54/113 (47%), Gaps = 1/113 (0%)
Frame = +1
Query: 307 DISVKGRYKSRYSTMKKLLKDGRRPEDVNDVLGLRVVLNPKSRENALEA-GERACYRAHQ 365
++++ R KS YS K+ + + V D LRVV+ K+ AL + CY
Sbjct: 505 EVTLSSRLKSLYSIYSKMKRKDTSIDKVYDARALRVVVGDKN--GALHGPAVQCCYSLLD 678
Query: 366 IIQSMWKEIPSRTKDYISRPKGNGYRSLHMAVDVSEIGRTRPLMEIQIRTTEM 418
I+ +W I DYI PK +GY+SLH AV+ G + +QI T M
Sbjct: 679 IVHRLWTPIDGEFDDYIINPKPSGYQSLHTAVE----GPDNSPLXVQIXTQRM 825
>TC77934 similar to GP|9965747|gb|AAG10150.1| calmodulin-like MSS3
{Arabidopsis thaliana}, partial (80%)
Length = 1325
Score = 52.4 bits (124), Expect = 5e-07
Identities = 27/90 (30%), Positives = 53/90 (58%), Gaps = 2/90 (2%)
Frame = +1
Query: 478 RVFRLLDKNGDGKISIEELTEVIEELG--APGEDAHDMMLLLDSNSDGSLSSDEFQMFQK 535
RVF++ D+N DG+I+ +EL + +E LG P ++ M+ +D N DG + +EF+ +
Sbjct: 295 RVFQMFDRNDDGRITKKELNDSLENLGIFIPDKELSQMIEKIDVNRDGCVDIEEFRELYE 474
Query: 536 QVEMVRNLEDRDDEYKKILDEKLHMADDSG 565
+ + +RD+E ++ + E ++ D +G
Sbjct: 475 SI-----MSERDEEEEEDMREAFNVFDQNG 549
Score = 39.3 bits (90), Expect = 0.004
Identities = 24/68 (35%), Positives = 36/68 (52%), Gaps = 4/68 (5%)
Frame = +1
Query: 472 EFDQRDRVFRLLDKNGDGKISIEELTEVIEELGAPG----EDAHDMMLLLDSNSDGSLSS 527
E D R+ F + D+NGDG IS++EL V+ LG ED M+ +D + +G +
Sbjct: 505 EEDMRE-AFNVFDQNGDGFISVDELRSVLVSLGLKQGRTVEDCKKMIGTVDVDGNGLVDY 681
Query: 528 DEFQMFQK 535
EF+ K
Sbjct: 682 KEFKQMMK 705
>TC87443 similar to GP|13397927|emb|CAC34625. putative calmodulin-related
protein {Medicago sativa}, partial (69%)
Length = 918
Score = 48.9 bits (115), Expect = 5e-06
Identities = 35/96 (36%), Positives = 47/96 (48%), Gaps = 2/96 (2%)
Frame = +3
Query: 474 DQRDRVFRLLDKNGDGKISIEELTEVIEELGA--PGEDAHDMMLLLDSNSDGSLSSDEFQ 531
D+ ++F DKNGDGKIS EL E++ LG+ E+ MM LD N DG + EF
Sbjct: 24 DEVRKIFSKFDKNGDGKISRSELKEMLLTLGSETTSEEVKRMMEELDQNGDGFIDLKEFA 203
Query: 532 MFQKQVEMVRNLEDRDDEYKKILDEKLHMADDSGLI 567
F +D E + D L+ D +GLI
Sbjct: 204 DF----HCTEPGKDESSELRDAFD--LYDLDKNGLI 293
>TC78640 homologue to GP|13397927|emb|CAC34625. putative calmodulin-related
protein {Medicago sativa}, partial (84%)
Length = 1003
Score = 48.1 bits (113), Expect = 9e-06
Identities = 28/72 (38%), Positives = 41/72 (56%), Gaps = 2/72 (2%)
Frame = +2
Query: 461 KDFPSANHKGIEFDQRDRVFRLLDKNGDGKISIEELTEVIEELGA--PGEDAHDMMLLLD 518
KD S+ + ++ ++ ++F DKNGDGKIS EL E++ LG+ E+ MM LD
Sbjct: 383 KDINSSISQAMDQEEVRKIFNKFDKNGDGKISRTELKEMMTALGSKTTTEEVTRMMEELD 562
Query: 519 SNSDGSLSSDEF 530
N DG + EF
Sbjct: 563 RNGDGYIDLKEF 598
Score = 32.0 bits (71), Expect = 0.67
Identities = 25/117 (21%), Positives = 56/117 (47%), Gaps = 16/117 (13%)
Frame = +2
Query: 454 ELAALRLKDFPSANHKGIEFDQRDRVFRLLDKNGDGKISIEELTEVIEELGAPGEDAHDM 513
+++ LK+ +A ++ R+ LD+NGDG I ++E E + G ++ +
Sbjct: 470 KISRTELKEMMTALGSKTTTEEVTRMMEELDRNGDGYIDLKEFGE-LHNGGGDTKELREA 646
Query: 514 MLLLDSNSDGSLSSDEFQMFQKQV----------EMVRNLE-DRD-----DEYKKIL 554
+ D + +G +S+ E +++ +M+ N++ D D +E+KK++
Sbjct: 647 FEMYDLDKNGLISAKELHAVMRRLGEKCSLGDCRKMIGNVDADADGNVNFEEFKKMM 817
>TC80225 similar to GP|14589311|emb|CAC43238. calcium binding protein
{Sesbania rostrata}, partial (87%)
Length = 704
Score = 47.4 bits (111), Expect = 2e-05
Identities = 34/105 (32%), Positives = 52/105 (49%), Gaps = 5/105 (4%)
Frame = +3
Query: 472 EFDQRDRVFRLLDKNGDGKISIEELTEVIEELGA--PGEDAHDMMLLLDSNSDGSLSSDE 529
+ D+ VF D NGDGKIS+ EL ++ LG+ P ++ +M LD++ DG ++ E
Sbjct: 75 DMDELKTVFTRFDTNGDGKISVTELDNILRSLGSTVPKDELQRVMEDLDTDRDGFINLAE 254
Query: 530 FQMFQKQVEM---VRNLEDRDDEYKKILDEKLHMADDSGLIQVYN 571
F F + V L + D Y K +K + + L QV N
Sbjct: 255 FAAFCRSGSADGDVSELREAFDLYDK---DKNGLISATELCQVLN 380
Score = 39.7 bits (91), Expect = 0.003
Identities = 21/54 (38%), Positives = 34/54 (62%), Gaps = 2/54 (3%)
Frame = +3
Query: 480 FRLLDKNGDGKISIEELTEVIEELG--APGEDAHDMMLLLDSNSDGSLSSDEFQ 531
F L DK+ +G IS EL +V+ LG E+ H M+ +DS+ DG+++ +EF+
Sbjct: 315 FDLYDKDKNGLISATELCQVLNTLGMKCSVEECHTMIKSVDSDGDGNVNFEEFK 476
>BF518649 weakly similar to GP|12963415|gb| regulator of gene silencing
{Nicotiana tabacum}, partial (30%)
Length = 297
Score = 46.2 bits (108), Expect = 3e-05
Identities = 25/58 (43%), Positives = 35/58 (60%), Gaps = 2/58 (3%)
Frame = +3
Query: 478 RVFRLLDKNGDGKISIEELTEVIEELGA--PGEDAHDMMLLLDSNSDGSLSSDEFQMF 533
RVF D+NGDGKIS +L + +E +GA E+A L+DS+ DG + D+F F
Sbjct: 105 RVFNHFDENGDGKISSSDLRQCVEAIGAKMSNEEAEMAEELVDSDRDGMIVLDDFVKF 278
>TC82277 weakly similar to PIR|A84532|A84532 probable calmodulin-like
protein [imported] - Arabidopsis thaliana, partial (91%)
Length = 780
Score = 46.2 bits (108), Expect = 3e-05
Identities = 29/83 (34%), Positives = 42/83 (49%), Gaps = 4/83 (4%)
Frame = +3
Query: 458 LRLKDFPSANHKGIEFDQRD--RVFRLLDKNGDGKISIEELTEVIEEL--GAPGEDAHDM 513
+ ++F A KG D FR DKNGDGKIS EE+ E++ +L ED M
Sbjct: 330 INFEEFMEAQKKGGGIRSLDIQTAFRTFDKNGDGKISAEEIKEMLWKLEERCSLEDCRRM 509
Query: 514 MLLLDSNSDGSLSSDEFQMFQKQ 536
+ +D++ DG + +EF Q
Sbjct: 510 VRAVDTDGDGMVDMNEFVAMMTQ 578
Score = 37.7 bits (86), Expect = 0.012
Identities = 22/68 (32%), Positives = 36/68 (52%), Gaps = 1/68 (1%)
Frame = +3
Query: 479 VFRLLDKNGDGKISIEELTEVIEELGA-PGEDAHDMMLLLDSNSDGSLSSDEFQMFQKQV 537
+FR++D +GDG I+ EE E ++ G D D N DG +S++E + ++
Sbjct: 294 IFRVVDLDGDGFINFEEFMEAQKKGGGIRSLDIQTAFRTFDKNGDGKISAEEIKEMLWKL 473
Query: 538 EMVRNLED 545
E +LED
Sbjct: 474 EERCSLED 497
>TC82447 similar to GP|14589311|emb|CAC43238. calcium binding protein
{Sesbania rostrata}, partial (79%)
Length = 660
Score = 46.2 bits (108), Expect = 3e-05
Identities = 41/144 (28%), Positives = 63/144 (43%), Gaps = 11/144 (7%)
Frame = +2
Query: 431 LYKAGLTNPEEAKRLKTIMLAAAELAALRLKDFPSANHKGIEFDQRDRVFRLLDKNGDGK 490
++ TNP+ K +A E + P+ + + D+ +VF D NGDGK
Sbjct: 47 IFSPNATNPKPNKLQTFKPMATNESIPVESDPTPTPSPLLGDMDELKKVFNNFDANGDGK 226
Query: 491 ISIEELTEVIEELGA---PGEDAHDMMLLLDSNSDGSLSSDEFQMFQKQVEMVRN---LE 544
IS+ EL V+ L + E+ +M L ++ DG ++ EF F + L
Sbjct: 227 ISVNELETVLRTLRSDVPQQEELRRVMEELSTDRDGFINLSEFAAFCRSDTTDGGDSALR 406
Query: 545 DRDDEYKK-----ILDEKLHMADD 563
D D Y K I E+LH+A D
Sbjct: 407 DAFDLYDKDKNGLISTEELHLALD 478
Score = 41.6 bits (96), Expect = 9e-04
Identities = 23/54 (42%), Positives = 34/54 (62%), Gaps = 2/54 (3%)
Frame = +2
Query: 480 FRLLDKNGDGKISIEELTEVIEELG--APGEDAHDMMLLLDSNSDGSLSSDEFQ 531
F L DK+ +G IS EEL ++ LG E+ DM+ +DS+ DGS++ DEF+
Sbjct: 413 FDLYDKDKNGLISTEELHLALDRLGMKCSVEECQDMIKSVDSDGDGSVNFDEFK 574
>TC88696 weakly similar to PIR|T10620|T10620 probable calcium-binding
protein F21C20.130 - Arabidopsis thaliana, partial (17%)
Length = 648
Score = 40.8 bits (94), Expect = 0.001
Identities = 21/58 (36%), Positives = 34/58 (58%), Gaps = 4/58 (6%)
Frame = +1
Query: 478 RVFRLLDKNGDGKISIEELTEVIEELG----APGEDAHDMMLLLDSNSDGSLSSDEFQ 531
+ F++ D +GDG I+ +EL V++ LG + G+D M+ D+N DG L EF+
Sbjct: 325 KTFKVFDLDGDGFITSQELECVLKRLGFLDESSGKDCRSMIRFYDTNLDGRLDFQEFK 498
Score = 34.3 bits (77), Expect = 0.14
Identities = 26/96 (27%), Positives = 44/96 (45%), Gaps = 8/96 (8%)
Frame = +1
Query: 478 RVFRLLDKNGDGKISIEELTEVIEELGAPGE--DAHDMMLLLDSNSDGSLSSDEFQMFQK 535
R+F LD N DG +S+EEL V++ + ++ L++ SL +EF F
Sbjct: 76 RIFEKLDTNCDGFVSLEELNSVLQRICNTSSQFSLEELESLVEKK---SLDFNEFLFFYN 246
Query: 536 QVEMVRNL------EDRDDEYKKILDEKLHMADDSG 565
+ +N ED +DE ++ L + + D G
Sbjct: 247 SISKEKNEENRGGDEDENDELERDLVKTFKVFDLDG 354
>TC88855 PIR|A30900|MCBH calmodulin - barley, complete
Length = 889
Score = 40.0 bits (92), Expect = 0.002
Identities = 28/90 (31%), Positives = 46/90 (51%), Gaps = 2/90 (2%)
Frame = +1
Query: 480 FRLLDKNGDGKISIEELTEVIEELGAPGEDA--HDMMLLLDSNSDGSLSSDEFQMFQKQV 537
F L DK+GDG I+ +EL V+ LG +A DM+ +D++ +G++ EF
Sbjct: 124 FSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDFPEFLNL---- 291
Query: 538 EMVRNLEDRDDEYKKILDEKLHMADDSGLI 567
M R ++D D E + ++ D +G I
Sbjct: 292 -MARKMKDTDSEEELKEAFRVFDKDQNGFI 378
>TC76783 calmodulin 1 [Medicago truncatula]
Length = 957
Score = 40.0 bits (92), Expect = 0.002
Identities = 28/90 (31%), Positives = 46/90 (51%), Gaps = 2/90 (2%)
Frame = +3
Query: 480 FRLLDKNGDGKISIEELTEVIEELGAPGEDA--HDMMLLLDSNSDGSLSSDEFQMFQKQV 537
F L DK+GDG I+ +EL V+ LG +A DM+ +D++ +G++ EF
Sbjct: 168 FSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDFPEFLNL---- 335
Query: 538 EMVRNLEDRDDEYKKILDEKLHMADDSGLI 567
M R ++D D E + ++ D +G I
Sbjct: 336 -MARKMKDTDSEEELKEAFRVFDKDQNGFI 422
>TC77023 calmodulin 2 [Medicago truncatula]
Length = 999
Score = 40.0 bits (92), Expect = 0.002
Identities = 28/90 (31%), Positives = 46/90 (51%), Gaps = 2/90 (2%)
Frame = +1
Query: 480 FRLLDKNGDGKISIEELTEVIEELGAPGEDA--HDMMLLLDSNSDGSLSSDEFQMFQKQV 537
F L DK+GDG I+ +EL V+ LG +A DM+ +D++ +G++ EF
Sbjct: 133 FSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDFPEFLNL---- 300
Query: 538 EMVRNLEDRDDEYKKILDEKLHMADDSGLI 567
M R ++D D E + ++ D +G I
Sbjct: 301 -MARKMKDTDSEEELKEAFRVFDKDQNGFI 387
>BF520634 weakly similar to GP|17428592|emb PROBABLE GTP PYROPHOSPHOKINASE
(ATP:GTP 3'-PYROPHOSPHOTRANSFERASE) PROTEIN {Ralstonia
solanacearum}, partial (6%)
Length = 542
Score = 38.9 bits (89), Expect = 0.006
Identities = 23/53 (43%), Positives = 31/53 (58%)
Frame = +3
Query: 381 YISRPKGNGYRSLHMAVDVSEIGRTRPLMEIQIRTTEMDRLAVGGMASHSLYK 433
YI K +GY+SL AV+ + R +E+QIRT M A G+A+H LYK
Sbjct: 3 YIINAKPSGYQSLLTAVEGPDNSR----VEVQIRTQRMHEYAEHGLAAHWLYK 149
>TC84210 homologue to PIR|T07122|T07122 calmodulin CAM5 - soybean, partial
(96%)
Length = 627
Score = 38.1 bits (87), Expect = 0.009
Identities = 24/75 (32%), Positives = 42/75 (56%), Gaps = 3/75 (4%)
Frame = +1
Query: 480 FRLLDKNGDGKISIEELTEVIEEL--GAPGEDAHDMMLLLDSNSDGSLSSDEF-QMFQKQ 536
F L DK+GDG I++EEL VI L E+ +M+ +D++ +G++ EF + K+
Sbjct: 31 FCLFDKDGDGCITVEELATVIRSLDQNPTEEELQEMINEVDADGNGTIEFVEFLNLMAKK 210
Query: 537 VEMVRNLEDRDDEYK 551
++ ED + +K
Sbjct: 211 MKETDADEDLKEAFK 255
>TC84597
Length = 513
Score = 37.7 bits (86), Expect = 0.012
Identities = 40/168 (23%), Positives = 74/168 (43%), Gaps = 15/168 (8%)
Frame = +2
Query: 380 DYISRPKGNGYR----SLHMAVDVSEIGRTRPLMEI-----QIRTTEMDRLAVGGMASHS 430
D+I+ P N + S H++V++ + + L+E ++ +GG+
Sbjct: 2 DHINIPSFNSCKMAQVSSHLSVEMETLNKVLSLIEAFKAFDGDNDGSINEAELGGIMGSL 181
Query: 431 LYKAGLTNPEEAKRLKTIMLAAAELAALRLKDFPSANHKGIEFDQRDRV----FRLLDKN 486
YKA E+ R + L + +F N KG+E V F LD++
Sbjct: 182 GYKAS----EQEVRAMMQQGDKNKDGLLCISEFLEMNTKGLETGNLANVLSAAFEALDED 349
Query: 487 GDGKISIEELTEVIEELGAPG--EDAHDMMLLLDSNSDGSLSSDEFQM 532
G+ ++ EEL E + G E +++ LD + DG++S D+F++
Sbjct: 350 GNEILTGEELHEAMGNFGLALSLEKCENVVASLDMDGDGAVSLDDFRL 493
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.319 0.135 0.379
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,319,879
Number of Sequences: 36976
Number of extensions: 199496
Number of successful extensions: 1216
Number of sequences better than 10.0: 79
Number of HSP's better than 10.0 without gapping: 1184
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1209
length of query: 580
length of database: 9,014,727
effective HSP length: 101
effective length of query: 479
effective length of database: 5,280,151
effective search space: 2529192329
effective search space used: 2529192329
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 61 (28.1 bits)
Medicago: description of AC146862.24


