

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146862.19 - phase: 0
(1287 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
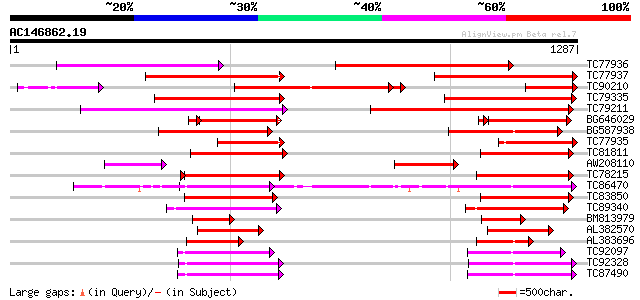
Score E
Sequences producing significant alignments: (bits) Value
TC77936 weakly similar to PIR|T52319|T52319 P-glycoprotein-like ... 747 0.0
TC77937 similar to GP|14715462|dbj|BAB62040. CjMDR1 {Coptis japo... 566 e-161
TC90210 similar to GP|14715462|dbj|BAB62040. CjMDR1 {Coptis japo... 456 e-134
TC79335 similar to GP|14715462|dbj|BAB62040. CjMDR1 {Coptis japo... 461 e-130
TC79211 similar to GP|4204793|gb|AAD10836.1| P-glycoprotein {Sol... 412 e-115
BG646029 similar to GP|14715462|d CjMDR1 {Coptis japonica}, part... 314 8e-91
BG587938 homologue to GP|22655312|g putative ABC transporter {Ar... 321 1e-87
TC77935 similar to PIR|E85023|E85023 probable P-glycoprotein-lik... 302 5e-82
TC81811 similar to GP|6671365|gb|AAF23176.1| P-glycoprotein {Gos... 270 3e-72
AW208110 similar to GP|14715462|d CjMDR1 {Coptis japonica}, part... 251 2e-66
TC78215 similar to GP|10177549|dbj|BAB10828. ABC transporter-lik... 243 9e-65
TC86470 similar to PIR|F84919|F84919 glutathione-conjugate trans... 240 2e-63
TC83850 weakly similar to GP|7300071|gb|AAF55241.1| CG4225 gene ... 207 3e-53
TC89340 weakly similar to GP|17427488|emb|CAD14006. PROBABLE COM... 185 1e-46
BM813979 homologue to PIR|E85023|E85 probable P-glycoprotein-lik... 171 2e-42
AL382570 weakly similar to GP|10241621|em putative P-glycoprotei... 169 5e-42
AL383696 homologue to PIR|T47671|T4 P-glycoprotein-like - Arabid... 145 1e-34
TC92097 similar to GP|22553016|emb|CAD44995. multidrug-resistanc... 144 2e-34
TC92328 similar to PIR|T52080|T52080 multi resistance protein [i... 138 1e-32
TC87490 similar to PIR|T52080|T52080 multi resistance protein [i... 134 2e-31
>TC77936 weakly similar to PIR|T52319|T52319 P-glycoprotein-like protein pgp3
[imported] - Arabidopsis thaliana, partial (32%)
Length = 1219
Score = 747 bits (1929), Expect = 0.0
Identities = 382/405 (94%), Positives = 392/405 (96%)
Frame = +1
Query: 739 GLLVSKMISTFYKPADELRHDSKVWAIVFVAVAVASLLIIPCRFYFFGVAGGKLIQRIRK 798
GLL+SKMI+ FYKPA ELRHDSKVWAIVFVAVAVA+LLIIPCRFYFFGVAGGKLIQRIR
Sbjct: 1 GLLISKMINIFYKPAHELRHDSKVWAIVFVAVAVATLLIIPCRFYFFGVAGGKLIQRIRN 180
Query: 799 LCFEKVVHMEVSWFDDVEHSSGALGARLSTDAASVRALVGDALGLLVQNIATIIVGMVIA 858
+CFEKVVHMEVSWFD+ EHSSGALGARLSTDAASVRALVGDALGLLVQNIAT I G+VI+
Sbjct: 181 MCFEKVVHMEVSWFDEAEHSSGALGARLSTDAASVRALVGDALGLLVQNIATAIAGLVIS 360
Query: 859 FQASWQLAFIVLALAPLLGLNGYVQVKVLKGFSADAKKLYEEASQVANDAVGSIRTVSSF 918
FQASWQLAFIVLALAPLLGLNGYVQVKVLKGFSADAKKLYEEASQVANDAVGSIRTV+SF
Sbjct: 361 FQASWQLAFIVLALAPLLGLNGYVQVKVLKGFSADAKKLYEEASQVANDAVGSIRTVASF 540
Query: 919 CAEEKVMELYKQKCEGPIKKGVRRGIISGLGFGSSFFMLYAVDACVFYAGARLVEDGKST 978
CAE+KVMELYKQKCEGPIKKGVRRGIISG GFG SFFMLYAVDACVFYAGARLVEDGKST
Sbjct: 541 CAEKKVMELYKQKCEGPIKKGVRRGIISGFGFGLSFFMLYAVDACVFYAGARLVEDGKST 720
Query: 979 FSDVFLVFFALSMAAMGVSQSGTLVPDSTNAKSAAASIFAILDQKSQIDSSDESGMTLEE 1038
FSDVFLVFFAL+MAAMGVSQSGTLVPDSTNAKSAAASIFAILD SQIDSSDESGMTLEE
Sbjct: 721 FSDVFLVFFALTMAAMGVSQSGTLVPDSTNAKSAAASIFAILDHNSQIDSSDESGMTLEE 900
Query: 1039 VKGDIEFNHVSFKYPTRLDVQIFNDLCLNIRSGKTVALVGESGSGKSTVISLLQRFYDPD 1098
VKGDIEFNHVSFKYPT LDVQIFNDLCLNIRSGKTVALVGESGSGKSTVISLLQRFYDPD
Sbjct: 901 VKGDIEFNHVSFKYPTTLDVQIFNDLCLNIRSGKTVALVGESGSGKSTVISLLQRFYDPD 1080
Query: 1099 SGHITLDGIEIQRMQVKWLRQQMGLVSQEPILFNDTVRANIAYGK 1143
SGHITLDGIEIQRMQVKWLRQQMGLVSQEPILFNDTV ANIAY K
Sbjct: 1081 SGHITLDGIEIQRMQVKWLRQQMGLVSQEPILFNDTVXANIAYXK 1215
Score = 254 bits (648), Expect = 2e-67
Identities = 138/381 (36%), Positives = 223/381 (58%), Gaps = 2/381 (0%)
Frame = +1
Query: 107 SLKFVYLAAGTFVASFLQLTCWMITGERQSARIRGLYLKTILRQDVSFFDK-ETNTGEVV 165
++ FV +A T + + + + G + RIR + + ++ +VS+FD+ E ++G +
Sbjct: 76 AIVFVAVAVATLLIIPCRFYFFGVAGGKLIQRIRNMCFEKVVHMEVSWFDEAEHSSGALG 255
Query: 166 GRMSGDTVLIKDAMGEKVGQFIQFMSTFIGGFVIAFTKGWLLTVVMLSSIPLLILSGSMT 225
R+S D ++ +G+ +G +Q ++T I G VI+F W L ++L+ PLL L+G +
Sbjct: 256 ARLSTDAASVRALVGDALGLLVQNIATAIAGLVISFQASWQLAFIVLALAPLLGLNGYVQ 435
Query: 226 SMVIAKASSTGQAAYSKSAGVVEQTIGSIRTVASFTGEKQATANYNRSLIKVYKTAVQEA 285
V+ S+ + Y +++ V +GSIRTVASF EK+ Y + K V+
Sbjct: 436 VKVLKGFSADAKKLYEEASQVANDAVGSIRTVASFCAEKKVMELYKQKCEGPIKKGVRRG 615
Query: 286 LASGVGFGTLFFVFICSYGLAVWFGGKMIIEKGY-TGGDVMTVIFAVLIGSTCLGQTSPS 344
+ SG GFG FF+ + + V++ G ++E G T DV V FA+ + + + Q+
Sbjct: 616 IISGFGFGLSFFM-LYAVDACVFYAGARLVEDGKSTFSDVFLVFFALTMAAMGVSQSGTL 792
Query: 345 LSAFAAGQAAAFKMFETINRKPEIDAYDTSGKKLDDIRGDIELRDVCFSYPTRPDELIFN 404
+ ++AA +F ++ +ID+ D SG L++++GDIE V F YPT D IFN
Sbjct: 793 VPDSTNAKSAAASIFAILDHNSQIDSSDESGMTLEEVKGDIEFNHVSFKYPTTLDVQIFN 972
Query: 405 GFSLSLPSGTTAALVGQSGSGKSTVVSLIERFYDPTDGEVLIDGINLKEFQLKWIRQKIG 464
L++ SG T ALVG+SGSGKSTV+SL++RFYDP G + +DGI ++ Q+KW+RQ++G
Sbjct: 973 DLCLNIRSGKTVALVGESGSGKSTVISLLQRFYDPDSGHITLDGIEIQRMQVKWLRQQMG 1152
Query: 465 LVSQEPVLFTCSIKENIAYGK 485
LVSQEP+LF ++ NIAY K
Sbjct: 1153LVSQEPILFNDTVXANIAYXK 1215
>TC77937 similar to GP|14715462|dbj|BAB62040. CjMDR1 {Coptis japonica},
partial (24%)
Length = 1164
Score = 567 bits (1460), Expect = e-161
Identities = 290/323 (89%), Positives = 310/323 (95%)
Frame = +3
Query: 965 FYAGARLVEDGKSTFSDVFLVFFALSMAAMGVSQSGTLVPDSTNAKSAAASIFAILDQKS 1024
FYAGARLVEDGKS+FSDVF VFFALSMAA+G+SQSG+L+PDST AKSA ASIFAILD+KS
Sbjct: 15 FYAGARLVEDGKSSFSDVFRVFFALSMAAIGLSQSGSLLPDSTKAKSAVASIFAILDRKS 194
Query: 1025 QIDSSDESGMTLEEVKGDIEFNHVSFKYPTRLDVQIFNDLCLNIRSGKTVALVGESGSGK 1084
ID +DESG+TLEEVKG+IEF HV+FKYPTR D+QIF DLCLNI SGKTVALVGESGSGK
Sbjct: 195 LIDPTDESGITLEEVKGEIEFKHVNFKYPTRPDIQIFRDLCLNIHSGKTVALVGESGSGK 374
Query: 1085 STVISLLQRFYDPDSGHITLDGIEIQRMQVKWLRQQMGLVSQEPILFNDTVRANIAYGKG 1144
STVISL+QRFYDPDSGHITLDG EIQ +QVKWLRQQMGLVSQEP+LFNDT+RANIAYGKG
Sbjct: 375 STVISLIQRFYDPDSGHITLDGKEIQSLQVKWLRQQMGLVSQEPVLFNDTIRANIAYGKG 554
Query: 1145 GDATEAEIVAAAELANAHQFIGSLQKGYDTIVGERGIQLSGGQKQRVAIARAIVKNPKIL 1204
GDA+EAEI+AAAELANAH+FI SLQKGYDT+VGERG+QLSGGQKQRVAIARAIVKNPKIL
Sbjct: 555 GDASEAEIIAAAELANAHKFISSLQKGYDTVVGERGVQLSGGQKQRVAIARAIVKNPKIL 734
Query: 1205 LLDEATSALDAESEKVVQDALDRVMVERTTIIVAHRLSTIKGADLIAVVKNGVIAEKGKH 1264
LLDEATSALDAESEKVVQDALDRVMVERTTIIVAHRLSTIKGADLIAVVKNGVIAEKGKH
Sbjct: 735 LLDEATSALDAESEKVVQDALDRVMVERTTIIVAHRLSTIKGADLIAVVKNGVIAEKGKH 914
Query: 1265 EALLHKGGDYASLVALHTSDSTS 1287
EALLHKGGDYASLVALHTS STS
Sbjct: 915 EALLHKGGDYASLVALHTSASTS 983
Score = 320 bits (821), Expect = 2e-87
Identities = 170/317 (53%), Positives = 225/317 (70%), Gaps = 2/317 (0%)
Frame = +3
Query: 308 WFGGKMIIEKGYTG-GDVMTVIFAVLIGSTCLGQTSPSLSAFAAGQAAAFKMFETINRKP 366
++ G ++E G + DV V FA+ + + L Q+ L ++A +F ++RK
Sbjct: 15 FYAGARLVEDGKSSFSDVFRVFFALSMAAIGLSQSGSLLPDSTKAKSAVASIFAILDRKS 194
Query: 367 EIDAYDTSGKKLDDIRGDIELRDVCFSYPTRPDELIFNGFSLSLPSGTTAALVGQSGSGK 426
ID D SG L++++G+IE + V F YPTRPD IF L++ SG T ALVG+SGSGK
Sbjct: 195 LIDPTDESGITLEEVKGEIEFKHVNFKYPTRPDIQIFRDLCLNIHSGKTVALVGESGSGK 374
Query: 427 STVVSLIERFYDPTDGEVLIDGINLKEFQLKWIRQKIGLVSQEPVLFTCSIKENIAYGKD 486
STV+SLI+RFYDP G + +DG ++ Q+KW+RQ++GLVSQEPVLF +I+ NIAYGK
Sbjct: 375 STVISLIQRFYDPDSGHITLDGKEIQSLQVKWLRQQMGLVSQEPVLFNDTIRANIAYGKG 554
Query: 487 C-ATDEEIRVAAELANAAKFIDKLPQGLDTMVGEHGTQLSGGQKQRVAIARAILKDPRIL 545
A++ EI AAELANA KFI L +G DT+VGE G QLSGGQKQRVAIARAI+K+P+IL
Sbjct: 555 GDASEAEIIAAAELANAHKFISSLQKGYDTVVGERGVQLSGGQKQRVAIARAIVKNPKIL 734
Query: 546 LLDEATSALDAESERIVQEALNRIMINRTTIVVAHRLSTIRNVDTIAVIHQGKIVERGSH 605
LLDEATSALDAESE++VQ+AL+R+M+ RTTI+VAHRLSTI+ D IAV+ G I E+G H
Sbjct: 735 LLDEATSALDAESEKVVQDALDRVMVERTTIIVAHRLSTIKGADLIAVVKNGVIAEKGKH 914
Query: 606 AELTNDPNGAYSQLIRL 622
L + G Y+ L+ L
Sbjct: 915 EALLH-KGGDYASLVAL 962
>TC90210 similar to GP|14715462|dbj|BAB62040. CjMDR1 {Coptis japonica},
partial (26%)
Length = 1158
Score = 456 bits (1173), Expect(2) = e-134
Identities = 236/362 (65%), Positives = 289/362 (79%), Gaps = 1/362 (0%)
Frame = +2
Query: 510 PQGLDTMVGEHGTQLSGGQKQRVAIARAILKDPRILLLDEATSALDAESERIVQEALNRI 569
PQGLDTMVG+HGTQLSGGQKQR+AIARAILK+PRILLLDEATSALDAESER+VQEAL+RI
Sbjct: 2 PQGLDTMVGDHGTQLSGGQKQRIAIARAILKNPRILLLDEATSALDAESERVVQEALDRI 181
Query: 570 MINRTTIVVAHRLSTIRNVDTIAVIHQGKIVERGSHAELTNDPNGAYSQLIRLQEMKRSE 629
M+NRTT+VVAHRLST+RN D IAVIH+GK+VE+G+H+EL DP GAYSQLIRLQE+ +
Sbjct: 182 MVNRTTVVVAHRLSTVRNADMIAVIHRGKMVEKGTHSELLKDPEGAYSQLIRLQEVNKES 361
Query: 630 QNDANDKNKPNSIVHSGRQSSQRSFSLRSISQGSA-GNSGRHSFSASYVAPTTDGFLETE 688
+ + K S RQSSQR RSIS+GS+ GNS RHSFS S+ PT +
Sbjct: 362 EETTDHHGKRELSAESFRQSSQRKSLQRSISRGSSIGNSSRHSFSVSFGLPTG---VNVA 532
Query: 689 DGGPQASPSKNSSPPEVPLYRLAYFNKPEIPVLLMGTITAVLHGAIMPVIGLLVSKMIST 748
D + P+K EVPL RLA NKPEIPVLL+G++ A+ +G I+P+ G+L+S +I T
Sbjct: 533 DPDLEKVPTKEKEQ-EVPLRRLASLNKPEIPVLLIGSLAAIANGVILPIFGVLISSVIKT 709
Query: 749 FYKPADELRHDSKVWAIVFVAVAVASLLIIPCRFYFFGVAGGKLIQRIRKLCFEKVVHME 808
FY+P DE++ DSK WAI+F+ + +ASL++IP R YFF VAG KLIQRIR LCFEKVV+ME
Sbjct: 710 FYEPFDEMKKDSKFWAIMFMLLGLASLVVIPARGYFFSVAGCKLIQRIRLLCFEKVVNME 889
Query: 809 VSWFDDVEHSSGALGARLSTDAASVRALVGDALGLLVQNIATIIVGMVIAFQASWQLAFI 868
V W+D+ E+SSGA+GARLS DAASVRALVGDALGLLVQN+A+ + G++IAF ASWQLA I
Sbjct: 890 VGWYDEPENSSGAVGARLSADAASVRALVGDALGLLVQNLASALAGLIIAFIASWQLALI 1069
Query: 869 VL 870
+L
Sbjct: 1070IL 1075
Score = 149 bits (377), Expect = 5e-36
Identities = 76/119 (63%), Positives = 95/119 (78%), Gaps = 1/119 (0%)
Frame = +2
Query: 1170 KGYDTIVGERGIQLSGGQKQRVAIARAIVKNPKILLLDEATSALDAESEKVVQDALDRVM 1229
+G DT+VG+ G QLSGGQKQR+AIARAI+KNP+ILLLDEATSALDAESE+VVQ+ALDR+M
Sbjct: 5 QGLDTMVGDHGTQLSGGQKQRIAIARAILKNPRILLLDEATSALDAESERVVQEALDRIM 184
Query: 1230 VERTTIIVAHRLSTIKGADLIAVVKNGVIAEKGKHEALL-HKGGDYASLVALHTSDSTS 1287
V RTT++VAHRLST++ AD+IAV+ G + EKG H LL G Y+ L+ L + S
Sbjct: 185 VNRTTVVVAHRLSTVRNADMIAVIHRGKMVEKGTHSELLKDPEGAYSQLIRLQEVNKES 361
Score = 72.4 bits (176), Expect = 1e-12
Identities = 50/197 (25%), Positives = 101/197 (50%), Gaps = 1/197 (0%)
Frame = +2
Query: 17 VEDHDSNQDSEKSKDKDVTTKTVPLYKLFSFADPSDRLLMLMGTLGAIGNGLSIPLMILI 76
V D D + K K+++V PL +L S P +L L+G+L AI NG+ +P+ ++
Sbjct: 527 VADPDLEKVPTKEKEQEV-----PLRRLASLNKPEIPVL-LIGSLAAIANGVILPIFGVL 688
Query: 77 FGTMINAFGDSTNSKVVDEVSETTTYCDQVSLKFVYLAAGTFVASFLQLTCWMITGERQS 136
++I F + DE+ + + + ++ F+ L + V + + + G +
Sbjct: 689 ISSVIKTFYEP-----FDEMKKDSKFW---AIMFMLLGLASLVVIPARGYFFSVAGCKLI 844
Query: 137 ARIRGLYLKTILRQDVSFFDKETNTGEVVG-RMSGDTVLIKDAMGEKVGQFIQFMSTFIG 195
RIR L + ++ +V ++D+ N+ VG R+S D ++ +G+ +G +Q +++ +
Sbjct: 845 QRIRLLCFEKVVNMEVGWYDEPENSSGAVGARLSADAASVRALVGDALGLLVQNLASALA 1024
Query: 196 GFVIAFTKGWLLTVVML 212
G +IAF W L +++L
Sbjct: 1025GLIIAFIASWQLALIIL 1075
Score = 44.3 bits (103), Expect(2) = e-134
Identities = 21/36 (58%), Positives = 27/36 (74%)
Frame = +3
Query: 862 SWQLAFIVLALAPLLGLNGYVQVKVLKGFSADAKKL 897
SW+ F L + PL+GLNGYVQ+K +KGFS DAK +
Sbjct: 1056 SWR*LF--LCVIPLIGLNGYVQMKFMKGFSGDAKMM 1157
>TC79335 similar to GP|14715462|dbj|BAB62040. CjMDR1 {Coptis japonica},
partial (23%)
Length = 1135
Score = 461 bits (1187), Expect = e-130
Identities = 234/301 (77%), Positives = 270/301 (88%), Gaps = 1/301 (0%)
Frame = +2
Query: 987 FALSMAAMGVSQSGTLVPDSTNAKSAAASIFAILDQKSQIDSSDESGMTLEEVKGDIEFN 1046
FAL+MAA+G+SQS + PDS+ AKSA ASIF ++D+KS+ID S+ESG TL+ +KG+IE
Sbjct: 2 FALTMAAIGISQSSSFAPDSSKAKSATASIFGMIDKKSKIDPSEESGTTLDSIKGEIELR 181
Query: 1047 HVSFKYPTRLDVQIFNDLCLNIRSGKTVALVGESGSGKSTVISLLQRFYDPDSGHITLDG 1106
H+SFKYP+R D+QIF DL L I SGKTVALVGESGSGKSTVI+LLQRFYDPDSG ITLDG
Sbjct: 182 HISFKYPSRPDIQIFRDLNLTIHSGKTVALVGESGSGKSTVIALLQRFYDPDSGEITLDG 361
Query: 1107 IEIQRMQVKWLRQQMGLVSQEPILFNDTVRANIAYGKGGDATEAEIVAAAELANAHQFIG 1166
IEI+++Q+KWLRQQMGLVSQEP+LFNDT+RANIAYGKGG ATEAEI+AAAELANAH+FI
Sbjct: 362 IEIRQLQLKWLRQQMGLVSQEPVLFNDTIRANIAYGKGGIATEAEIIAAAELANAHRFIS 541
Query: 1167 SLQKGYDTIVGERGIQLSGGQKQRVAIARAIVKNPKILLLDEATSALDAESEKVVQDALD 1226
LQ+GYDTIVGERG QLSGGQKQRVAIARAI+K+PKILLLDEATSALDAESE+VVQDALD
Sbjct: 542 GLQQGYDTIVGERGTQLSGGQKQRVAIARAIIKSPKILLLDEATSALDAESERVVQDALD 721
Query: 1227 RVMVERTTIIVAHRLSTIKGADLIAVVKNGVIAEKGKHEALLH-KGGDYASLVALHTSDS 1285
+VMV RTT++VAHRLSTIK AD+IAVVKNGVI EKG+HE L++ K G YASLV LHTS
Sbjct: 722 KVMVNRTTVVVAHRLSTIKNADVIAVVKNGVIVEKGRHETLINVKDGFYASLVQLHTSAK 901
Query: 1286 T 1286
T
Sbjct: 902 T 904
Score = 336 bits (862), Expect = 3e-92
Identities = 171/295 (57%), Positives = 225/295 (75%), Gaps = 1/295 (0%)
Frame = +2
Query: 329 FAVLIGSTCLGQTSPSLSAFAAGQAAAFKMFETINRKPEIDAYDTSGKKLDDIRGDIELR 388
FA+ + + + Q+S + ++A +F I++K +ID + SG LD I+G+IELR
Sbjct: 2 FALTMAAIGISQSSSFAPDSSKAKSATASIFGMIDKKSKIDPSEESGTTLDSIKGEIELR 181
Query: 389 DVCFSYPTRPDELIFNGFSLSLPSGTTAALVGQSGSGKSTVVSLIERFYDPTDGEVLIDG 448
+ F YP+RPD IF +L++ SG T ALVG+SGSGKSTV++L++RFYDP GE+ +DG
Sbjct: 182 HISFKYPSRPDIQIFRDLNLTIHSGKTVALVGESGSGKSTVIALLQRFYDPDSGEITLDG 361
Query: 449 INLKEFQLKWIRQKIGLVSQEPVLFTCSIKENIAYGKD-CATDEEIRVAAELANAAKFID 507
I +++ QLKW+RQ++GLVSQEPVLF +I+ NIAYGK AT+ EI AAELANA +FI
Sbjct: 362 IEIRQLQLKWLRQQMGLVSQEPVLFNDTIRANIAYGKGGIATEAEIIAAAELANAHRFIS 541
Query: 508 KLPQGLDTMVGEHGTQLSGGQKQRVAIARAILKDPRILLLDEATSALDAESERIVQEALN 567
L QG DT+VGE GTQLSGGQKQRVAIARAI+K P+ILLLDEATSALDAESER+VQ+AL+
Sbjct: 542 GLQQGYDTIVGERGTQLSGGQKQRVAIARAIIKSPKILLLDEATSALDAESERVVQDALD 721
Query: 568 RIMINRTTIVVAHRLSTIRNVDTIAVIHQGKIVERGSHAELTNDPNGAYSQLIRL 622
++M+NRTT+VVAHRLSTI+N D IAV+ G IVE+G H L N +G Y+ L++L
Sbjct: 722 KVMVNRTTVVVAHRLSTIKNADVIAVVKNGVIVEKGRHETLINVKDGFYASLVQL 886
>TC79211 similar to GP|4204793|gb|AAD10836.1| P-glycoprotein {Solanum
tuberosum}, partial (36%)
Length = 1667
Score = 412 bits (1060), Expect = e-115
Identities = 215/465 (46%), Positives = 317/465 (67%), Gaps = 3/465 (0%)
Frame = +1
Query: 819 SGALGARLSTDAASVRALVGDALGLLVQNIATIIVGMVIAFQASWQLAFIVLALAPLLGL 878
S + ARL+ DA +VR+ +GD + ++VQN A ++V F W+LA +++A+ P++
Sbjct: 7 SARISARLALDANNVRSAIGDRISVIVQNTALMLVACTAGFVLQWRLALVLIAVFPVVVA 186
Query: 879 NGYVQVKVLKGFSADAKKLYEEASQVANDAVGSIRTVSSFCAEEKVMELYKQKCEGPIKK 938
+Q + GFS D + + +A+Q+A +A+ ++RTV++F +E K++ L+ E P+++
Sbjct: 187 ATVLQKMFMTGFSGDLEAAHAKATQLAGEAIANVRTVAAFNSESKIVRLFASNLETPLQR 366
Query: 939 GVRRGIISGLGFGSSFFMLYAVDACVFYAGARLVEDGKSTFSDVFLVFFALSMAAMGVSQ 998
+G ISG G+G + F LYA A + + LV+ G S FS VF L ++A G ++
Sbjct: 367 CFWKGQISGSGYGIAQFALYASYALGLWYASWLVKHGISDFSKTIRVFMVLMVSANGAAE 546
Query: 999 SGTLVPDSTNAKSAAASIFAILDQKSQIDSSDESGMTL-EEVKGDIEFNHVSFKYPTRLD 1057
+ TL PD A S+F +LD++++I+ D+ + + ++G++E HV F YPTR D
Sbjct: 547 TLTLAPDFIKGGRAMRSVFDLLDRQTEIEPDDQDATPVPDRLRGEVELKHVDFSYPTRPD 726
Query: 1058 VQIFNDLCLNIRSGKTVALVGESGSGKSTVISLLQRFYDPDSGHITLDGIEIQRMQVKWL 1117
+ +F DL L IR+GKT+ALVG SG GKS+VI+L+QRFYDP SG I +DG +I++ +K L
Sbjct: 727 MPVFRDLNLRIRAGKTLALVGPSGCGKSSVIALIQRFYDPTSGRIMIDGKDIRKYNLKSL 906
Query: 1118 RQQMGLVSQEPILFNDTVRANIAYGKGGDATEAEIVAAAELANAHQFIGSLQKGYDTIVG 1177
R+ + +V QEP LF T+ NIAYG ATEAEI+ AA LANAH+FI SL GY T VG
Sbjct: 907 RRHISVVPQEPCLFATTIYENIAYGHDS-ATEAEIIEAATLANAHKFISSLPDGYKTFVG 1083
Query: 1178 ERGIQLSGGQKQRVAIARAIVKNPKILLLDEATSALDAESEKVVQDALDRVMVERTTIIV 1237
ERG+QLSGGQKQR+A+ARA ++ +++LLDEATSALDAESE+ VQ+ALDR +TTIIV
Sbjct: 1084 ERGVQLSGGQKQRIAVARAFLRKAELMLLDEATSALDAESERSVQEALDRASTGKTTIIV 1263
Query: 1238 AHRLSTIKGADLIAVVKNGVIAEKGKHEALL--HKGGDYASLVAL 1280
AHRLSTI+ A++IAV+ +G +AE+G H L+ H+ G YA ++ L
Sbjct: 1264 AHRLSTIRNANVIAVIDDGKVAEQGSHSQLMKNHQDGIYARMIQL 1398
Score = 350 bits (899), Expect = 2e-96
Identities = 187/471 (39%), Positives = 285/471 (59%), Gaps = 2/471 (0%)
Frame = +1
Query: 161 TGEVVGRMSGDTVLIKDAMGEKVGQFIQFMSTFIGGFVIAFTKGWLLTVVMLSSIPLLIL 220
+ + R++ D ++ A+G+++ +Q + + F W L +V+++ P+++
Sbjct: 7 SARISARLALDANNVRSAIGDRISVIVQNTALMLVACTAGFVLQWRLALVLIAVFPVVVA 186
Query: 221 SGSMTSMVIAKASSTGQAAYSKSAGVVEQTIGSIRTVASFTGEKQATANYNRSLIKVYKT 280
+ + M + S +AA++K+ + + I ++RTVA+F E + + +L +
Sbjct: 187 ATVLQKMFMTGFSGDLEAAHAKATQLAGEAIANVRTVAAFNSESKIVRLFASNLETPLQR 366
Query: 281 AVQEALASGVGFGTLFFVFICSYGLAVWFGGKMIIEKGYTGGDVMTVIFAVLIGSTCLGQ 340
+ SG G+G F SY L +W+ ++ + V +++ + +
Sbjct: 367 CFWKGQISGSGYGIAQFALYASYALGLWYASWLVKHGISDFSKTIRVFMVLMVSANGAAE 546
Query: 341 TSPSLSAFAAGQAAAFKMFETINRKPEIDAYDTSGKKLDD-IRGDIELRDVCFSYPTRPD 399
T F G A +F+ ++R+ EI+ D + D +RG++EL+ V FSYPTRPD
Sbjct: 547 TLTLAPDFIKGGRAMRSVFDLLDRQTEIEPDDQDATPVPDRLRGEVELKHVDFSYPTRPD 726
Query: 400 ELIFNGFSLSLPSGTTAALVGQSGSGKSTVVSLIERFYDPTDGEVLIDGINLKEFQLKWI 459
+F +L + +G T ALVG SG GKS+V++LI+RFYDPT G ++IDG +++++ LK +
Sbjct: 727 MPVFRDLNLRIRAGKTLALVGPSGCGKSSVIALIQRFYDPTSGRIMIDGKDIRKYNLKSL 906
Query: 460 RQKIGLVSQEPVLFTCSIKENIAYGKDCATDEEIRVAAELANAAKFIDKLPQGLDTMVGE 519
R+ I +V QEP LF +I ENIAYG D AT+ EI AA LANA KFI LP G T VGE
Sbjct: 907 RRHISVVPQEPCLFATTIYENIAYGHDSATEAEIIEAATLANAHKFISSLPDGYKTFVGE 1086
Query: 520 HGTQLSGGQKQRVAIARAILKDPRILLLDEATSALDAESERIVQEALNRIMINRTTIVVA 579
G QLSGGQKQR+A+ARA L+ ++LLDEATSALDAESER VQEAL+R +TTI+VA
Sbjct: 1087RGVQLSGGQKQRIAVARAFLRKAELMLLDEATSALDAESERSVQEALDRASTGKTTIIVA 1266
Query: 580 HRLSTIRNVDTIAVIHQGKIVERGSHAEL-TNDPNGAYSQLIRLQEMKRSE 629
HRLSTIRN + IAVI GK+ E+GSH++L N +G Y+++I+LQ +E
Sbjct: 1267HRLSTIRNANVIAVIDDGKVAEQGSHSQLMKNHQDGIYARMIQLQRFTHNE 1419
>BG646029 similar to GP|14715462|d CjMDR1 {Coptis japonica}, partial (16%)
Length = 680
Score = 314 bits (804), Expect(2) = 8e-91
Identities = 158/188 (84%), Positives = 174/188 (92%)
Frame = -3
Query: 1088 ISLLQRFYDPDSGHITLDGIEIQRMQVKWLRQQMGLVSQEPILFNDTVRANIAYGKGGDA 1147
ISLLQRFYDPDSG I LDG EIQ++Q+KW RQQMGLVSQEP+LFNDT+RANIAYGKGG+A
Sbjct: 606 ISLLQRFYDPDSGQIKLDGTEIQKLQLKWFRQQMGLVSQEPVLFNDTIRANIAYGKGGNA 427
Query: 1148 TEAEIVAAAELANAHQFIGSLQKGYDTIVGERGIQLSGGQKQRVAIARAIVKNPKILLLD 1207
TEAE++AAAELANAH FI SLQ+GYDTIVGERGIQLSGGQKQ VAIARAIV P+ILLLD
Sbjct: 426 TEAEVIAAAELANAHNFISSLQQGYDTIVGERGIQLSGGQKQPVAIARAIVNRPRILLLD 247
Query: 1208 EATSALDAESEKVVQDALDRVMVERTTIIVAHRLSTIKGADLIAVVKNGVIAEKGKHEAL 1267
EATSALDAESEKVVQDALDRV V+RTTI+VAHRLSTIKGA+ IAVVKNGVI EKGKH+ L
Sbjct: 246 EATSALDAESEKVVQDALDRVRVDRTTIVVAHRLSTIKGANSIAVVKNGVIEEKGKHDIL 67
Query: 1268 LHKGGDYA 1275
++KGG YA
Sbjct: 66 INKGGTYA 43
Score = 224 bits (572), Expect(2) = 7e-63
Identities = 117/189 (61%), Positives = 147/189 (76%), Gaps = 1/189 (0%)
Frame = -3
Query: 430 VSLIERFYDPTDGEVLIDGINLKEFQLKWIRQKIGLVSQEPVLFTCSIKENIAYGKDC-A 488
+SL++RFYDP G++ +DG +++ QLKW RQ++GLVSQEPVLF +I+ NIAYGK A
Sbjct: 606 ISLLQRFYDPDSGQIKLDGTEIQKLQLKWFRQQMGLVSQEPVLFNDTIRANIAYGKGGNA 427
Query: 489 TDEEIRVAAELANAAKFIDKLPQGLDTMVGEHGTQLSGGQKQRVAIARAILKDPRILLLD 548
T+ E+ AAELANA FI L QG DT+VGE G QLSGGQKQ VAIARAI+ PRILLLD
Sbjct: 426 TEAEVIAAAELANAHNFISSLQQGYDTIVGERGIQLSGGQKQPVAIARAIVNRPRILLLD 247
Query: 549 EATSALDAESERIVQEALNRIMINRTTIVVAHRLSTIRNVDTIAVIHQGKIVERGSHAEL 608
EATSALDAESE++VQ+AL+R+ ++RTTIVVAHRLSTI+ ++IAV+ G I E+G H L
Sbjct: 246 EATSALDAESEKVVQDALDRVRVDRTTIVVAHRLSTIKGANSIAVVKNGVIEEKGKHDIL 67
Query: 609 TNDPNGAYS 617
N G Y+
Sbjct: 66 IN-KGGTYA 43
Score = 40.0 bits (92), Expect(2) = 8e-91
Identities = 20/24 (83%), Positives = 21/24 (87%)
Frame = -2
Query: 1064 LCLNIRSGKTVALVGESGSGKSTV 1087
L L I SG+TVALVGESGSGKSTV
Sbjct: 679 LSLTIHSGQTVALVGESGSGKSTV 608
Score = 36.2 bits (82), Expect(2) = 7e-63
Identities = 17/29 (58%), Positives = 23/29 (78%)
Frame = -2
Query: 407 SLSLPSGTTAALVGQSGSGKSTVVSLIER 435
SL++ SG T ALVG+SGSGKSTV ++ +
Sbjct: 676 SLTIHSGQTVALVGESGSGKSTVDLIVAK 590
>BG587938 homologue to GP|22655312|g putative ABC transporter {Arabidopsis
thaliana}, partial (19%)
Length = 780
Score = 321 bits (822), Expect = 1e-87
Identities = 162/259 (62%), Positives = 205/259 (78%)
Frame = +3
Query: 338 LGQTSPSLSAFAAGQAAAFKMFETINRKPEIDAYDTSGKKLDDIRGDIELRDVCFSYPTR 397
LGQ++PS++AF + AA K+F I+ +P ID SG +L+ + G +EL++V FSYP+R
Sbjct: 3 LGQSAPSMAAFTKARVAAAKIFRIIDHQPGIDRNSESGLELETVTGLVELKNVDFSYPSR 182
Query: 398 PDELIFNGFSLSLPSGTTAALVGQSGSGKSTVVSLIERFYDPTDGEVLIDGINLKEFQLK 457
P+ LI N FSLS+P+G T ALVG SGSGKSTVVSLIERFYDPT G+V++DG ++K +LK
Sbjct: 183 PEVLILNDFSLSVPAGKTIALVGSSGSGKSTVVSLIERFYDPTSGQVMLDGHDIKTLKLK 362
Query: 458 WIRQKIGLVSQEPVLFTCSIKENIAYGKDCATDEEIRVAAELANAAKFIDKLPQGLDTMV 517
W+RQ+IGLVSQEP LF +I+ENI G+ A EI AA +ANA FI KLP+G +T V
Sbjct: 363 WLRQQIGLVSQEPALFATTIRENILLGRPDANQVEIEEAARVANAHSFIIKLPEGFETQV 542
Query: 518 GEHGTQLSGGQKQRVAIARAILKDPRILLLDEATSALDAESERIVQEALNRIMINRTTIV 577
GE G QLSGGQKQR+AIARA+LK+P ILLLDEATSALD+ESE++VQEAL+R MI RTT+V
Sbjct: 543 GERGLQLSGGQKQRIAIARAMLKNPAILLLDEATSALDSESEKLVQEALDRFMIGRTTLV 722
Query: 578 VAHRLSTIRNVDTIAVIHQ 596
+AHRLSTIR D +AVIH+
Sbjct: 723 IAHRLSTIRKADLVAVIHK 779
Score = 302 bits (773), Expect = 6e-82
Identities = 154/258 (59%), Positives = 200/258 (76%)
Frame = +3
Query: 996 VSQSGTLVPDSTNAKSAAASIFAILDQKSQIDSSDESGMTLEEVKGDIEFNHVSFKYPTR 1055
+ QS + T A+ AAA IF I+D + ID + ESG+ LE V G +E +V F YP+R
Sbjct: 3 LGQSAPSMAAFTKARVAAAKIFRIIDHQPGIDRNSESGLELETVTGLVELKNVDFSYPSR 182
Query: 1056 LDVQIFNDLCLNIRSGKTVALVGESGSGKSTVISLLQRFYDPDSGHITLDGIEIQRMQVK 1115
+V I ND L++ +GKT+ALVG SGSGKSTV+SL++RFYDP SG + LDG +I+ +++K
Sbjct: 183 PEVLILNDFSLSVPAGKTIALVGSSGSGKSTVVSLIERFYDPTSGQVMLDGHDIKTLKLK 362
Query: 1116 WLRQQMGLVSQEPILFNDTVRANIAYGKGGDATEAEIVAAAELANAHQFIGSLQKGYDTI 1175
WLRQQ+GLVSQEP LF T+R NI G+ DA + EI AA +ANAH FI L +G++T
Sbjct: 363 WLRQQIGLVSQEPALFATTIRENILLGR-PDANQVEIEEAARVANAHSFIIKLPEGFETQ 539
Query: 1176 VGERGIQLSGGQKQRVAIARAIVKNPKILLLDEATSALDAESEKVVQDALDRVMVERTTI 1235
VGERG+QLSGGQKQR+AIARA++KNP ILLLDEATSALD+ESEK+VQ+ALDR M+ RTT+
Sbjct: 540 VGERGLQLSGGQKQRIAIARAMLKNPAILLLDEATSALDSESEKLVQEALDRFMIGRTTL 719
Query: 1236 IVAHRLSTIKGADLIAVV 1253
++AHRLSTI+ ADL+AV+
Sbjct: 720 VIAHRLSTIRKADLVAVI 773
>TC77935 similar to PIR|E85023|E85023 probable P-glycoprotein-like protein
[imported] - Arabidopsis thaliana, partial (12%)
Length = 869
Score = 302 bits (774), Expect = 5e-82
Identities = 164/178 (92%), Positives = 165/178 (92%)
Frame = +2
Query: 1110 QRMQVKWLRQQMGLVSQEPILFNDTVRANIAYGKGGDATEAEIVAAAELANAHQFIGSLQ 1169
QRMQVK + LFNDTVRANIAYGKGGDATEAEIVAAAELANAHQFIGSLQ
Sbjct: 8 QRMQVKC*DNRWDC-KPRAYLFNDTVRANIAYGKGGDATEAEIVAAAELANAHQFIGSLQ 184
Query: 1170 KGYDTIVGERGIQLSGGQKQRVAIARAIVKNPKILLLDEATSALDAESEKVVQDALDRVM 1229
KGYDTIVGERGIQLSGGQKQRVAIARAIVKNPKILLLDEATSALDAESEKVVQDALDRVM
Sbjct: 185 KGYDTIVGERGIQLSGGQKQRVAIARAIVKNPKILLLDEATSALDAESEKVVQDALDRVM 364
Query: 1230 VERTTIIVAHRLSTIKGADLIAVVKNGVIAEKGKHEALLHKGGDYASLVALHTSDSTS 1287
VERTTIIVAHRLSTIKGADLIAVVKNGVIAEKGKHEALLHKGGDYASLVALHTSDSTS
Sbjct: 365 VERTTIIVAHRLSTIKGADLIAVVKNGVIAEKGKHEALLHKGGDYASLVALHTSDSTS 538
Score = 177 bits (448), Expect = 3e-44
Identities = 95/152 (62%), Positives = 118/152 (77%), Gaps = 1/152 (0%)
Frame = +2
Query: 472 LFTCSIKENIAYGKDC-ATDEEIRVAAELANAAKFIDKLPQGLDTMVGEHGTQLSGGQKQ 530
LF +++ NIAYGK AT+ EI AAELANA +FI L +G DT+VGE G QLSGGQKQ
Sbjct: 65 LFNDTVRANIAYGKGGDATEAEIVAAAELANAHQFIGSLQKGYDTIVGERGIQLSGGQKQ 244
Query: 531 RVAIARAILKDPRILLLDEATSALDAESERIVQEALNRIMINRTTIVVAHRLSTIRNVDT 590
RVAIARAI+K+P+ILLLDEATSALDAESE++VQ+AL+R+M+ RTTI+VAHRLSTI+ D
Sbjct: 245 RVAIARAIVKNPKILLLDEATSALDAESEKVVQDALDRVMVERTTIIVAHRLSTIKGADL 424
Query: 591 IAVIHQGKIVERGSHAELTNDPNGAYSQLIRL 622
IAV+ G I E+G H L + G Y+ L+ L
Sbjct: 425 IAVVKNGVIAEKGKHEALLH-KGGDYASLVAL 517
>TC81811 similar to GP|6671365|gb|AAF23176.1| P-glycoprotein {Gossypium
hirsutum}, partial (17%)
Length = 850
Score = 270 bits (690), Expect = 3e-72
Identities = 137/220 (62%), Positives = 172/220 (77%)
Frame = +1
Query: 410 LPSGTTAALVGQSGSGKSTVVSLIERFYDPTDGEVLIDGINLKEFQLKWIRQKIGLVSQE 469
+PSG + ALVGQSGSGKS+V+SLI RFYDPT G+VLIDG ++ LK + + IGLV QE
Sbjct: 1 VPSGKSVALVGQSGSGKSSVISLILRFYDPTSGKVLIDGKDITRINLKSLTKHIGLVQQE 180
Query: 470 PVLFTCSIKENIAYGKDCATDEEIRVAAELANAAKFIDKLPQGLDTMVGEHGTQLSGGQK 529
P LF SI ENI YGK+ A+D E+ AA+LANA FI LP+G T VGE G QLSGGQ+
Sbjct: 181 PALFATSIYENILYGKEGASDSEVIEAAKLANAHNFISALPEGYSTKVGERGVQLSGGQR 360
Query: 530 QRVAIARAILKDPRILLLDEATSALDAESERIVQEALNRIMINRTTIVVAHRLSTIRNVD 589
QRVAIARA+LK+P ILLLDEATSALD ESERIVQ+AL+R+M NRTT++VAHRLSTIRN D
Sbjct: 361 QRVAIARAVLKNPEILLLDEATSALDVESERIVQQALDRLMQNRTTVMVAHRLSTIRNAD 540
Query: 590 TIAVIHQGKIVERGSHAELTNDPNGAYSQLIRLQEMKRSE 629
I+V+ GKI+E+G+H+ L + +G Y +L+ LQ+ + +
Sbjct: 541 QISVLQDGKIIEQGTHSSLIENKDGPYYKLVNLQQQQNHQ 660
Score = 253 bits (647), Expect = 3e-67
Identities = 131/214 (61%), Positives = 172/214 (80%), Gaps = 1/214 (0%)
Frame = +1
Query: 1068 IRSGKTVALVGESGSGKSTVISLLQRFYDPDSGHITLDGIEIQRMQVKWLRQQMGLVSQE 1127
+ SGK+VALVG+SGSGKS+VISL+ RFYDP SG + +DG +I R+ +K L + +GLV QE
Sbjct: 1 VPSGKSVALVGQSGSGKSSVISLILRFYDPTSGKVLIDGKDITRINLKSLTKHIGLVQQE 180
Query: 1128 PILFNDTVRANIAYGKGGDATEAEIVAAAELANAHQFIGSLQKGYDTIVGERGIQLSGGQ 1187
P LF ++ NI YGK G A+++E++ AA+LANAH FI +L +GY T VGERG+QLSGGQ
Sbjct: 181 PALFATSIYENILYGKEG-ASDSEVIEAAKLANAHNFISALPEGYSTKVGERGVQLSGGQ 357
Query: 1188 KQRVAIARAIVKNPKILLLDEATSALDAESEKVVQDALDRVMVERTTIIVAHRLSTIKGA 1247
+QRVAIARA++KNP+ILLLDEATSALD ESE++VQ ALDR+M RTT++VAHRLSTI+ A
Sbjct: 358 RQRVAIARAVLKNPEILLLDEATSALDVESERIVQQALDRLMQNRTTVMVAHRLSTIRNA 537
Query: 1248 DLIAVVKNGVIAEKGKHEALL-HKGGDYASLVAL 1280
D I+V+++G I E+G H +L+ +K G Y LV L
Sbjct: 538 DQISVLQDGKIIEQGTHSSLIENKDGPYYKLVNL 639
>AW208110 similar to GP|14715462|d CjMDR1 {Coptis japonica}, partial (11%)
Length = 449
Score = 251 bits (640), Expect = 2e-66
Identities = 130/147 (88%), Positives = 134/147 (90%)
Frame = +2
Query: 873 APLLGLNGYVQVKVLKGFSADAKKLYEEASQVANDAVGSIRTVSSFCAEEKVMELYKQKC 932
APLLGL+GYVQVK LKGFSADAKKLYEEASQVANDAVG IRTVSSFCAEEKVMELY+QKC
Sbjct: 8 APLLGLDGYVQVKFLKGFSADAKKLYEEASQVANDAVGCIRTVSSFCAEEKVMELYEQKC 187
Query: 933 EGPIKKGVRRGIISGLGFGSSFFMLYAVDACVFYAGARLVEDGKSTFSDVFLVFFALSMA 992
EGPIKKG+RRGIISGLGFG S F+LYAV AC FYAGARLVEDGKSTFSDVFLV FAL MA
Sbjct: 188 EGPIKKGIRRGIISGLGFGLSCFLLYAVYACCFYAGARLVEDGKSTFSDVFLVIFALGMA 367
Query: 993 AMGVSQSGTLVPDSTNAKSAAASIFAI 1019
A GVSQ GTLVPD NAKSA ASIFAI
Sbjct: 368 ASGVSQLGTLVPDLINAKSATASIFAI 448
Score = 61.2 bits (147), Expect = 2e-09
Identities = 40/144 (27%), Positives = 66/144 (45%), Gaps = 3/144 (2%)
Frame = +2
Query: 216 PLLILSGSMTSMVIAKASSTGQAAYSKSAGVVEQTIGSIRTVASFTGEKQATANYNRSLI 275
PLL L G + + S+ + Y +++ V +G IRTV+SF E++ Y +
Sbjct: 11 PLLGLDGYVQVKFLKGFSADAKKLYEEASQVANDAVGCIRTVSSFCAEEKVMELYEQKCE 190
Query: 276 KVYKTAVQEALASGVGFGTLFFVFICSYGLAVWFGGKMIIEKGYTGGDVMTVIFAV---L 332
K ++ + SG+GFG F+ Y + G +++ + T DV VIFA+
Sbjct: 191 GPIKKGIRRGIISGLGFGLSCFLLYAVYACCFYAGARLVEDGKSTFSDVFLVIFALGMAA 370
Query: 333 IGSTCLGQTSPSLSAFAAGQAAAF 356
G + LG P L + A+ F
Sbjct: 371 SGVSQLGTLVPDLINAKSATASIF 442
>TC78215 similar to GP|10177549|dbj|BAB10828. ABC transporter-like protein
{Arabidopsis thaliana}, partial (36%)
Length = 1176
Score = 243 bits (620), Expect(2) = 9e-65
Identities = 124/227 (54%), Positives = 166/227 (72%), Gaps = 1/227 (0%)
Frame = +1
Query: 398 PDELIFNGFSLSLPSGTTAALVGQSGSGKSTVVSLIERFYDPTDGEVLIDGINLKEFQLK 457
P ++ G ++ L G+ ALVG SG GK+T+ +LIERFYDPT G +L++G+ L E K
Sbjct: 34 P*YMVLKGINIKLQPGSKVALVGPSGGGKTTIANLIERFYDPTKGNILVNGVPLVEISHK 213
Query: 458 WIRQKIGLVSQEPVLFTCSIKENIAYGKDCAT-DEEIRVAAELANAAKFIDKLPQGLDTM 516
+ +KI +VSQEP LF CSI+ENIAYG D D +I AA++ANA +FI P+ T
Sbjct: 214 HLHRKISIVSQEPTLFNCSIEENIAYGFDGKIEDADIENAAKMANAHEFISNFPEKYKTF 393
Query: 517 VGEHGTQLSGGQKQRVAIARAILKDPRILLLDEATSALDAESERIVQEALNRIMINRTTI 576
VGE G +LSGGQKQR+AIARA+L DP+ILLLDEATSALDAESE +VQ+A++ IM RT +
Sbjct: 394 VGERGIRLSGGQKQRIAIARALLMDPKILLLDEATSALDAESEYLVQDAMDSIMKGRTVL 573
Query: 577 VVAHRLSTIRNVDTIAVIHQGKIVERGSHAELTNDPNGAYSQLIRLQ 623
V+AHRLST++ +T+AV+ G+IVE G+H EL + NG Y+ L++ Q
Sbjct: 574 VIAHRLSTVKTANTVAVVSDGQIVESGTHDELL-EKNGVYTALVKRQ 711
Score = 236 bits (602), Expect = 4e-62
Identities = 116/219 (52%), Positives = 162/219 (73%)
Frame = +1
Query: 1060 IFNDLCLNIRSGKTVALVGESGSGKSTVISLLQRFYDPDSGHITLDGIEIQRMQVKWLRQ 1119
+ + + ++ G VALVG SG GK+T+ +L++RFYDP G+I ++G+ + + K L +
Sbjct: 46 VLKGINIKLQPGSKVALVGPSGGGKTTIANLIERFYDPTKGNILVNGVPLVEISHKHLHR 225
Query: 1120 QMGLVSQEPILFNDTVRANIAYGKGGDATEAEIVAAAELANAHQFIGSLQKGYDTIVGER 1179
++ +VSQEP LFN ++ NIAYG G +A+I AA++ANAH+FI + + Y T VGER
Sbjct: 226 KISIVSQEPTLFNCSIEENIAYGFDGKIEDADIENAAKMANAHEFISNFPEKYKTFVGER 405
Query: 1180 GIQLSGGQKQRVAIARAIVKNPKILLLDEATSALDAESEKVVQDALDRVMVERTTIIVAH 1239
GI+LSGGQKQR+AIARA++ +PKILLLDEATSALDAESE +VQDA+D +M RT +++AH
Sbjct: 406 GIRLSGGQKQRIAIARALLMDPKILLLDEATSALDAESEYLVQDAMDSIMKGRTVLVIAH 585
Query: 1240 RLSTIKGADLIAVVKNGVIAEKGKHEALLHKGGDYASLV 1278
RLST+K A+ +AVV +G I E G H+ LL K G Y +LV
Sbjct: 586 RLSTVKTANTVAVVSDGQIVESGTHDELLEKNGVYTALV 702
Score = 23.9 bits (50), Expect(2) = 9e-65
Identities = 9/13 (69%), Positives = 11/13 (84%)
Frame = +2
Query: 387 LRDVCFSYPTRPD 399
L DV FSYP+RP+
Sbjct: 2 LNDVWFSYPSRPN 40
>TC86470 similar to PIR|F84919|F84919 glutathione-conjugate transporter AtMRP4
[imported] - Arabidopsis thaliana, partial (57%)
Length = 3175
Score = 240 bits (613), Expect = 2e-63
Identities = 227/926 (24%), Positives = 410/926 (43%), Gaps = 24/926 (2%)
Frame = +1
Query: 385 IELRDVCFSYPTRPDELIFNGFSLSLPSGTTAALVGQSGSGKSTVVSLIERFYDPTDGEV 444
++++D FS+ E +L + G A+VG GSGKS++++ I G+V
Sbjct: 154 VDVQDGTFSWDDEGLEQDLKNINLKVNKGELTAIVGTVGSGKSSLLASILGEMHRNSGKV 333
Query: 445 LIDGINLKEFQLKWIRQKIGLVSQEPVLFTCSIKENIAYGKDCAT---DEEIRVAAELAN 501
+ G Q WI+ +I+ENI +G +E IRV
Sbjct: 334 QVCGSTAYVAQTSWIQNG-------------TIEENILFGLPMNRQKYNEIIRVCC---- 462
Query: 502 AAKFIDKLPQGLDTMVGEHGTQLSGGQKQRVAIARAILKDPRILLLDEATSALDAES-ER 560
K ++ + G T +GE G LSGGQKQR+ +ARA+ +D I LLD+ SA+DA +
Sbjct: 463 LEKDLEMMEYGDQTEIGERGINLSGGQKQRIQLARAVYQDCDIYLLDDVFSAVDAHTGTE 642
Query: 561 IVQEALNRIMINRTTIVVAHRLSTIRNVDTIAVIHQGKIVERGSHAELTNDPNGAYSQLI 620
I +E + + +T ++V H++ + NVD I V+ G IV+ G + +L D + L+
Sbjct: 643 IFKECVRGALKGKTIVLVTHQVDFLHNVDRIVVMRDGMIVQSGRYNDLL-DSGLDFGVLV 819
Query: 621 RLQE--MKRSEQNDANDKNKPNSIVHSGRQSSQRSFSLRSISQGSAGNSGRHSFSASYVA 678
E M+ EQ A N ++ S +S S+ + NS
Sbjct: 820 AAHETSMELVEQGAAVPGENSNKLMIS------KSASINNRETNGESNS----------- 948
Query: 679 PTTDGFLETEDGGPQASPSKNSSPPEVPLYRLAYFNKPEIPVLLMGTITAVLHGAIMPVI 738
L+ + +S E + + I AVL +++
Sbjct: 949 ------LDQPNSAKGSSKLVKEEERETGKVSFNIYKRYCTEAFGWAGILAVLFLSVLWQA 1110
Query: 739 GLLVSKMISTFYKPADELR-HDSKVWAIVFVAVAVASLLIIPCRFYFFGVAGGKLIQRIR 797
++ S F + + V+ ++ A+ + S+++I R Y + G K Q
Sbjct: 1111 SMMASDYWLAFETSVERAEVFNPVVFISIYAAITIVSVILIVVRSYSVTIFGLKTAQIFF 1290
Query: 798 KLCFEKVVHMEVSWFDDVEHSSGALGARLSTDAASVRALVGDALGLLVQNIATIIVGMVI 857
++H +S++D SG + +R STD +V + + +V T+I ++I
Sbjct: 1291 NQILTSILHAPMSFYDTTP--SGRILSRASTDQTNVDIFIPLFINFVVAMYITVISIVII 1464
Query: 858 AFQASWQLAFIVLALAPLLGLNGYVQVKVLKGFSADAKKLYEEASQVA--------NDAV 909
Q SW AF+++ PL+ LN + +G+ + + ++++
Sbjct: 1465 TCQNSWPTAFLLI---PLVWLNIWY-----RGYFLSTSRELTRLDSITKAPVIVHFSESI 1620
Query: 910 GSIRTVSSFCAEEKV-MELYKQKCEGPIKKGVRRGIISGLGFGSSFFMLYAVDACVFYAG 968
+ TV +F +++ +E +K+ + +R + F L + + VF
Sbjct: 1621 SGVMTVRAFRKQKEFRLENFKR-----VNSNLRMDFHNYSSNAWLGFRLELLGSLVFCLS 1785
Query: 969 ARLVEDGKSTFSDVFLVFFALSMAAMGVSQSGTL---VPDSTNAKSAAASI-----FAIL 1020
A + S V +LS G+S + L + S ++ S+ F+ +
Sbjct: 1786 ALFMILLPSNIIKPENVGLSLSY---GLSLNSVLFWAIYMSCFIENKMVSVERIKQFSNI 1956
Query: 1021 DQKSQIDSSDESGMTLEEVKGDIEFNHVSFKYPTRLDVQIFNDLCLNIRSGKTVALVGES 1080
++ + D S +G ++ + +Y + + + L+I G+ V +VG +
Sbjct: 1957 PSEAAWNIKDRSPPPNWPGQGHVDIKDLQVRYRPNTPL-VLKGITLSISGGEKVGVVGRT 2133
Query: 1081 GSGKSTVISLLQRFYDPDSGHITLDGIEIQRMQVKWLRQQMGLVSQEPILFNDTVRANIA 1140
GSGKST+I + R +P G I +DGI+I + + LR + G++ QEP+LF TVR+NI
Sbjct: 2134 GSGKSTLIQVFFRLVEPTGGKIIIDGIDICALGLHDLRSRFGIIPQEPVLFEGTVRSNI- 2310
Query: 1141 YGKGGDATEAEIVAAAELANAHQFIGSLQKGYDTIVGERGIQLSGGQKQRVAIARAIVKN 1200
G T+ EI + + + S + D++V + G S GQ+Q + + R ++K
Sbjct: 2311 -DPTGQYTDDEIWKSLDRCQLKDTVASKPEKLDSLVVDNGDNWSVGQRQLLCLGRVMLKQ 2487
Query: 1201 PKILLLDEATSALDAESEKVVQDALDRVMVERTTIIVAHRLSTIKGADLIAVVKNGVIAE 1260
++L +DEAT+++D++++ V+Q + RT I +AHR+ T+ D + VV G E
Sbjct: 2488 SRLLFMDEATASVDSQTDAVIQKIIREDFAARTIISIAHRIPTVMDCDRVLVVDAGRAKE 2667
Query: 1261 KGKHEALLHKGGDYASLVALHTSDST 1286
K LL + +A+LV + + ST
Sbjct: 2668 FDKPSNLLQRQSLFAALVQEYANRST 2745
Score = 144 bits (364), Expect = 2e-34
Identities = 118/471 (25%), Positives = 224/471 (47%), Gaps = 13/471 (2%)
Frame = +1
Query: 144 LKTILRQDVSFFDKETNTGEVVGRMSGDTVLIKDAMGEKVGQFIQFMSTFIGGFVIAFTK 203
L +IL +SF+D T +G ++ R S D + + + + T I +I
Sbjct: 1300 LTSILHAPMSFYDT-TPSGRILSRASTDQTNVDIFIPLFINFVVAMYITVISIVIITCQN 1476
Query: 204 GWLLTVVMLSSIPLLILSGSMTSMVIAKASSTGQAAYSKSAGVV---EQTIGSIRTVASF 260
W ++ IPL+ L+ ++ + + A V+ ++I + TV +F
Sbjct: 1477 SWPTAFLL---IPLVWLNIWYRGYFLSTSRELTRLDSITKAPVIVHFSESISGVMTVRAF 1647
Query: 261 TGEKQ-ATANYNRSLIKVYKTAVQEALASGVGF-----GTLFFVFICSYGLAVWFGGKMI 314
+K+ N+ R + + + +GF G+L F C L + I
Sbjct: 1648 RKQKEFRLENFKRVNSNLRMDFHNYSSNAWLGFRLELLGSLVF---CLSALFMILLPSNI 1818
Query: 315 IEKGYTGGDVMTVIFAVLIGSTCLGQTSPSLSAFAAGQAAA---FKMFETINRKPEIDAY 371
I+ G +++ + + + S +S F + + K F I + +
Sbjct: 1819 IKPENVG---LSLSYGLSLNSVLFWAIY--MSCFIENKMVSVERIKQFSNIPSEAAWNIK 1983
Query: 372 DTSGKKLDDIRGDIELRDVCFSY-PTRPDELIFNGFSLSLPSGTTAALVGQSGSGKSTVV 430
D S +G ++++D+ Y P P L+ G +LS+ G +VG++GSGKST++
Sbjct: 1984 DRSPPPNWPGQGHVDIKDLQVRYRPNTP--LVLKGITLSISGGEKVGVVGRTGSGKSTLI 2157
Query: 431 SLIERFYDPTDGEVLIDGINLKEFQLKWIRQKIGLVSQEPVLFTCSIKENIAYGKDCATD 490
+ R +PT G+++IDGI++ L +R + G++ QEPVLF +++ NI TD
Sbjct: 2158 QVFFRLVEPTGGKIIIDGIDICALGLHDLRSRFGIIPQEPVLFEGTVRSNIDPTGQY-TD 2334
Query: 491 EEIRVAAELANAAKFIDKLPQGLDTMVGEHGTQLSGGQKQRVAIARAILKDPRILLLDEA 550
+EI + + + P+ LD++V ++G S GQ+Q + + R +LK R+L +DEA
Sbjct: 2335 DEIWKSLDRCQLKDTVASKPEKLDSLVVDNGDNWSVGQRQLLCLGRVMLKQSRLLFMDEA 2514
Query: 551 TSALDAESERIVQEALNRIMINRTTIVVAHRLSTIRNVDTIAVIHQGKIVE 601
T+++D++++ ++Q+ + RT I +AHR+ T+ + D + V+ G+ E
Sbjct: 2515 TASVDSQTDAVIQKIIREDFAARTIISIAHRIPTVMDCDRVLVVDAGRAKE 2667
>TC83850 weakly similar to GP|7300071|gb|AAF55241.1| CG4225 gene product
{Drosophila melanogaster}, partial (23%)
Length = 816
Score = 207 bits (526), Expect = 3e-53
Identities = 106/213 (49%), Positives = 147/213 (68%)
Frame = +2
Query: 396 TRPDELIFNGFSLSLPSGTTAALVGQSGSGKSTVVSLIERFYDPTDGEVLIDGINLKEFQ 455
T+ E +G S ++ GT A+VG+SGSGKST + L+ RFYD +G V +DG ++K +
Sbjct: 5 TKKGEPALDGVSFTVAPGTKTAIVGESGSGKSTCLKLLFRFYDVQEGAVTVDGHDIKHLK 184
Query: 456 LKWIRQKIGLVSQEPVLFTCSIKENIAYGKDCATDEEIRVAAELANAAKFIDKLPQGLDT 515
L +R+ IG+V Q+ VLF +I N+ Y A+ ++ A + AN + I P G +T
Sbjct: 185 LDSLRRNIGVVPQDTVLFNATIMYNLLYANPKASQADVFEACKAANIHERILNFPDGYET 364
Query: 516 MVGEHGTQLSGGQKQRVAIARAILKDPRILLLDEATSALDAESERIVQEALNRIMINRTT 575
VGE G +LSGG++QR+AIARAILKD RILLLDEAT++LD+ +ER +Q+AL R+ RTT
Sbjct: 365 KVGERGLKLSGGERQRIAIARAILKDARILLLDEATASLDSHTERQIQDALERVTAGRTT 544
Query: 576 IVVAHRLSTIRNVDTIAVIHQGKIVERGSHAEL 608
I +AHRLSTI D I V+H+ KIVERG+H+EL
Sbjct: 545 ITIAHRLSTITTSDQIVVLHKSKIVERGTHSEL 643
Score = 201 bits (511), Expect = 2e-51
Identities = 103/211 (48%), Positives = 149/211 (69%)
Frame = +2
Query: 1068 IRSGKTVALVGESGSGKSTVISLLQRFYDPDSGHITLDGIEIQRMQVKWLRQQMGLVSQE 1127
+ G A+VGESGSGKST + LL RFYD G +T+DG +I+ +++ LR+ +G+V Q+
Sbjct: 47 VAPGTKTAIVGESGSGKSTCLKLLFRFYDVQEGAVTVDGHDIKHLKLDSLRRNIGVVPQD 226
Query: 1128 PILFNDTVRANIAYGKGGDATEAEIVAAAELANAHQFIGSLQKGYDTIVGERGIQLSGGQ 1187
+LFN T+ N+ Y A++A++ A + AN H+ I + GY+T VGERG++LSGG+
Sbjct: 227 TVLFNATIMYNLLYANP-KASQADVFEACKAANIHERILNFPDGYETKVGERGLKLSGGE 403
Query: 1188 KQRVAIARAIVKNPKILLLDEATSALDAESEKVVQDALDRVMVERTTIIVAHRLSTIKGA 1247
+QR+AIARAI+K+ +ILLLDEAT++LD+ +E+ +QDAL+RV RTTI +AHRLSTI +
Sbjct: 404 RQRIAIARAILKDARILLLDEATASLDSHTERQIQDALERVTAGRTTITIAHRLSTITTS 583
Query: 1248 DLIAVVKNGVIAEKGKHEALLHKGGDYASLV 1278
D I V+ I E+G H LL GG Y ++V
Sbjct: 584 DQIVVLHKSKIVERGTHSELLALGGRYHAMV 676
>TC89340 weakly similar to GP|17427488|emb|CAD14006. PROBABLE COMPOSITE
ATP-BINDING TRANSMEMBRANE ABC TRANSPORTER PROTEIN
{Ralstonia solanacearum}, partial (32%)
Length = 795
Score = 185 bits (469), Expect = 1e-46
Identities = 103/233 (44%), Positives = 148/233 (63%)
Frame = +1
Query: 1036 LEEVKGDIEFNHVSFKYPTRLDVQIFNDLCLNIRSGKTVALVGESGSGKSTVISLLQRFY 1095
L + G+I + +V F Y + + DL + G T A VGESG GKSTV L+ R+Y
Sbjct: 85 LRDCNGNIRWKNVGFNYDPKN--KALKDLDFECKPGSTTAFVGESGGGKSTVFRLMFRYY 258
Query: 1096 DPDSGHITLDGIEIQRMQVKWLRQQMGLVSQEPILFNDTVRANIAYGKGGDATEAEIVAA 1155
+ SG I +DG +++ + + +R +G+V Q+ LFN+++ N+ Y + +AT+ EI A
Sbjct: 259 NTTSGSIEIDGRDVKDLTIDSVRSYIGVVPQDTTLFNESLMYNLRYARP-EATDEEIYDA 435
Query: 1156 AELANAHQFIGSLQKGYDTIVGERGIQLSGGQKQRVAIARAIVKNPKILLLDEATSALDA 1215
A+ H I S YDT VGERG++LSGG++QRVAIAR I+K+PKI++LDEATSALDA
Sbjct: 436 CRAASIHDRIMSFPDRYDTKVGERGLKLSGGERQRVAIARTIIKSPKIIMLDEATSALDA 615
Query: 1216 ESEKVVQDALDRVMVERTTIIVAHRLSTIKGADLIAVVKNGVIAEKGKHEALL 1268
+E+ +QD L + RT +I+AHRLSTI AD I V + KG H+ L+
Sbjct: 616 HTEQQIQDRLGHLGQGRTLLIIAHRLSTITHADQIIVPQQWDY**KGHHKELV 774
Score = 181 bits (460), Expect = 1e-45
Identities = 105/260 (40%), Positives = 150/260 (57%)
Frame = +1
Query: 357 KMFETINRKPEIDAYDTSGKKLDDIRGDIELRDVCFSYPTRPDELIFNGFSLSLPSGTTA 416
++ E KP + T+ + L D G+I ++V F+Y P G+T
Sbjct: 25 RLLELFKVKPTVVDSPTA-EPLRDCNGNIRWKNVGFNYD--PKNKALKDLDFECKPGSTT 195
Query: 417 ALVGQSGSGKSTVVSLIERFYDPTDGEVLIDGINLKEFQLKWIRQKIGLVSQEPVLFTCS 476
A VG+SG GKSTV L+ R+Y+ T G + IDG ++K+ + +R IG+V Q+ LF S
Sbjct: 196 AFVGESGGGKSTVFRLMFRYYNTTSGSIEIDGRDVKDLTIDSVRSYIGVVPQDTTLFNES 375
Query: 477 IKENIAYGKDCATDEEIRVAAELANAAKFIDKLPQGLDTMVGEHGTQLSGGQKQRVAIAR 536
+ N+ Y + ATDEEI A A+ I P DT VGE G +LSGG++QRVAIAR
Sbjct: 376 LMYNLRYARPEATDEEIYDACRAASIHDRIMSFPDRYDTKVGERGLKLSGGERQRVAIAR 555
Query: 537 AILKDPRILLLDEATSALDAESERIVQEALNRIMINRTTIVVAHRLSTIRNVDTIAVIHQ 596
I+K P+I++LDEATSALDA +E+ +Q+ L + RT +++AHRLSTI + D I V Q
Sbjct: 556 TIIKSPKIIMLDEATSALDAHTEQQIQDRLGHLGQGRTLLIIAHRLSTITHADQIIVPQQ 735
Query: 597 GKIVERGSHAELTNDPNGAY 616
+G H EL +G +
Sbjct: 736 WDY**KGHHKELVMPMDGTH 795
>BM813979 homologue to PIR|E85023|E85 probable P-glycoprotein-like protein
[imported] - Arabidopsis thaliana, partial (7%)
Length = 434
Score = 171 bits (432), Expect = 2e-42
Identities = 83/99 (83%), Positives = 94/99 (94%)
Frame = +3
Query: 1072 KTVALVGESGSGKSTVISLLQRFYDPDSGHITLDGIEIQRMQVKWLRQQMGLVSQEPILF 1131
+TVALVGESGSGKSTVI+LLQRFYDPDSG ITLDGIEI+++Q+KWLRQQMGLVSQEP+LF
Sbjct: 135 QTVALVGESGSGKSTVIALLQRFYDPDSGEITLDGIEIRQLQLKWLRQQMGLVSQEPVLF 314
Query: 1132 NDTVRANIAYGKGGDATEAEIVAAAELANAHQFIGSLQK 1170
NDT+RANIAYGKGG ATEAEI+AAAELANAH+FI L +
Sbjct: 315 NDTIRANIAYGKGGIATEAEIIAAAELANAHRFISGLHR 431
Score = 124 bits (310), Expect = 3e-28
Identities = 61/96 (63%), Positives = 78/96 (80%), Gaps = 1/96 (1%)
Frame = +3
Query: 415 TAALVGQSGSGKSTVVSLIERFYDPTDGEVLIDGINLKEFQLKWIRQKIGLVSQEPVLFT 474
T ALVG+SGSGKSTV++L++RFYDP GE+ +DGI +++ QLKW+RQ++GLVSQEPVLF
Sbjct: 138 TVALVGESGSGKSTVIALLQRFYDPDSGEITLDGIEIRQLQLKWLRQQMGLVSQEPVLFN 317
Query: 475 CSIKENIAYGK-DCATDEEIRVAAELANAAKFIDKL 509
+I+ NIAYGK AT+ EI AAELANA +FI L
Sbjct: 318 DTIRANIAYGKGGIATEAEIIAAAELANAHRFISGL 425
>AL382570 weakly similar to GP|10241621|em putative P-glycoprotein {Oryza
sativa}, partial (17%)
Length = 481
Score = 169 bits (429), Expect = 5e-42
Identities = 85/150 (56%), Positives = 111/150 (73%)
Frame = +3
Query: 426 KSTVVSLIERFYDPTDGEVLIDGINLKEFQLKWIRQKIGLVSQEPVLFTCSIKENIAYGK 485
KS+V++L+ +FYDP G+VLIDG +L+E+ L+W+R +IGLV QEP+LF CSI+ENI YG
Sbjct: 3 KSSVLALLLKFYDPVVGKVLIDGKDLREYNLRWLRTQIGLVQQEPLLFNCSIRENICYGN 182
Query: 486 DCATDEEIRVAAELANAAKFIDKLPQGLDTMVGEHGTQLSGGQKQRVAIARAILKDPRIL 545
+ A + EI A AN +F+ LP G +T+VGE G QLSGGQKQR+AIAR +LK P IL
Sbjct: 183 NGAFESEIVEVAREANIHEFVSNLPNGYNTVVGEKGCQLSGGQKQRIAIARTLLKKPAIL 362
Query: 546 LLDEATSALDAESERIVQEALNRIMINRTT 575
LLDEATSALDA SER + A+ + + T
Sbjct: 363 LLDEATSALDA*SERTIVNAIKAMNLKEET 452
Score = 166 bits (419), Expect = 7e-41
Identities = 81/151 (53%), Positives = 113/151 (74%)
Frame = +3
Query: 1084 KSTVISLLQRFYDPDSGHITLDGIEIQRMQVKWLRQQMGLVSQEPILFNDTVRANIAYGK 1143
KS+V++LL +FYDP G + +DG +++ ++WLR Q+GLV QEP+LFN ++R NI YG
Sbjct: 3 KSSVLALLLKFYDPVVGKVLIDGKDLREYNLRWLRTQIGLVQQEPLLFNCSIRENICYGN 182
Query: 1144 GGDATEAEIVAAAELANAHQFIGSLQKGYDTIVGERGIQLSGGQKQRVAIARAIVKNPKI 1203
G A E+EIV A AN H+F+ +L GY+T+VGE+G QLSGGQKQR+AIAR ++K P I
Sbjct: 183 NG-AFESEIVEVAREANIHEFVSNLPNGYNTVVGEKGCQLSGGQKQRIAIARTLLKKPAI 359
Query: 1204 LLLDEATSALDAESEKVVQDALDRVMVERTT 1234
LLLDEATSALDA SE+ + +A+ + ++ T
Sbjct: 360 LLLDEATSALDA*SERTIVNAIKAMNLKEET 452
>AL383696 homologue to PIR|T47671|T4 P-glycoprotein-like - Arabidopsis
thaliana, partial (9%)
Length = 391
Score = 145 bits (365), Expect = 1e-34
Identities = 68/130 (52%), Positives = 94/130 (72%)
Frame = +1
Query: 401 LIFNGFSLSLPSGTTAALVGQSGSGKSTVVSLIERFYDPTDGEVLIDGINLKEFQLKWIR 460
L+ + FSL + G T A+VG SGSGKST++SLIERFYDP G+VL+DG +LK + L+W+R
Sbjct: 1 LVLSNFSLKVNGGQTVAVVGVSGSGKSTIISLIERFYDPVAGQVLLDGRDLKLYNLRWLR 180
Query: 461 QKIGLVSQEPVLFTCSIKENIAYGKDCATDEEIRVAAELANAAKFIDKLPQGLDTMVGEH 520
+GL+ QEP++F+ +I+ENI Y + A++ E++ AA +ANA FI LP G DT VG
Sbjct: 181 SHLGLIQQEPIIFSTTIRENIIYARHNASEAEMKEAARIANAHHFISSLPHGYDTHVGMR 360
Query: 521 GTQLSGGQKQ 530
G L+ GQKQ
Sbjct: 361 GVDLTPGQKQ 390
Score = 143 bits (361), Expect = 4e-34
Identities = 68/130 (52%), Positives = 96/130 (73%)
Frame = +1
Query: 1060 IFNDLCLNIRSGKTVALVGESGSGKSTVISLLQRFYDPDSGHITLDGIEIQRMQVKWLRQ 1119
+ ++ L + G+TVA+VG SGSGKST+ISL++RFYDP +G + LDG +++ ++WLR
Sbjct: 4 VLSNFSLKVNGGQTVAVVGVSGSGKSTIISLIERFYDPVAGQVLLDGRDLKLYNLRWLRS 183
Query: 1120 QMGLVSQEPILFNDTVRANIAYGKGGDATEAEIVAAAELANAHQFIGSLQKGYDTIVGER 1179
+GL+ QEPI+F+ T+R NI Y + +A+EAE+ AA +ANAH FI SL GYDT VG R
Sbjct: 184 HLGLIQQEPIIFSTTIRENIIYAR-HNASEAEMKEAARIANAHHFISSLPHGYDTHVGMR 360
Query: 1180 GIQLSGGQKQ 1189
G+ L+ GQKQ
Sbjct: 361 GVDLTPGQKQ 390
>TC92097 similar to GP|22553016|emb|CAD44995. multidrug-resistance related
protein {Arabidopsis thaliana}, partial (25%)
Length = 1011
Score = 144 bits (363), Expect = 2e-34
Identities = 83/221 (37%), Positives = 132/221 (59%), Gaps = 1/221 (0%)
Frame = +2
Query: 382 RGDIELRDVCFSYPTRPDE-LIFNGFSLSLPSGTTAALVGQSGSGKSTVVSLIERFYDPT 440
+G I+L+ + Y RP+ L+ G + G+ +VG++GSGKST++S + R +P+
Sbjct: 347 KGKIDLQGLEIRY--RPNSPLVLKGIICTFKEGSRVGVVGRTGSGKSTLISALFRLVEPS 520
Query: 441 DGEVLIDGINLKEFQLKWIRQKIGLVSQEPVLFTCSIKENIAYGKDCATDEEIRVAAELA 500
G++LIDGIN+ LK +R K+ ++ QEP LF SI+ N+ +D+EI A E
Sbjct: 521 RGDILIDGINICSIGLKDLRTKLSIIPQEPTLFKGSIRTNLD-PLGLYSDDEIWKAVEKC 697
Query: 501 NAAKFIDKLPQGLDTMVGEHGTQLSGGQKQRVAIARAILKDPRILLLDEATSALDAESER 560
+ I KLP LD+ V + G S GQ+Q + R +LK RIL+LDEAT+++D+ ++
Sbjct: 698 QLKETISKLPSLLDSSVSDEGGNWSLGQRQLFCLGRVLLKRNRILVLDEATASIDSATDA 877
Query: 561 IVQEALNRIMINRTTIVVAHRLSTIRNVDTIAVIHQGKIVE 601
I+Q + + T I VAHR+ T+ + D + V+ GK+VE
Sbjct: 878 ILQRIIRQEFEECTVITVAHRVPTVIDSDMVMVLSYGKLVE 1000
Score = 125 bits (313), Expect = 1e-28
Identities = 73/221 (33%), Positives = 122/221 (55%)
Frame = +2
Query: 1040 KGDIEFNHVSFKYPTRLDVQIFNDLCLNIRSGKTVALVGESGSGKSTVISLLQRFYDPDS 1099
KG I+ + +Y + + +C + G V +VG +GSGKST+IS L R +P
Sbjct: 347 KGKIDLQGLEIRYRPNSPLVLKGIIC-TFKEGSRVGVVGRTGSGKSTLISALFRLVEPSR 523
Query: 1100 GHITLDGIEIQRMQVKWLRQQMGLVSQEPILFNDTVRANIAYGKGGDATEAEIVAAAELA 1159
G I +DGI I + +K LR ++ ++ QEP LF ++R N+ G ++ EI A E
Sbjct: 524 GDILIDGINICSIGLKDLRTKLSIIPQEPTLFKGSIRTNL--DPLGLYSDDEIWKAVEKC 697
Query: 1160 NAHQFIGSLQKGYDTIVGERGIQLSGGQKQRVAIARAIVKNPKILLLDEATSALDAESEK 1219
+ I L D+ V + G S GQ+Q + R ++K +IL+LDEAT+++D+ ++
Sbjct: 698 QLKETISKLPSLLDSSVSDEGGNWSLGQRQLFCLGRVLLKRNRILVLDEATASIDSATDA 877
Query: 1220 VVQDALDRVMVERTTIIVAHRLSTIKGADLIAVVKNGVIAE 1260
++Q + + E T I VAHR+ T+ +D++ V+ G + E
Sbjct: 878 ILQRIIRQEFEECTVITVAHRVPTVIDSDMVMVLSYGKLVE 1000
>TC92328 similar to PIR|T52080|T52080 multi resistance protein [imported] -
Arabidopsis thaliana, partial (18%)
Length = 1133
Score = 138 bits (348), Expect = 1e-32
Identities = 76/238 (31%), Positives = 138/238 (57%)
Frame = +1
Query: 383 GDIELRDVCFSYPTRPDELIFNGFSLSLPSGTTAALVGQSGSGKSTVVSLIERFYDPTDG 442
G IE+ D+ Y L+ +G S + P G +VG++GSGKST++ + R +P DG
Sbjct: 100 GTIEIFDLKVRYKENLP-LVLHGVSCTFPGGKNIGIVGRTGSGKSTLIQALFRLIEPADG 276
Query: 443 EVLIDGINLKEFQLKWIRQKIGLVSQEPVLFTCSIKENIAYGKDCATDEEIRVAAELANA 502
+ ID IN+ E L +R + ++ Q+P LF +I+ N+ ++ +D++I A + +
Sbjct: 277 SIHIDNINIFEIGLHDLRSHLSIIPQDPTLFEGTIRGNLDPLEE-HSDKDIWEALDKSQL 453
Query: 503 AKFIDKLPQGLDTMVGEHGTQLSGGQKQRVAIARAILKDPRILLLDEATSALDAESERIV 562
+ I + Q LDT V E+G S GQ+Q V++ RA+LK +IL+LDEAT+++D ++ ++
Sbjct: 454 GEIIREKGQKLDTPVIENGDNWSVGQRQLVSLGRALLKQSKILVLDEATASVDTATDNLI 633
Query: 563 QEALNRIMINRTTIVVAHRLSTIRNVDTIAVIHQGKIVERGSHAELTNDPNGAYSQLI 620
Q+ + + T + +AHR+ T+ + D + V+ G++ E + L D + + +L+
Sbjct: 634 QKIIRTEFKDCTVLTIAHRIPTVIDSDQVLVLSDGRVAEFDTPLRLLEDRSSMFLKLV 807
Score = 120 bits (302), Expect = 3e-27
Identities = 76/247 (30%), Positives = 132/247 (52%), Gaps = 1/247 (0%)
Frame = +1
Query: 1041 GDIEFNHVSFKYPTRLDVQIFNDLCLNIRSGKTVALVGESGSGKSTVISLLQRFYDPDSG 1100
G IE + +Y L + + C GK + +VG +GSGKST+I L R +P G
Sbjct: 100 GTIEIFDLKVRYKENLPLVLHGVSC-TFPGGKNIGIVGRTGSGKSTLIQALFRLIEPADG 276
Query: 1101 HITLDGIEIQRMQVKWLRQQMGLVSQEPILFNDTVRANIAYGKGGDATEAEIVAAAELAN 1160
I +D I I + + LR + ++ Q+P LF T+R N+ + + ++ +I A + +
Sbjct: 277 SIHIDNINIFEIGLHDLRSHLSIIPQDPTLFEGTIRGNLDPLE--EHSDKDIWEALDKSQ 450
Query: 1161 AHQFIGSLQKGYDTIVGERGIQLSGGQKQRVAIARAIVKNPKILLLDEATSALDAESEKV 1220
+ I + DT V E G S GQ+Q V++ RA++K KIL+LDEAT+++D ++ +
Sbjct: 451 LGEIIREKGQKLDTPVIENGDNWSVGQRQLVSLGRALLKQSKILVLDEATASVDTATDNL 630
Query: 1221 VQDALDRVMVERTTIIVAHRLSTIKGADLIAVVKNGVIAEKGKHEALLH-KGGDYASLVA 1279
+Q + + T + +AHR+ T+ +D + V+ +G +AE LL + + LV
Sbjct: 631 IQKIIRTEFKDCTVLTIAHRIPTVIDSDQVLVLSDGRVAEFDTPLRLLEDRSSMFLKLVT 810
Query: 1280 LHTSDST 1286
++S S+
Sbjct: 811 EYSSRSS 831
>TC87490 similar to PIR|T52080|T52080 multi resistance protein [imported] -
Arabidopsis thaliana, partial (19%)
Length = 927
Score = 134 bits (338), Expect = 2e-31
Identities = 74/240 (30%), Positives = 138/240 (56%)
Frame = +1
Query: 381 IRGDIELRDVCFSYPTRPDELIFNGFSLSLPSGTTAALVGQSGSGKSTVVSLIERFYDPT 440
+ G I+L D+ Y ++ +G S + P G +VG++GSGKST++ + R +P
Sbjct: 154 VNGTIQLIDLKVRYKENLP-MVLHGVSCTFPGGKKIGIVGRTGSGKSTLIQALFRLVEPA 330
Query: 441 DGEVLIDGINLKEFQLKWIRQKIGLVSQEPVLFTCSIKENIAYGKDCATDEEIRVAAELA 500
G +LID I++ L +R + ++ Q+P LF +I+ N+ ++ +D+EI A + +
Sbjct: 331 AGSILIDNIDISGIGLHDLRSHLSIIPQDPTLFEGTIRGNLDPLEE-HSDKEIWEALDKS 507
Query: 501 NAAKFIDKLPQGLDTMVGEHGTQLSGGQKQRVAIARAILKDPRILLLDEATSALDAESER 560
+ I + Q LDT V ++G S GQ+Q VA+ RA+LK +IL+LDEAT+++D+ ++
Sbjct: 508 QLGEIIREKGQKLDTPVLQNGDNWSVGQRQLVALGRALLKQSKILVLDEATASVDSATDN 687
Query: 561 IVQEALNRIMINRTTIVVAHRLSTIRNVDTIAVIHQGKIVERGSHAELTNDPNGAYSQLI 620
++Q+ + + T +AHR+ T+ + D + V++ G + E + L D + + +L+
Sbjct: 688 LIQKVIREEFRDCTVCTIAHRIPTVIDSDLVLVLNDGLLAEFDTPLRLLEDKSSMFLKLV 867
Score = 129 bits (325), Expect = 6e-30
Identities = 80/249 (32%), Positives = 137/249 (54%), Gaps = 1/249 (0%)
Frame = +1
Query: 1039 VKGDIEFNHVSFKYPTRLDVQIFNDLCLNIRSGKTVALVGESGSGKSTVISLLQRFYDPD 1098
V G I+ + +Y L + + C GK + +VG +GSGKST+I L R +P
Sbjct: 154 VNGTIQLIDLKVRYKENLPMVLHGVSC-TFPGGKKIGIVGRTGSGKSTLIQALFRLVEPA 330
Query: 1099 SGHITLDGIEIQRMQVKWLRQQMGLVSQEPILFNDTVRANIAYGKGGDATEAEIVAAAEL 1158
+G I +D I+I + + LR + ++ Q+P LF T+R N+ + + ++ EI A +
Sbjct: 331 AGSILIDNIDISGIGLHDLRSHLSIIPQDPTLFEGTIRGNLDPLE--EHSDKEIWEALDK 504
Query: 1159 ANAHQFIGSLQKGYDTIVGERGIQLSGGQKQRVAIARAIVKNPKILLLDEATSALDAESE 1218
+ + I + DT V + G S GQ+Q VA+ RA++K KIL+LDEAT+++D+ ++
Sbjct: 505 SQLGEIIREKGQKLDTPVLQNGDNWSVGQRQLVALGRALLKQSKILVLDEATASVDSATD 684
Query: 1219 KVVQDALDRVMVERTTIIVAHRLSTIKGADLIAVVKNGVIAEKGKHEALLH-KGGDYASL 1277
++Q + + T +AHR+ T+ +DL+ V+ +G++AE LL K + L
Sbjct: 685 NLIQKVIREEFRDCTVCTIAHRIPTVIDSDLVLVLNDGLLAEFDTPLRLLEDKSSMFLKL 864
Query: 1278 VALHTSDST 1286
V ++S ST
Sbjct: 865 VTEYSSRST 891
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.319 0.135 0.377
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 30,986,719
Number of Sequences: 36976
Number of extensions: 372569
Number of successful extensions: 2221
Number of sequences better than 10.0: 137
Number of HSP's better than 10.0 without gapping: 1952
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2139
length of query: 1287
length of database: 9,014,727
effective HSP length: 107
effective length of query: 1180
effective length of database: 5,058,295
effective search space: 5968788100
effective search space used: 5968788100
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 64 (29.3 bits)
Medicago: description of AC146862.19


