

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146862.18 - phase: 0 /pseudo
(155 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
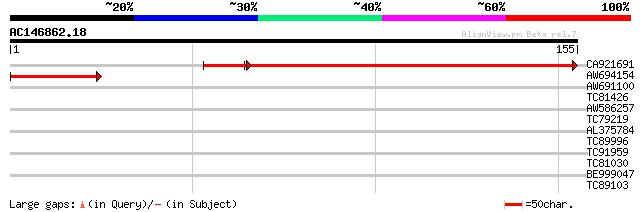
CA921691 similar to GP|17064896|gb Unknown protein {Arabidopsis ... 179 3e-48
AW694154 similar to GP|10177185|db contains similarity to 5'-nuc... 55 9e-09
AW691100 homologue to GP|20260154|gb| cytosolic IMP-GMP specific... 30 0.39
TC81426 similar to GP|6016719|gb|AAF01545.1| putative cell divis... 29 0.87
AW586257 weakly similar to GP|14532668|gb putative RB-binding pr... 28 1.5
TC79219 similar to GP|15810000|gb|AAL06927.1 AT5g23340/MKD15_20 ... 27 2.5
AL375784 homologue to SP|P31167|ADT1_ ADP ATP carrier protein 1 ... 27 3.3
TC89996 similar to GP|13430502|gb|AAK25873.1 unknown protein {Ar... 26 7.4
TC91959 GP|22653454|gb|AAN04071.1 Orf284 {Amoebidium parasiticum... 25 9.6
TC81030 similar to GP|6006872|gb|AAF00648.1| s-syntaxin-like pro... 25 9.6
BE999047 similar to GP|21740855|emb OSJNBb0072N21.4 {Oryza sativ... 25 9.6
TC89103 similar to GP|16930441|gb|AAL31906.1 At2g42810/F7D19.19 ... 25 9.6
>CA921691 similar to GP|17064896|gb Unknown protein {Arabidopsis thaliana},
partial (20%)
Length = 763
Score = 179 bits (453), Expect(2) = 3e-48
Identities = 89/91 (97%), Positives = 91/91 (99%)
Frame = -3
Query: 65 QKEVVGDLFNQLRLALQRRSKDRPAQTLAATNMDDEDLTESMQKLLIVMQRLDEKIAPML 124
++EVVGDLFNQLRLALQRRSKDRPAQTLAATNMDDEDLTESMQKLLIVMQRLDEKIAPML
Sbjct: 722 KREVVGDLFNQLRLALQRRSKDRPAQTLAATNMDDEDLTESMQKLLIVMQRLDEKIAPML 543
Query: 125 EADGELFNSRWGFLSRAGLWDKSHLMRQIEK 155
EADGELFNSRWGFLSRAGLWDKSHLMRQIEK
Sbjct: 542 EADGELFNSRWGFLSRAGLWDKSHLMRQIEK 450
Score = 28.9 bits (63), Expect(2) = 3e-48
Identities = 13/13 (100%), Positives = 13/13 (100%)
Frame = -2
Query: 54 GDRESLVELINQK 66
GDRESLVELINQK
Sbjct: 756 GDRESLVELINQK 718
>AW694154 similar to GP|10177185|db contains similarity to
5'-nucleotidase~gene_id:K19E20.8 {Arabidopsis thaliana},
partial (29%)
Length = 581
Score = 55.5 bits (132), Expect = 9e-09
Identities = 25/25 (100%), Positives = 25/25 (100%)
Frame = +2
Query: 1 MVENSLGIHGDEILYVGDHIYTDVS 25
MVENSLGIHGDEILYVGDHIYTDVS
Sbjct: 488 MVENSLGIHGDEILYVGDHIYTDVS 562
>AW691100 homologue to GP|20260154|gb| cytosolic IMP-GMP specific
5-nucleotidase putative {Arabidopsis thaliana},
partial (5%)
Length = 539
Score = 30.0 bits (66), Expect = 0.39
Identities = 11/17 (64%), Positives = 15/17 (87%)
Frame = +1
Query: 12 EILYVGDHIYTDVSQSK 28
++LYVGDHIY D+ +SK
Sbjct: 37 QVLYVGDHIYGDILRSK 87
>TC81426 similar to GP|6016719|gb|AAF01545.1| putative cell division control
protein {Arabidopsis thaliana}, partial (21%)
Length = 965
Score = 28.9 bits (63), Expect = 0.87
Identities = 29/98 (29%), Positives = 41/98 (41%), Gaps = 2/98 (2%)
Frame = +2
Query: 4 NSLGIHGDEIL--YVGDHIYTDVSQSKVHLRWRTALICRELEDEYSALIRCRGDRESLVE 61
N + I G E+L YVG+ ++++ + R RT C DE AL RG V
Sbjct: 68 NFIHIKGPELLNKYVGE---SELAVRTLFNRARTCAPCVLFFDEVDALTTKRGKEGGWV- 235
Query: 62 LINQKEVVGDLFNQLRLALQRRSKDRPAQTLAATNMDD 99
+ L NQL + L + R + ATN D
Sbjct: 236 -------IERLLNQLLIELDGAEQRRGVFVIGATNRPD 328
>AW586257 weakly similar to GP|14532668|gb putative RB-binding protein
{Arabidopsis thaliana}, partial (3%)
Length = 590
Score = 28.1 bits (61), Expect = 1.5
Identities = 17/77 (22%), Positives = 41/77 (53%)
Frame = +1
Query: 58 SLVELINQKEVVGDLFNQLRLALQRRSKDRPAQTLAATNMDDEDLTESMQKLLIVMQRLD 117
+++E+ +K+ + L L +A++ + + A+ D ED+ + + + +++ LD
Sbjct: 238 AMLEIEGEKQFIS-LSCVLGVAMRWEERAGAILSAEASISDFEDMIRASENIFVILASLD 414
Query: 118 EKIAPMLEADGELFNSR 134
+ +LEA+ L NS+
Sbjct: 415 DVNKALLEANSWLRNSK 465
>TC79219 similar to GP|15810000|gb|AAL06927.1 AT5g23340/MKD15_20 {Arabidopsis
thaliana}, partial (78%)
Length = 1619
Score = 27.3 bits (59), Expect = 2.5
Identities = 16/52 (30%), Positives = 26/52 (49%)
Frame = -3
Query: 65 QKEVVGDLFNQLRLALQRRSKDRPAQTLAATNMDDEDLTESMQKLLIVMQRL 116
Q+ G L L+L LQ+ K +PA + A + DLT + L + +R+
Sbjct: 1335 QRRTAGSLMLPLKLTLQKSGKCKPASSRPAFVICGHDLTSKYFR*LHLPKRM 1180
>AL375784 homologue to SP|P31167|ADT1_ ADP ATP carrier protein 1
mitochondrial precursor (ADP/ATP translocase 1), partial
(10%)
Length = 502
Score = 26.9 bits (58), Expect = 3.3
Identities = 14/27 (51%), Positives = 18/27 (65%), Gaps = 1/27 (3%)
Frame = +3
Query: 13 ILYVGDHIYTDVSQSKVHLR-WRTALI 38
I Y+G + TDVSQS +H R W+ LI
Sbjct: 390 ISYMGQSVETDVSQSYIHQRIWQF*LI 470
>TC89996 similar to GP|13430502|gb|AAK25873.1 unknown protein {Arabidopsis
thaliana}, partial (5%)
Length = 628
Score = 25.8 bits (55), Expect = 7.4
Identities = 15/38 (39%), Positives = 22/38 (57%), Gaps = 2/38 (5%)
Frame = +3
Query: 92 LAATNMDDE--DLTESMQKLLIVMQRLDEKIAPMLEAD 127
L A ++DE T ++QKL + Q+L K P +EAD
Sbjct: 39 LEAAQLEDEVEAATSTLQKLAEMSQQLLNKAMPSVEAD 152
>TC91959 GP|22653454|gb|AAN04071.1 Orf284 {Amoebidium parasiticum}, partial
(4%)
Length = 900
Score = 25.4 bits (54), Expect = 9.6
Identities = 14/36 (38%), Positives = 19/36 (51%)
Frame = -2
Query: 31 LRWRTALICRELEDEYSALIRCRGDRESLVELINQK 66
L W +C D+YS RCR R+S+ L+ QK
Sbjct: 584 LEWN*ESVC----DQYSHNNRCRRHRKSIKMLVLQK 489
>TC81030 similar to GP|6006872|gb|AAF00648.1| s-syntaxin-like protein
{Arabidopsis thaliana}, partial (93%)
Length = 1212
Score = 25.4 bits (54), Expect = 9.6
Identities = 18/73 (24%), Positives = 37/73 (50%)
Frame = +1
Query: 61 ELINQKEVVGDLFNQLRLALQRRSKDRPAQTLAATNMDDEDLTESMQKLLIVMQRLDEKI 120
E I++ GD + A+Q + + + TLA E + + +KLL + Q + I
Sbjct: 634 ETIDRLIETGDSEQIFQKAIQEQGRGQVMDTLAEIQERHEAVRDVERKLLDLQQTFMD-I 810
Query: 121 APMLEADGELFNS 133
A +++A G++ ++
Sbjct: 811 AVLVDAQGDMLDN 849
>BE999047 similar to GP|21740855|emb OSJNBb0072N21.4 {Oryza sativa}, partial
(3%)
Length = 540
Score = 25.4 bits (54), Expect = 9.6
Identities = 15/39 (38%), Positives = 23/39 (58%)
Frame = -1
Query: 101 DLTESMQKLLIVMQRLDEKIAPMLEADGELFNSRWGFLS 139
D++ +K LI R+ + P++E +GELF RW LS
Sbjct: 363 DISFH*KKYLISSHRI--LLLPIVEFEGELFACRWEPLS 253
>TC89103 similar to GP|16930441|gb|AAL31906.1 At2g42810/F7D19.19
{Arabidopsis thaliana}, partial (66%)
Length = 1107
Score = 25.4 bits (54), Expect = 9.6
Identities = 9/23 (39%), Positives = 16/23 (69%)
Frame = -1
Query: 24 VSQSKVHLRWRTALICRELEDEY 46
+S S +H RWR AL C++ + ++
Sbjct: 420 ISSSYIHGRWRHALKCKQSKCDF 352
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.320 0.136 0.393
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,091,351
Number of Sequences: 36976
Number of extensions: 43228
Number of successful extensions: 172
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 172
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 172
length of query: 155
length of database: 9,014,727
effective HSP length: 88
effective length of query: 67
effective length of database: 5,760,839
effective search space: 385976213
effective search space used: 385976213
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 54 (25.4 bits)
Medicago: description of AC146862.18


