

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146855.3 - phase: 0
(186 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
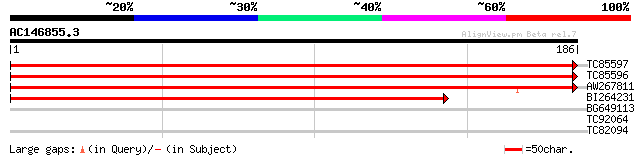
TC85597 similar to GP|22795244|gb|AAN08216.1 ribosomal protein L... 383 e-107
TC85596 similar to GP|22795244|gb|AAN08216.1 ribosomal protein L... 383 e-107
AW267811 similar to SP|O65082|R15B 60S ribosomal protein L15-2. ... 260 2e-70
BI264231 similar to SP|O65050|R15A 60S ribosomal protein L15-1. ... 237 2e-63
BG649113 similar to GP|18033111|gb functional candidate resistan... 27 3.6
TC92064 similar to PIR|T08935|T08935 COP1-interacting protein 7 ... 27 6.1
TC82094 similar to GP|20259195|gb|AAM14313.1 unknown protein {Ar... 26 8.0
>TC85597 similar to GP|22795244|gb|AAN08216.1 ribosomal protein L15 {Oryza
sativa (japonica cultivar-group)}, complete
Length = 904
Score = 383 bits (983), Expect = e-107
Identities = 186/186 (100%), Positives = 186/186 (100%)
Frame = +2
Query: 1 MRFMQRVRCWEYRQQSSIVRLTRPTRPDKARRLGYKAKQGYVVYRVRVRRGGRKRPVSKG 60
MRFMQRVRCWEYRQQSSIVRLTRPTRPDKARRLGYKAKQGYVVYRVRVRRGGRKRPVSKG
Sbjct: 122 MRFMQRVRCWEYRQQSSIVRLTRPTRPDKARRLGYKAKQGYVVYRVRVRRGGRKRPVSKG 301
Query: 61 IVYGKPTNQGVTQLKFQRSKRSVAEERAGRKLGGLRVLNSYWVNEDSTFKYFEVILVDVA 120
IVYGKPTNQGVTQLKFQRSKRSVAEERAGRKLGGLRVLNSYWVNEDSTFKYFEVILVDVA
Sbjct: 302 IVYGKPTNQGVTQLKFQRSKRSVAEERAGRKLGGLRVLNSYWVNEDSTFKYFEVILVDVA 481
Query: 121 HSAIRNDPRINWLTNPVHKHRELRGLTSAGKENRGLSGKGHRYHKARPSRRANWKRNNTL 180
HSAIRNDPRINWLTNPVHKHRELRGLTSAGKENRGLSGKGHRYHKARPSRRANWKRNNTL
Sbjct: 482 HSAIRNDPRINWLTNPVHKHRELRGLTSAGKENRGLSGKGHRYHKARPSRRANWKRNNTL 661
Query: 181 SLRRYR 186
SLRRYR
Sbjct: 662 SLRRYR 679
>TC85596 similar to GP|22795244|gb|AAN08216.1 ribosomal protein L15 {Oryza
sativa (japonica cultivar-group)}, complete
Length = 879
Score = 383 bits (983), Expect = e-107
Identities = 186/186 (100%), Positives = 186/186 (100%)
Frame = +1
Query: 1 MRFMQRVRCWEYRQQSSIVRLTRPTRPDKARRLGYKAKQGYVVYRVRVRRGGRKRPVSKG 60
MRFMQRVRCWEYRQQSSIVRLTRPTRPDKARRLGYKAKQGYVVYRVRVRRGGRKRPVSKG
Sbjct: 121 MRFMQRVRCWEYRQQSSIVRLTRPTRPDKARRLGYKAKQGYVVYRVRVRRGGRKRPVSKG 300
Query: 61 IVYGKPTNQGVTQLKFQRSKRSVAEERAGRKLGGLRVLNSYWVNEDSTFKYFEVILVDVA 120
IVYGKPTNQGVTQLKFQRSKRSVAEERAGRKLGGLRVLNSYWVNEDSTFKYFEVILVDVA
Sbjct: 301 IVYGKPTNQGVTQLKFQRSKRSVAEERAGRKLGGLRVLNSYWVNEDSTFKYFEVILVDVA 480
Query: 121 HSAIRNDPRINWLTNPVHKHRELRGLTSAGKENRGLSGKGHRYHKARPSRRANWKRNNTL 180
HSAIRNDPRINWLTNPVHKHRELRGLTSAGKENRGLSGKGHRYHKARPSRRANWKRNNTL
Sbjct: 481 HSAIRNDPRINWLTNPVHKHRELRGLTSAGKENRGLSGKGHRYHKARPSRRANWKRNNTL 660
Query: 181 SLRRYR 186
SLRRYR
Sbjct: 661 SLRRYR 678
>AW267811 similar to SP|O65082|R15B 60S ribosomal protein L15-2. [Black
spruce] {Picea mariana}, partial (96%)
Length = 725
Score = 260 bits (665), Expect = 2e-70
Identities = 127/187 (67%), Positives = 149/187 (78%), Gaps = 1/187 (0%)
Frame = +2
Query: 1 MRFMQRVRCWEYRQQSSIVRLTRPTRPDKARRLGYKAKQGYVVYRVRVRRGGRKRPVSKG 60
+RF+ RVR WEYRQ I R+T PTR DKARRLGYKAKQG+V+ RV VRRGGRKRP SKG
Sbjct: 71 LRFLLRVRAWEYRQLPVIHRVTGPTRVDKARRLGYKAKQGFVIVRVAVRRGGRKRPNSKG 250
Query: 61 IVYGKPTNQGVTQLKFQRSKRSVAEERAGRKLGGLRVLNSYWVNEDSTFKYFEVILVDVA 120
IVYGKP + G+ +LKF+R+ RSVAEERAG K LRVLNSYWVN+D+T+KYFE+I+VD +
Sbjct: 251 IVYGKPKHHGINELKFERNLRSVAEERAGNKFTNLRVLNSYWVNQDATYKYFEIIMVDHS 430
Query: 121 HSAIRNDPRINWLTNPVHKHRELRGLTSAGKENRGLSGKGHRYHK-ARPSRRANWKRNNT 179
H IR DPRINW+ N VHK R+ RGLTSAG++ RGL +GHR +K S R NWKR NT
Sbjct: 431 HKVIRRDPRINWIANAVHKGRQQRGLTSAGRQGRGLRKRGHRANKIIGSSFRQNWKRRNT 610
Query: 180 LSLRRYR 186
LSLRRYR
Sbjct: 611 LSLRRYR 631
>BI264231 similar to SP|O65050|R15A 60S ribosomal protein L15-1. [Black
spruce] {Picea mariana}, partial (79%)
Length = 505
Score = 237 bits (604), Expect = 2e-63
Identities = 109/144 (75%), Positives = 125/144 (86%)
Frame = +3
Query: 1 MRFMQRVRCWEYRQQSSIVRLTRPTRPDKARRLGYKAKQGYVVYRVRVRRGGRKRPVSKG 60
+RF+ RVRCWEYRQ S+ RLT P+RPDKARRLGYKAKQGYVVYRVRVRRGGRK+PV G
Sbjct: 72 LRFLLRVRCWEYRQLPSLARLTGPSRPDKARRLGYKAKQGYVVYRVRVRRGGRKKPVKGG 251
Query: 61 IVYGKPTNQGVTQLKFQRSKRSVAEERAGRKLGGLRVLNSYWVNEDSTFKYFEVILVDVA 120
+VYGKP NQG++QLK R+ +SVAEER GRK+G LRVLNSYWVN+DST+KY+EVILVD
Sbjct: 252 VVYGKPVNQGISQLKPTRNLKSVAEERVGRKIGSLRVLNSYWVNQDSTYKYYEVILVDPX 431
Query: 121 HSAIRNDPRINWLTNPVHKHRELR 144
H+AIR DPR NW+ NP HKHRELR
Sbjct: 432 HNAIRRDPRXNWIXNPXHKHRELR 503
>BG649113 similar to GP|18033111|gb functional candidate resistance protein
KR1 {Glycine max}, partial (2%)
Length = 782
Score = 27.3 bits (59), Expect = 3.6
Identities = 12/39 (30%), Positives = 21/39 (53%)
Frame = -3
Query: 137 VHKHRELRGLTSAGKENRGLSGKGHRYHKARPSRRANWK 175
+HKHR+ + L + +GL+ + +HK +A WK
Sbjct: 729 IHKHRDFQDLVESLYFLKGLAVPHNNFHKMVTFTKALWK 613
>TC92064 similar to PIR|T08935|T08935 COP1-interacting protein 7 -
Arabidopsis thaliana, partial (4%)
Length = 844
Score = 26.6 bits (57), Expect = 6.1
Identities = 13/39 (33%), Positives = 18/39 (45%)
Frame = +2
Query: 139 KHRELRGLTSAGKENRGLSGKGHRYHKARPSRRANWKRN 177
KH+ + S K NR S + + H A+P N RN
Sbjct: 260 KHKVEEAVASLEKRNRPTSRQHKKQHSAKPHDMLNGSRN 376
>TC82094 similar to GP|20259195|gb|AAM14313.1 unknown protein {Arabidopsis
thaliana}, partial (83%)
Length = 1759
Score = 26.2 bits (56), Expect = 8.0
Identities = 12/24 (50%), Positives = 14/24 (58%)
Frame = +3
Query: 161 HRYHKARPSRRANWKRNNTLSLRR 184
H Y SRR N+KR TL+L R
Sbjct: 1431 HTYKLLMASRRRNYKREGTLNLPR 1502
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.322 0.136 0.415
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,525,455
Number of Sequences: 36976
Number of extensions: 68619
Number of successful extensions: 347
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 345
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 346
length of query: 186
length of database: 9,014,727
effective HSP length: 90
effective length of query: 96
effective length of database: 5,686,887
effective search space: 545941152
effective search space used: 545941152
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 55 (25.8 bits)
Medicago: description of AC146855.3


