

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
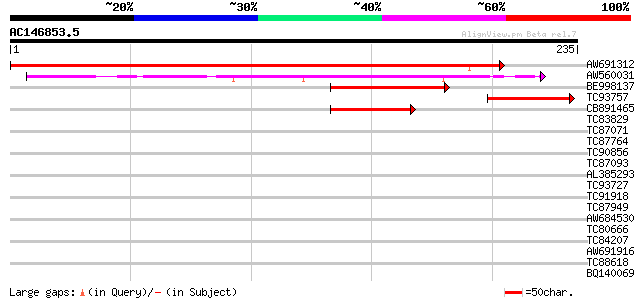
Query= AC146853.5 - phase: 0 /pseudo
(235 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AW691312 similar to GP|14586363|emb lateral root primordia (LRP1... 406 e-114
AW560031 similar to GP|882341|gb|AA LRP1 {Arabidopsis thaliana},... 129 1e-30
BE998137 similar to GP|10177429|db zinc finger protein SHI-like ... 88 2e-18
TC93757 weakly similar to GP|15450615|gb|AAK96579.1 AT3g54430/T1... 73 1e-13
CB891465 similar to PIR|G84600|G846 hypothetical protein At2g214... 62 2e-10
TC83829 similar to GP|18377855|gb|AAL67114.1 AT4g28140/F26K10_20... 35 0.025
TC87071 weakly similar to GP|14532748|gb|AAK64075.1 unknown prot... 35 0.025
TC87764 35 0.033
TC90856 weakly similar to GP|16648921|gb|AAL24312.1 putative pro... 34 0.056
TC87093 homologue to GP|10177817|dbj|BAB11183. gene_id:MKD15.14~... 32 0.16
AL385293 similar to GP|5737842|gb| adenylyl cyclase {Dictyosteli... 32 0.21
TC93727 weakly similar to GP|12963432|gb|AAK11263.1 ribosomal pr... 32 0.28
TC91918 homologue to GP|9369375|gb|AAF87124.1| F10A5.29 {Arabido... 31 0.36
TC87949 similar to GP|16904224|gb|AAL30819.1 calcium/calmodulin-... 31 0.36
AW684530 similar to GP|15724197|gb At2g39050/T7F6.22 {Arabidopsi... 30 0.61
TC80666 similar to PIR|E85112|E85112 probable methyltransferase ... 30 0.61
TC84207 homologue to GP|16649151|gb|AAL24427.1 Unknown protein {... 30 0.61
AW691916 weakly similar to PIR|T05215|T052 peroxidase homolog F1... 30 0.61
TC88618 weakly similar to PIR|T48397|T48397 S-receptor kinase-li... 30 0.61
BQ140069 similar to GP|4031|emb|CAA Negative growth regulatory p... 29 1.8
>AW691312 similar to GP|14586363|emb lateral root primordia (LRP1)
{Arabidopsis thaliana}, partial (14%)
Length = 652
Score = 406 bits (1043), Expect = e-114
Identities = 194/206 (94%), Positives = 197/206 (95%), Gaps = 1/206 (0%)
Frame = +1
Query: 1 MSMLGLRDLVLIAPSPSSLQHHHQQNQNQNQNQNQPISHDHNSNHSLPSSASLSVGFGIF 60
MSMLGLRDLVLIAPSPSSLQHHHQQNQNQNQNQNQPISHDHNSNHSLPSSASLSVGFGIF
Sbjct: 25 MSMLGLRDLVLIAPSPSSLQHHHQQNQNQNQNQNQPISHDHNSNHSLPSSASLSVGFGIF 204
Query: 61 PLLTATPCMPQQQSQNNEVQENPSNNNNFWNLRMCPEVVNLPKKGVITEEENHGKIAMME 120
PLLTATPCMPQQQSQNNEVQENPSNNNNFWNLRMCPEVVNLPKKGVITEEENHGKIAMME
Sbjct: 205 PLLTATPCMPQQQSQNNEVQENPSNNNNFWNLRMCPEVVNLPKKGVITEEENHGKIAMME 384
Query: 121 SEENGVYGSEYRVCQDCGNRAKKDCVFRRCRTCCKGRGYDCSTHLKSTWIPSTRRREREV 180
SEENGVYGSEYRVCQDCGNRAKKDCVFRRCRTCCKGRGYDCSTHLKSTWIPSTRRREREV
Sbjct: 385 SEENGVYGSEYRVCQDCGNRAKKDCVFRRCRTCCKGRGYDCSTHLKSTWIPSTRRREREV 564
Query: 181 EMFAGGGDG-EGCSGVKRQKGLLGSS 205
EMFAGGGDG ++ QKGL+ S
Sbjct: 565 EMFAGGGDG*RVVLVLREQKGLIRGS 642
>AW560031 similar to GP|882341|gb|AA LRP1 {Arabidopsis thaliana}, partial
(24%)
Length = 756
Score = 129 bits (323), Expect = 1e-30
Identities = 81/231 (35%), Positives = 112/231 (48%), Gaps = 16/231 (6%)
Frame = +1
Query: 8 DLVLIAPSPSSLQHHHQQNQNQNQNQNQPISHDHNSNHSLPSSASLSVGFGIFPLLTATP 67
++ + +PSS HHH N + + P HSL + +L G G+ PLLTA+P
Sbjct: 121 EMGMFVVAPSSFHHHHHNNNHHEALMSDP--------HSLNPATAL--GVGVIPLLTASP 270
Query: 68 CMPQQQSQNNEVQENPSNNNNFWN-------LRMCPEVVNLPKKGVITEEENHGKIAMME 120
C+ + N+ + FW+ + P L K+ + N ++
Sbjct: 271 CLENENMLNSRRNQQ---GIQFWHEQHHQQQQQQSPHSHYLKKQQGFLDHHNTSPTNLVH 441
Query: 121 ----SEENGVYGSEYRVCQDCGNRAKKDCVFRRCRTCCKGRGYDCSTHLKSTWIPSTRRR 176
+ G CQDCGN+AKKDC RRCRTCCK RG+DC TH+KSTW+P+ RRR
Sbjct: 442 GGGLTASGTSSGGGTTTCQDCGNQAKKDCSNRRCRTCCKSRGFDCPTHVKSTWVPAARRR 621
Query: 177 ER-----EVEMFAGGGDGEGCSGVKRQKGLLGSSQNAAATSHSSNSNGTTP 222
ER + AGGG G K+ + L+ S TSH+S SN TTP
Sbjct: 622 ERLSSTATTTVVAGGGSSGSTFGAKKPR-LIASQ----TTSHTSTSN-TTP 756
>BE998137 similar to GP|10177429|db zinc finger protein SHI-like {Arabidopsis
thaliana}, partial (36%)
Length = 592
Score = 88.2 bits (217), Expect = 2e-18
Identities = 33/49 (67%), Positives = 42/49 (85%)
Frame = +1
Query: 134 CQDCGNRAKKDCVFRRCRTCCKGRGYDCSTHLKSTWIPSTRRREREVEM 182
CQDCGN+AKKDC RCRTCCK RG+ C TH+KSTW+P++RRRER+ ++
Sbjct: 37 CQDCGNQAKKDCPHMRCRTCCKSRGFQCQTHVKSTWVPASRRRERQQQL 183
>TC93757 weakly similar to GP|15450615|gb|AAK96579.1 AT3g54430/T12E18_120
{Arabidopsis thaliana}, partial (15%)
Length = 391
Score = 72.8 bits (177), Expect = 1e-13
Identities = 35/36 (97%), Positives = 35/36 (97%)
Frame = +1
Query: 199 KGLLGSSQNAAATSHSSNSNGTTPKSFATSSCHQGA 234
KGLLGSSQNAAATSHSSNSNGTTPKSFATSSCHQ A
Sbjct: 1 KGLLGSSQNAAATSHSSNSNGTTPKSFATSSCHQDA 108
>CB891465 similar to PIR|G84600|G846 hypothetical protein At2g21400
[imported] - Arabidopsis thaliana, partial (20%)
Length = 607
Score = 62.0 bits (149), Expect = 2e-10
Identities = 20/35 (57%), Positives = 30/35 (85%)
Frame = -1
Query: 134 CQDCGNRAKKDCVFRRCRTCCKGRGYDCSTHLKST 168
CQDCGN+AKKDC + RCR+CCK + ++C TH++++
Sbjct: 403 CQDCGNQAKKDCAYSRCRSCCKNKDFNCQTHIRTS 299
>TC83829 similar to GP|18377855|gb|AAL67114.1 AT4g28140/F26K10_20
{Arabidopsis thaliana}, partial (9%)
Length = 1054
Score = 35.0 bits (79), Expect = 0.025
Identities = 26/66 (39%), Positives = 32/66 (48%), Gaps = 2/66 (3%)
Frame = +1
Query: 24 QQNQNQNQNQNQPISHDHNSNHSLPSSASLSVGFGIFPLL--TATPCMPQQQSQNNEVQE 81
++NQNQNQNQ QP S PSSAS S +FP + PQQ NN +
Sbjct: 667 RENQNQNQNQFQPF--------SFPSSAS-SSSRHVFPFAFDHNSEQFPQQFGPNNNLPF 819
Query: 82 NPSNNN 87
+P N
Sbjct: 820 HPPTQN 837
>TC87071 weakly similar to GP|14532748|gb|AAK64075.1 unknown protein
{Arabidopsis thaliana}, partial (21%)
Length = 1162
Score = 35.0 bits (79), Expect = 0.025
Identities = 25/83 (30%), Positives = 33/83 (39%), Gaps = 17/83 (20%)
Frame = +1
Query: 20 QHHHQQNQNQNQNQNQPISHDHNSNHSLPSS---------ASLSVGFGIF--------PL 62
QHHHQQ Q Q N P + SN L S L FGIF L
Sbjct: 376 QHHHQQQQQQQPQNNDPPIQSYPSNPQLQFSFRRLFHITPLKLFTFFGIFLFALFLYGSL 555
Query: 63 LTATPCMPQQQSQNNEVQENPSN 85
+ T + + +NN+ + PS+
Sbjct: 556 FSTTTILELRGVKNNDGDDEPSS 624
>TC87764
Length = 1479
Score = 34.7 bits (78), Expect = 0.033
Identities = 20/48 (41%), Positives = 27/48 (55%), Gaps = 1/48 (2%)
Frame = +2
Query: 14 PSPSSLQHHHQQNQNQNQNQNQPISHD-HNSNHSLPSSASLSVGFGIF 60
P S HHHQ NQNQNQNQN + + SN P+ +L+ G++
Sbjct: 11 PRRKSRSHHHQ-NQNQNQNQNHVLPNTLCFSNQLAPNLITLTTKKGLY 151
>TC90856 weakly similar to GP|16648921|gb|AAL24312.1 putative protein
{Arabidopsis thaliana}, partial (25%)
Length = 619
Score = 33.9 bits (76), Expect = 0.056
Identities = 16/36 (44%), Positives = 20/36 (55%), Gaps = 2/36 (5%)
Frame = -2
Query: 21 HHHQQNQNQN--QNQNQPISHDHNSNHSLPSSASLS 54
H H Q QNQN QN N ++H NH LP++ S
Sbjct: 537 HXHAQKQNQNHSQN*NSNVNHSPKQNHHLPNNYPFS 430
>TC87093 homologue to GP|10177817|dbj|BAB11183. gene_id:MKD15.14~unknown
protein {Arabidopsis thaliana}, partial (49%)
Length = 1313
Score = 32.3 bits (72), Expect = 0.16
Identities = 12/24 (50%), Positives = 14/24 (58%)
Frame = +3
Query: 22 HHQQNQNQNQNQNQPISHDHNSNH 45
HHQQ Q Q Q+Q Q H H +H
Sbjct: 615 HHQQQQQQQQHQQQQQQHHHQQHH 686
>AL385293 similar to GP|5737842|gb| adenylyl cyclase {Dictyostelium
discoideum}, partial (1%)
Length = 467
Score = 32.0 bits (71), Expect = 0.21
Identities = 20/67 (29%), Positives = 28/67 (40%)
Frame = +1
Query: 22 HHQQNQNQNQNQNQPISHDHNSNHSLPSSASLSVGFGIFPLLTATPCMPQQQSQNNEVQE 81
H QQ+Q Q Q QNQ S D + LPS +T P +Q N+++
Sbjct: 61 HQQQHQQQQQQQNQQNSQDQSQQIQLPSKDD-----------DSTFIAPTPPTQLNQIRT 207
Query: 82 NPSNNNN 88
N N +
Sbjct: 208 NVVNTGS 228
>TC93727 weakly similar to GP|12963432|gb|AAK11263.1 ribosomal protein P2
{Podospora anserina}, partial (77%)
Length = 442
Score = 31.6 bits (70), Expect = 0.28
Identities = 19/60 (31%), Positives = 25/60 (41%)
Frame = +1
Query: 128 GSEYRVCQDCGNRAKKDCVFRRCRTCCKGRGYDCSTHLKSTWIPSTRRREREVEMFAGGG 187
G + R C + + D +RRCR CC R + W P R R R+ GGG
Sbjct: 166 GGQGR*RAGCSGQGEADEGWRRCRRCCSRR--------RCRWCPRRRCRPRQG*EAQGGG 321
>TC91918 homologue to GP|9369375|gb|AAF87124.1| F10A5.29 {Arabidopsis
thaliana}, partial (38%)
Length = 1171
Score = 31.2 bits (69), Expect = 0.36
Identities = 21/68 (30%), Positives = 32/68 (46%), Gaps = 7/68 (10%)
Frame = +2
Query: 66 TPCMPQQQSQNNEVQENPSNNNNFWN-LRMCPEV------VNLPKKGVITEEENHGKIAM 118
TP QQQ Q+ +Q P+NNNN N +C + N P ++ ++ G I
Sbjct: 341 TPPQMQQQQQHPHIQIQPNNNNNINNSFNICNNINMNNNNNNSPVFVSLSSQQVGGDITP 520
Query: 119 MESEENGV 126
+ S +N V
Sbjct: 521 IYSSDNSV 544
>TC87949 similar to GP|16904224|gb|AAL30819.1 calcium/calmodulin-dependent
protein kinase CaMK2 {Nicotiana tabacum}, partial (62%)
Length = 1413
Score = 31.2 bits (69), Expect = 0.36
Identities = 13/32 (40%), Positives = 18/32 (55%)
Frame = +3
Query: 148 RRCRTCCKGRGYDCSTHLKSTWIPSTRRRERE 179
+RCR CCK CS+ + TW ++R RE
Sbjct: 147 KRCRRCCKADAEGCSS-VSFTWFGTSRHEARE 239
>AW684530 similar to GP|15724197|gb At2g39050/T7F6.22 {Arabidopsis thaliana},
partial (33%)
Length = 643
Score = 30.4 bits (67), Expect = 0.61
Identities = 16/51 (31%), Positives = 20/51 (38%)
Frame = +3
Query: 6 LRDLVLIAPSPSSLQHHHQQNQNQNQNQNQPISHDHNSNHSLPSSASLSVG 56
+ LV PS+ HHH N H HN+NH P S + G
Sbjct: 102 IHHLVTTIYLPSTNHHHHHIINNHLSTLTHHTHHHHNNNHINPKLKSSTPG 254
>TC80666 similar to PIR|E85112|E85112 probable methyltransferase [imported]
- Arabidopsis thaliana, partial (39%)
Length = 1616
Score = 30.4 bits (67), Expect = 0.61
Identities = 16/48 (33%), Positives = 23/48 (47%), Gaps = 6/48 (12%)
Frame = +3
Query: 3 MLGLRDLVLIAPSPSSLQHHHQQNQ------NQNQNQNQPISHDHNSN 44
+L + L+ P + HH QQN+ N NQNQ Q ++ N N
Sbjct: 792 VLNIERARLLREFPETAHHHQQQNESSLGDGNMNQNQQQIVTGSTNVN 935
>TC84207 homologue to GP|16649151|gb|AAL24427.1 Unknown protein {Arabidopsis
thaliana}, partial (4%)
Length = 694
Score = 30.4 bits (67), Expect = 0.61
Identities = 21/75 (28%), Positives = 32/75 (42%), Gaps = 3/75 (4%)
Frame = +1
Query: 20 QHHHQQNQNQNQNQNQPISHDH---NSNHSLPSSASLSVGFGIFPLLTATPCMPQQQSQN 76
++H N+NQNQNQ + H N N+ + VGF + + S N
Sbjct: 217 ENHETSNENQNQNQTFFENQYHRIENWNNETTNDYIKKVGFMVNSQSFLGQGSGSEYSPN 396
Query: 77 NEVQENPSNNNNFWN 91
+Q N S++ WN
Sbjct: 397 ETLQSNFSHSVRTWN 441
>AW691916 weakly similar to PIR|T05215|T052 peroxidase homolog F17I5.60 -
Arabidopsis thaliana, partial (13%)
Length = 658
Score = 30.4 bits (67), Expect = 0.61
Identities = 18/45 (40%), Positives = 25/45 (55%)
Frame = +2
Query: 10 VLIAPSPSSLQHHHQQNQNQNQNQNQPISHDHNSNHSLPSSASLS 54
+L+ SPS HHHQ+N+ Q+ +S H S PS +SLS
Sbjct: 167 LLLFSSPSLQHHHHQENEEILHLQHMDLSCIHCS----PSCSSLS 289
>TC88618 weakly similar to PIR|T48397|T48397 S-receptor kinase-like protein -
Arabidopsis thaliana, partial (38%)
Length = 2304
Score = 30.4 bits (67), Expect = 0.61
Identities = 18/43 (41%), Positives = 25/43 (57%), Gaps = 2/43 (4%)
Frame = -1
Query: 9 LVLIAPSPSSLQHHHQQNQNQNQNQ-NQPISHDHNSN-HSLPS 49
L+L+ +P SL H N N N N NQ +HD N+N + +PS
Sbjct: 1353 LILLFHTPFSLFPPHYTNHNTNNNYPNQNRNHDSNNNPYPIPS 1225
>BQ140069 similar to GP|4031|emb|CAA Negative growth regulatory protein
{Saccharomyces cerevisiae}, partial (3%)
Length = 606
Score = 28.9 bits (63), Expect = 1.8
Identities = 23/81 (28%), Positives = 30/81 (36%), Gaps = 1/81 (1%)
Frame = +3
Query: 10 VLIAPSPSSLQHHHQQNQNQNQNQNQPISHDHNSNH-SLPSSASLSVGFGIFPLLTATPC 68
+LI+ LQ HQ Q Q ++ N SL SS S S
Sbjct: 303 ILISQEDQLLQQQHQHQHQHQQLDQQDFFYESNDRLLSLSSSCSXS-------------- 440
Query: 69 MPQQQSQNNEVQENPSNNNNF 89
Q +E QEN +NN N+
Sbjct: 441 -NSNSDQEDEDQENNNNNCNY 500
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.313 0.128 0.389
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,384,938
Number of Sequences: 36976
Number of extensions: 162350
Number of successful extensions: 1769
Number of sequences better than 10.0: 131
Number of HSP's better than 10.0 without gapping: 1578
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1705
length of query: 235
length of database: 9,014,727
effective HSP length: 93
effective length of query: 142
effective length of database: 5,575,959
effective search space: 791786178
effective search space used: 791786178
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 57 (26.6 bits)
Medicago: description of AC146853.5


