

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
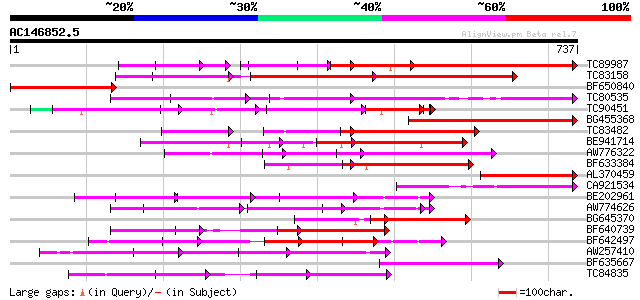
Query= AC146852.5 + phase: 2 /pseudo
(737 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC89987 weakly similar to PIR|D86443|D86443 probable PPR-repeat ... 293 8e-93
TC83158 weakly similar to PIR|H86191|H86191 hypothetical protein... 300 1e-81
BF650840 homologue to SP|O48609|Y65L_ Ycf65-like protein chloro... 273 2e-73
TC80535 weakly similar to GP|14335136|gb|AAK59848.1 At2g15690/F9... 219 4e-57
TC90451 similar to PIR|H84508|H84508 hypothetical protein At2g13... 135 4e-52
BG455368 weakly similar to GP|9759524|dbj selenium-binding prote... 187 2e-47
TC83482 weakly similar to GP|9758453|dbj|BAB08982.1 selenium-bin... 169 5e-42
BE941714 weakly similar to GP|11994734|dbj selenium-binding prot... 161 7e-40
AW776322 weakly similar to PIR|H84859|H84 hypothetical protein A... 149 3e-36
BF633384 similar to PIR|T46179|T46 hypothetical protein T8H10.30... 148 8e-36
AL370459 weakly similar to GP|20146256|d hypothetical protein {O... 139 3e-33
CA921534 similar to PIR|D96516|D96 F16N3.14 [imported] - Arabido... 136 3e-32
BE202961 weakly similar to GP|6016735|gb|A hypothetical protein ... 127 2e-29
AW774626 similar to PIR|T47909|T47 hypothetical protein T20K12.7... 126 3e-29
BG645370 weakly similar to PIR|T00405|T004 hypothetical protein ... 124 1e-28
BF640739 weakly similar to GP|7363286|dbj Similar to Arabidopsis... 120 2e-27
BF642497 weakly similar to PIR|H96713|H96 hypothetical protein T... 110 2e-24
AW257410 weakly similar to GP|10177533|db selenium-binding prote... 110 2e-24
BF635667 weakly similar to PIR|C85359|C853 hypothetical protein ... 106 4e-23
TC84835 weakly similar to PIR|T02328|T02328 hypothetical protein... 103 2e-22
>TC89987 weakly similar to PIR|D86443|D86443 probable PPR-repeat protein
[imported] - Arabidopsis thaliana, partial (62%)
Length = 1559
Score = 293 bits (749), Expect(2) = 8e-93
Identities = 142/321 (44%), Positives = 208/321 (64%)
Frame = +2
Query: 417 LVKAREVFENMPRKNVISWSSMINAFAMHGNADSAIKLFRRMKEVNIEPNGVTFIGVLYA 476
L K VF+NM KN S++ MI+ A+HG A+K+F M E + P+ V ++GV A
Sbjct: 563 LRKGLRVFKNMSEKNRYSYTVMISGLAIHGRGKEALKVFSEMIEEGLAPDDVVYVGVFSA 742
Query: 477 CGHAGLVEEGEKLFSSMINEHGISPTREHYGCMVDLYCRANFLRKAIELIETMPFAPNVI 536
C HAGLVEEG + F SM EH I PT +HYGCMVDL R L++A ELI++M PN +
Sbjct: 743 CSHAGLVEEGLQCFKSMQFEHKIEPTVQHYGCMVDLLGRFGMLKEAYELIKSMSIKPNDV 922
Query: 537 IWGSLMSACQVHGEAELGEFAAKRLLELEPDHDGALVVLSNIYAKEKRWNDVGLIRKSMS 596
IW SL+SAC+VH E+G+ AA+ L L ++ G +VL+N+YAK ++W+DV IR ++
Sbjct: 923 IWRSLLSACKVHHNLEIGKIAAENLFMLNQNNSGDYLVLANMYAKAQKWDDVAKIRTKLA 1102
Query: 597 YKGISKEKASSRIEINNQVHMFMMADRYHKQSDEIYEKLDEVVSKLKLVGYKPSTSGILI 656
+ + + S IE +V+ F+ D+ Q + IYE + ++ ++K GY P TS +L+
Sbjct: 1103ERNLVQTPGFSLIEAKRKVYKFVSQDKSIPQWNIIYEMIHQMEWQVKFEGYIPDTSQVLL 1282
Query: 657 DLEEEDKKELVLWHSEKLAVCYGLISRRNESCIRIVKNLRICEDCHSFMKLVSKVYQIEI 716
D+++E+KKE + +HS+KLA+ +GLI +RI +NLR+C DCH++ K +S +Y+ EI
Sbjct: 1283DVDDEEKKERLKFHSQKLAIAFGLIHTSEGXPLRITRNLRMCSDCHTYTKYISMIYEREI 1462
Query: 717 VVRDRTRFHHCSGGICSCRDY 737
VRDR RF H G CSC+DY
Sbjct: 1463TVRDRXRFXHFKNGSCSCKDY 1525
Score = 74.3 bits (181), Expect(2) = 6e-27
Identities = 43/146 (29%), Positives = 72/146 (48%), Gaps = 1/146 (0%)
Frame = +1
Query: 142 AVSKVSAFNHGLEIHGLASKLGFVDDPFIQTGLIAMYASCRRIMDARLLFDKMCHPDAVA 201
A S + + G+++HG K+G D +Q LI MY C I +A +F+ M +
Sbjct: 130 ACSLLGVVDEGIQVHGHVFKMGLEGDVIVQNSLINMYGKCGEIKNACDVFNGMDEKSVAS 309
Query: 202 WNMIIDGYCQNGHYDDALRLFEDMRSSD-MKPDSVILCTVLSACGHAGNLSYGRTIHEFV 260
W+ II + +++ L L M S + + L VLSAC H G+ G+ IH +
Sbjct: 310 WSAIIGAHACVEMWNECLMLLGKMSSEGRCRVEESTLVNVLSACTHLGSPDLGKCIHGIL 489
Query: 261 KDNGYAIDSHLQTALINMYANCGAMD 286
N ++ ++T+LI+MY G ++
Sbjct: 490 LRNISELNVVVKTSLIDMYVKSGCLE 567
Score = 68.6 bits (166), Expect = 8e-12
Identities = 46/157 (29%), Positives = 76/157 (48%), Gaps = 4/157 (2%)
Frame = +1
Query: 375 ACSHVGALAQANWIHTYVDRSGFGRALSVNNALIDMYAKCGNLVKAREVFENMPRKNVIS 434
ACS +G + + +H +V + G + V N+LI+MY KCG + A +VF M K+V S
Sbjct: 130 ACSLLGVVDEGIQVHGHVFKMGLEGDVIVQNSLINMYGKCGEIKNACDVFNGMDEKSVAS 309
Query: 435 WSSMINAFAMHGNADSAIKLFRRM-KEVNIEPNGVTFIGVLYACGHAGLVEEGEKLFSSM 493
WS++I A A + + L +M E T + VL AC H G + G+ + +
Sbjct: 310 WSAIIGAHACVEMWNECLMLLGKMSSEGRCRVEESTLVNVLSACTHLGSPDLGKCIHGIL 489
Query: 494 ---INEHGISPTREHYGCMVDLYCRANFLRKAIELIE 527
I+E + ++D+Y ++ L K I+
Sbjct: 490 LRNISELNVVVKTS----LIDMYVKSGCLEKRFTRIQ 588
Score = 67.0 bits (162), Expect(2) = 8e-93
Identities = 40/121 (33%), Positives = 69/121 (56%), Gaps = 1/121 (0%)
Frame = +1
Query: 300 LIVSTAMLSGYAKLGMVKDARFIFDQMIERDLVCWSAMISGYAESDQPQEALKLFDEMLQ 359
+IV ++++ Y K G +K+A +F+ M E+ + WSA+I +A + E L L +M
Sbjct: 208 VIVQNSLINMYGKCGEIKNACDVFNGMDEKSVASWSAIIGAHACVEMWNECLMLLGKMSS 387
Query: 360 K-RSVPDQITMLSVISACSHVGALAQANWIHTYVDRSGFGRALSVNNALIDMYAKCGNLV 418
+ R ++ T+++V+SAC+H+G+ IH + R+ + V +LIDMY K G L
Sbjct: 388 EGRCRVEESTLVNVLSACTHLGSPDLGKCIHGILLRNISELNVVVKTSLIDMYVKSGCLE 567
Query: 419 K 419
K
Sbjct: 568 K 570
Score = 65.5 bits (158), Expect(2) = 6e-27
Identities = 37/141 (26%), Positives = 74/141 (52%), Gaps = 2/141 (1%)
Frame = +2
Query: 311 AKLGMVKDARFIFDQMIERDLVCWSAMISGYAESDQPQEALKLFDEMLQKRSVPDQITML 370
+++ +++ +F M E++ ++ MISG A + +EALK+F EM+++ PD + +
Sbjct: 548 SRVDVLRKGLRVFKNMSEKNRYSYTVMISGLAIHGRGKEALKVFSEMIEEGLAPDDVVYV 727
Query: 371 SVISACSHVGALAQA-NWIHTYVDRSGFGRALSVNNALIDMYAKCGNLVKAREVFENMPR 429
V SACSH G + + + + ++D+ + G L +A E+ ++M
Sbjct: 728 GVFSACSHAGLVEEGLQCFKSMQFEHKIEPTVQHYGCMVDLLGRFGMLKEAYELIKSMSI 907
Query: 430 K-NVISWSSMINAFAMHGNAD 449
K N + W S+++A +H N +
Sbjct: 908 KPNDVIWRSLLSACKVHHNLE 970
Score = 43.9 bits (102), Expect = 2e-04
Identities = 20/64 (31%), Positives = 34/64 (52%)
Frame = +2
Query: 190 LFDKMCHPDAVAWNMIIDGYCQNGHYDDALRLFEDMRSSDMKPDSVILCTVLSACGHAGN 249
+F M + ++ ++I G +G +AL++F +M + PD V+ V SAC HAG
Sbjct: 581 VFKNMSEKNRYSYTVMISGLAIHGRGKEALKVFSEMIEEGLAPDDVVYVGVFSACSHAGL 760
Query: 250 LSYG 253
+ G
Sbjct: 761 VEEG 772
Score = 38.9 bits (89), Expect = 0.007
Identities = 56/271 (20%), Positives = 106/271 (38%), Gaps = 16/271 (5%)
Frame = +2
Query: 107 LSRSSFPEKTIFLYHNLRAINAFALDRFSFPSLLKAVSKVSAFNHGLEIHGLASKLGFVD 166
+SR K + ++ N+ N R+S+ ++ ++ L++ + G
Sbjct: 545 MSRVDVLRKGLRVFKNMSEKN-----RYSYTVMISGLAIHGRGKEALKVFSEMIEEGLAP 709
Query: 167 DPFIQTGLIAMYASCRRIMDARLLFDKM-----CHPDAVAWNMIIDGYCQNGHYDDALRL 221
D + G+ + + + + F M P + ++D + G +A
Sbjct: 710 DDVVYVGVFSACSHAGLVEEGLQCFKSMQFEHKIEPTVQHYGCMVDLLGRFGMLKEA--- 880
Query: 222 FEDMRSSDMKPDSVILCTVLSACGHAGNLSYGRTIHE--FVKDNGYAIDSHLQTALINMY 279
+E ++S +KP+ VI ++LSAC NL G+ E F+ + + D L NMY
Sbjct: 881 YELIKSMSIKPNDVIWRSLLSACKVHHNLEIGKIAAENLFMLNQNNSGD---YLVLANMY 1051
Query: 280 ANCGAMDLARKIYDGLSSKHLIVSTAMLSGYAKLGMVK---------DARFIFDQMIERD 330
A D KI L+ ++L+ + AK + K I++ + + +
Sbjct: 1052AKAQKWDDVAKIRTKLAERNLVQTPGFSLIEAKRKVYKFVSQDKSIPQWNIIYEMIHQME 1231
Query: 331 LVCWSAMISGYAESDQPQEALKLFDEMLQKR 361
W GY D Q L + DE ++R
Sbjct: 1232---WQVKFEGYI-PDTSQVLLDVDDEEKKER 1312
>TC83158 weakly similar to PIR|H86191|H86191 hypothetical protein [imported]
- Arabidopsis thaliana, partial (42%)
Length = 1194
Score = 300 bits (768), Expect = 1e-81
Identities = 145/347 (41%), Positives = 227/347 (64%)
Frame = +2
Query: 314 GMVKDARFIFDQMIERDLVCWSAMISGYAESDQPQEALKLFDEMLQKRSVPDQITMLSVI 373
G V DA +FD++ +++V W+ +I G+ + + +EAL+ F EM VPD +T++++I
Sbjct: 2 GDVDDALKLFDKLPVKNVVSWTVVIGGFVKKECYEEALECFREMQLAGVVPDFVTVIAII 181
Query: 374 SACSHVGALAQANWIHTYVDRSGFGRALSVNNALIDMYAKCGNLVKAREVFENMPRKNVI 433
SAC+++GAL W+H V + F + V N+LIDMYA+CG + AR+VF+ M ++N++
Sbjct: 182 SACANLGALGLGLWVHRLVMKKEFRDNVKVLNSLIDMYARCGCIELARQVFDGMSQRNLV 361
Query: 434 SWSSMINAFAMHGNADSAIKLFRRMKEVNIEPNGVTFIGVLYACGHAGLVEEGEKLFSSM 493
SW+S+I FA++G AD A+ FR MK+ +EPNGV++ L AC HAGL++EG K+F+ +
Sbjct: 362 SWNSIIVGFAVNGLADKALSFFRSMKKEGLEPNGVSYTSALTACSHAGLIDEGLKIFADI 541
Query: 494 INEHGISPTREHYGCMVDLYCRANFLRKAIELIETMPFAPNVIIWGSLMSACQVHGEAEL 553
+H SP EHYGC+VDLY RA L++A ++I+ MP PN ++ GSL++AC+ G+ EL
Sbjct: 542 KRDHRNSPRIEHYGCLVDLYSRAGRLKEAWDVIKKMPMMPNEVVLGSLLAACRTQGDVEL 721
Query: 554 GEFAAKRLLELEPDHDGALVVLSNIYAKEKRWNDVGLIRKSMSYKGISKEKASSRIEINN 613
E K +EL P D V+ SNIYA +W+ +R+ M +G+ K A S IEI++
Sbjct: 722 AEKVMKYQVELYPGGDSNYVLFSNIYAAVGKWDGASKVRREMKERGLQKNLAFSSIEIDS 901
Query: 614 QVHMFMMADRYHKQSDEIYEKLDEVVSKLKLVGYKPSTSGILIDLEE 660
+H F+ D+YH+++D IY L+ + +L L GY P SG D+++
Sbjct: 902 GIHKFVSGDKYHEENDYIYSALELLSFELHLYGYVPDFSGKESDVDD 1042
Score = 124 bits (312), Expect = 1e-28
Identities = 78/297 (26%), Positives = 144/297 (48%), Gaps = 4/297 (1%)
Frame = +2
Query: 186 DARLLFDKMCHPDAVAWNMIIDGYCQNGHYDDALRLFEDMRSSDMKPDSVILCTVLSACG 245
DA LFDK+ + V+W ++I G+ + Y++AL F +M+ + + PD V + ++SAC
Sbjct: 14 DALKLFDKLPVKNVVSWTVVIGGFVKKECYEEALECFREMQLAGVVPDFVTVIAIISACA 193
Query: 246 HAGNLSYGRTIHEFVKDNGYAIDSHLQTALINMYANCGAMDLARKIYDGLSSKHLIVSTA 305
+ G L G +H V + + + +LI+MYA CG ++LAR+++DG+S ++L+
Sbjct: 194 NLGALGLGLWVHRLVMKKEFRDNVKVLNSLIDMYARCGCIELARQVFDGMSQRNLV---- 361
Query: 306 MLSGYAKLGMVKDARFIFDQMIERDLVCWSAMISGYAESDQPQEALKLFDEMLQKRSVPD 365
W+++I G+A + +AL F M ++ P+
Sbjct: 362 ---------------------------SWNSIIVGFAVNGLADKALSFFRSMKKEGLEPN 460
Query: 366 QITMLSVISACSHVGALAQANWIHTYVDRSGFGR-ALSVNNALIDMYAKCGNLVKAREVF 424
++ S ++ACSH G + + I + R + L+D+Y++ G L +A +V
Sbjct: 461 GVSYTSALTACSHAGLIDEGLKIFADIKRDHRNSPRIEHYGCLVDLYSRAGRLKEAWDVI 640
Query: 425 ENMP-RKNVISWSSMINAFAMHGNADSAIKLFRRMKEV--NIEPNGVTFIGVLYACG 478
+ MP N + S++ A G+ + A K+ + E+ + N V F + A G
Sbjct: 641 KKMPMMPNEVVLGSLLAACRTQGDVELAEKVMKYQVELYPGGDSNYVLFSNIYAAVG 811
Score = 83.2 bits (204), Expect = 3e-16
Identities = 52/159 (32%), Positives = 81/159 (50%), Gaps = 5/159 (3%)
Frame = +2
Query: 138 SLLKAVSKVSAFNHGLEIHGLASKLGFVDDPFIQTGLIAMYASCRRIMDARLLFDKMCHP 197
+++ A + + A GL +H L K F D+ + LI MYA C I AR +FD M
Sbjct: 173 AIISACANLGALGLGLWVHRLVMKKEFRDNVKVLNSLIDMYARCGCIELARQVFDGMSQR 352
Query: 198 DAVAWNMIIDGYCQNGHYDDALRLFEDMRSSDMKPDSVILCTVLSACGHAGNLSYGRTIH 257
+ V+WN II G+ NG D AL F M+ ++P+ V + L+AC HAG + G I
Sbjct: 353 NLVSWNSIIVGFAVNGLADKALSFFRSMKKEGLEPNGVSYTSALTACSHAGLIDEGLKIF 532
Query: 258 EFVK-DNGYAIDSHLQTALINMYANCG----AMDLARKI 291
+K D+ + L+++Y+ G A D+ +K+
Sbjct: 533 ADIKRDHRNSPRIEHYGCLVDLYSRAGRLKEAWDVIKKM 649
>BF650840 homologue to SP|O48609|Y65L_ Ycf65-like protein chloroplast
precursor. [Barley] {Hordeum vulgare}, partial (8%)
Length = 444
Score = 273 bits (697), Expect = 2e-73
Identities = 139/139 (100%), Positives = 139/139 (100%)
Frame = +3
Query: 1 *YLSSLLSMAIEMAMTTMSHTHNRITTHPQLLTTLSSSTTLSHLKQIHAQILHSNTTPEN 60
*YLSSLLSMAIEMAMTTMSHTHNRITTHPQLLTTLSSSTTLSHLKQIHAQILHSNTTPEN
Sbjct: 27 *YLSSLLSMAIEMAMTTMSHTHNRITTHPQLLTTLSSSTTLSHLKQIHAQILHSNTTPEN 206
Query: 61 TNTLLSKLALSICTLSSSSSSLHYALSVFSQIPNPHTHFSNQLLRHLSRSSFPEKTIFLY 120
TNTLLSKLALSICTLSSSSSSLHYALSVFSQIPNPHTHFSNQLLRHLSRSSFPEKTIFLY
Sbjct: 207 TNTLLSKLALSICTLSSSSSSLHYALSVFSQIPNPHTHFSNQLLRHLSRSSFPEKTIFLY 386
Query: 121 HNLRAINAFALDRFSFPSL 139
HNLRAINAFALDRFSFPSL
Sbjct: 387 HNLRAINAFALDRFSFPSL 443
>TC80535 weakly similar to GP|14335136|gb|AAK59848.1 At2g15690/F9O13.24
{Arabidopsis thaliana}, partial (55%)
Length = 1727
Score = 219 bits (557), Expect = 4e-57
Identities = 124/401 (30%), Positives = 214/401 (52%), Gaps = 1/401 (0%)
Frame = +2
Query: 338 ISGYAESDQPQEALKLFDEMLQKRSVPDQITMLSVISACSHVGALAQANWIHTYVDRSGF 397
++ + + + +EAL E+++K D + C ++ A +H Y +S F
Sbjct: 209 LTRFCQEGKVKEAL----ELMEKGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTF 376
Query: 398 GRALSVNNALIDMYAKCGNLVKAREVFENMPRKNVISWSSMINAFAMHGNADSAIKLFRR 457
++N +I+MY C ++ AR VF++MP +N+ SW MI +A D ++LF +
Sbjct: 377 RSDFKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQ 556
Query: 458 MKEVNIEPNGVTFIGVLYACGHAGLVEEGEKLFSSMINEHGISPTREHYGCMVDLYCRAN 517
M E+ +E T + VL ACG A VE+ SM +++GI P EHY ++D+ ++
Sbjct: 557 MNELGLEITSETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSG 736
Query: 518 FLRKAIELIETMPFAPNVIIWGSLMSACQVHGEAELGEFAAKRLLELEPDHDGALVVLSN 577
+L++A E IE +PF P V ++ +L + ++HG+ +L + + ++ L+P + V +
Sbjct: 737 YLKEAEEFIEQLPFEPTVTVFETLKNYARIHGDVDLEDHVEELIVSLDP----SKAVANK 904
Query: 578 IYAKEKRWNDVGLIRKSMSYKGISKEKASSR-IEINNQVHMFMMADRYHKQSDEIYEKLD 636
I + Y IS +R IE N +K +++
Sbjct: 905 IPTPPPK-----------KYTAISMLDGKNRIIEYKNPT--------LYKDDEKLI---- 1015
Query: 637 EVVSKLKLVGYKPSTSGILIDLEEEDKKELVLWHSEKLAVCYGLISRRNESCIRIVKNLR 696
++ +K GY P T +L D+++E K++ +L+HSE+LA+ YGLIS + +RI+KNLR
Sbjct: 1016-AMNSMKDAGYVPDTRYVLHDIDQEAKEQALLYHSERLAIAYGLISTPPRTPLRIIKNLR 1192
Query: 697 ICEDCHSFMKLVSKVYQIEIVVRDRTRFHHCSGGICSCRDY 737
+C DCH+ +K++S++ E++VRD RFHH G CSC DY
Sbjct: 1193VCGDCHNAIKIMSRIVGRELIVRDNKRFHHFKDGKCSCGDY 1315
Score = 74.3 bits (181), Expect = 2e-13
Identities = 59/244 (24%), Positives = 106/244 (43%), Gaps = 3/244 (1%)
Frame = +2
Query: 209 YCQNGHYDDALRLFEDMRSSDMKPDSVILCTVLSACGHAGNLSYGRTIHEFVKDNGYAID 268
+CQ G +AL L E +K D+ + CG + ++ + +H++ + + D
Sbjct: 218 FCQEGKVKEALELME----KGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSD 385
Query: 269 SHLQTALINMYANCGAMDLARKIYDGLSSKHLIVSTAMLSGYAKLGMVKDARFIFDQMIE 328
+ +I MY NC +M AR+++D + ++++ M+ GYA M + +F+QM E
Sbjct: 386 FKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNE 565
Query: 329 RDLVCWSAMISGYAESDQPQEALKLFDEMLQKRSVPDQITMLSVISACSHVGALAQANWI 388
L S TML+V+SAC A+ A +I
Sbjct: 566 LGLEITSE-------------------------------TMLAVLSACGSAEAVEDA-YI 649
Query: 389 HTYVDRSGFGRALSVNN--ALIDMYAKCGNLVKAREVFENMP-RKNVISWSSMINAFAMH 445
+ +S +G V + L+D+ + G L +A E E +P V + ++ N +H
Sbjct: 650 YLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIEQLPFEPTVTVFETLKNYARIH 829
Query: 446 GNAD 449
G+ D
Sbjct: 830 GDVD 841
Score = 67.0 bits (162), Expect = 2e-11
Identities = 47/184 (25%), Positives = 83/184 (44%), Gaps = 2/184 (1%)
Frame = +2
Query: 132 DRFSFPSLLKAVSKVSAFNHGLEIHGLASKLGFVDDPFIQTGLIAMYASCRRIMDARLLF 191
D F L K + ++H + F D + +I MY +C+ + DAR +F
Sbjct: 278 DANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYGNCKSMTDARRVF 457
Query: 192 DKMCHPDAVAWNMIIDGYCQNGHYDDALRLFEDMRSSDMKPDSVILCTVLSACGHAGNLS 251
D M + + +W+M+I GY + D+ L+LFE M ++ S + VLSACG A +
Sbjct: 458 DHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLEITSETMLAVLSACGSAEAVE 637
Query: 252 YGRTIHEFVKDNGYAIDSHLQ--TALINMYANCGAMDLARKIYDGLSSKHLIVSTAMLSG 309
E +K Y I+ ++ L+++ G + A + + L + + L
Sbjct: 638 DAYIYLESMKSK-YGIEPGVEHYMGLLDVLGQSGYLKEAEEFIEQLPFEPTVTVFETLKN 814
Query: 310 YAKL 313
YA++
Sbjct: 815 YARI 826
>TC90451 similar to PIR|H84508|H84508 hypothetical protein At2g13600
[imported] - Arabidopsis thaliana, partial (12%)
Length = 1362
Score = 135 bits (339), Expect(3) = 4e-52
Identities = 85/270 (31%), Positives = 131/270 (48%), Gaps = 2/270 (0%)
Frame = +2
Query: 196 HPDAVAWNMIIDGYCQNGHYDDALRLFEDMRSSDMKPDSVILCTVLSACGHAGNLSYGRT 255
+P + WN II Y + +ALR++ M + + PD L VL A + + G+
Sbjct: 371 NPASFNWNNIIRSYTRLESPQNALRIYVSMLRAGVLPDRYTLPIVLKAVSQSFAIQLGQQ 550
Query: 256 IHEFVKDNGYAIDSHLQTALINMYANCGAMDLARKIYDGLSSKHLIVSTAMLSGYAKLGM 315
+H + G + + ++ IN+Y G D A K+
Sbjct: 551 VHSYGIKLGLQSNEYCESGFINLYCKAGDFDSAHKV------------------------ 658
Query: 316 VKDARFIFDQMIERDLVCWSAMISGYAESDQPQEALKLFDEMLQKRSVPDQITMLSVISA 375
FD+ E L W+A+ISG ++ +A+ +F +M + PD ITM+SV+SA
Sbjct: 659 -------FDENHEPKLGSWNALISGLSQGGLAMDAIVVFVDMKRHGFEPDGITMVSVMSA 817
Query: 376 CSHVGALAQANWIHTYVDRSGFGR--ALSVNNALIDMYAKCGNLVKAREVFENMPRKNVI 433
C +G L A +H YV ++ + ++N+LIDMY KCG + A EVF M +NV
Sbjct: 818 CGSIGDLYLALQLHKYVFQAKTNEWTVILMSNSLIDMYGKCGRMDLAYEVFATMEDRNVS 997
Query: 434 SWSSMINAFAMHGNADSAIKLFRRMKEVNI 463
SW+SMI +AMHG+A A+ F MK V +
Sbjct: 998 SWTSMIVGYAMHGHAKEALGCFHCMKGVGV 1087
Score = 128 bits (322), Expect = 7e-30
Identities = 79/281 (28%), Positives = 142/281 (50%), Gaps = 10/281 (3%)
Frame = +2
Query: 56 TTPENTNTLLSKLALSICTLSSSSSSLHYALSVFSQI-------PNPHTHFSNQLLRHLS 108
+ P+++ T + I TL S+++ + +++ I NP + N ++R +
Sbjct: 236 SVPQHSITSGNDPVTVIATLLSNTTRIRDLNQIYAHILLTRFLESNPASFNWNNIIRSYT 415
Query: 109 RSSFPEKTIFLYHNLRAINAFALDRFSFPSLLKAVSKVSAFNHGLEIHGLASKLGFVDDP 168
R P+ + +Y ++ DR++ P +LKAVS+ A G ++H KLG +
Sbjct: 416 RLESPQNALRIYVSMLRAGVLP-DRYTLPIVLKAVSQSFAIQLGQQVHSYGIKLGLQSNE 592
Query: 169 FIQTGLIAMYASCRRIMDARLLFDKMCHPDAVAWNMIIDGYCQNGHYDDALRLFEDMRSS 228
+ ++G I +Y A +FD+ P +WN +I G Q G DA+ +F DM+
Sbjct: 593 YCESGFINLYCKAGDFDSAHKVFDENHEPKLGSWNALISGLSQGGLAMDAIVVFVDMKRH 772
Query: 229 DMKPDSVILCTVLSACGHAGNLSYGRTIHEFV---KDNGYAIDSHLQTALINMYANCGAM 285
+PD + + +V+SACG G+L +H++V K N + + + +LI+MY CG M
Sbjct: 773 GFEPDGITMVSVMSACGSIGDLYLALQLHKYVFQAKTNEWTV-ILMSNSLIDMYGKCGRM 949
Query: 286 DLARKIYDGLSSKHLIVSTAMLSGYAKLGMVKDARFIFDQM 326
DLA +++ + +++ T+M+ GYA G K+A F M
Sbjct: 950 DLAYEVFATMEDRNVSSWTSMIVGYAMHGHAKEALGCFHCM 1072
Score = 97.8 bits (242), Expect = 1e-20
Identities = 61/223 (27%), Positives = 109/223 (48%), Gaps = 4/223 (1%)
Frame = +2
Query: 334 WSAMISGYAESDQPQEALKLFDEMLQKRSVPDQITMLSVISACSHVGALAQANWIHTYVD 393
W+ +I Y + PQ AL+++ ML+ +PD+ T+ V+ A S A+ +H+Y
Sbjct: 389 WNNIIRSYTRLESPQNALRIYVSMLRAGVLPDRYTLPIVLKAVSQSFAIQLGQQVHSYGI 568
Query: 394 RSGFGRALSVNNALIDMYAKCGNLVKAREVFENMPRKNVISWSSMINAFAMHGNADSAIK 453
+ G + I++Y K G+ A +VF+ + SW+++I+ + G A AI
Sbjct: 569 KLGLQSNEYCESGFINLYCKAGDFDSAHKVFDENHEPKLGSWNALISGLSQGGLAMDAIV 748
Query: 454 LFRRMKEVNIEPNGVTFIGVLYACGHAG----LVEEGEKLFSSMINEHGISPTREHYGCM 509
+F MK EP+G+T + V+ ACG G ++ + +F + NE + +
Sbjct: 749 VFVDMKRHGFEPDGITMVSVMSACGSIGDLYLALQLHKYVFQAKTNEWTVILMS---NSL 919
Query: 510 VDLYCRANFLRKAIELIETMPFAPNVIIWGSLMSACQVHGEAE 552
+D+Y + + A E+ TM NV W S++ +HG A+
Sbjct: 920 IDMYGKCGRMDLAYEVFATME-DRNVSSWTSMIVGYAMHGHAK 1045
Score = 87.8 bits (216), Expect(3) = 4e-52
Identities = 40/77 (51%), Positives = 50/77 (63%)
Frame = +1
Query: 463 IEPNGVTFIGVLYACGHAGLVEEGEKLFSSMINEHGISPTREHYGCMVDLYCRANFLRKA 522
++PN VTFIGVL AC H G V+EG F M N +GI+P +HYGCMVDL RA A
Sbjct: 1087 VKPNYVTFIGVLSACVHGGTVQEGRFYFDMMKNIYGITPQLQHYGCMVDLLGRAGLFDDA 1266
Query: 523 IELIETMPFAPNVIIWG 539
++E MP PN ++WG
Sbjct: 1267 RRMVEEMPMKPNSVVWG 1317
Score = 43.5 bits (101), Expect = 3e-04
Identities = 44/202 (21%), Positives = 81/202 (39%), Gaps = 2/202 (0%)
Frame = +2
Query: 27 THPQLLTTLSSSTTLSHLKQIHAQILHSNTTPENTNTLLSKLALSICTLSSSSSSLHYAL 86
T P +L +S S + +Q+H+ + +N +++ + S H
Sbjct: 491 TLPIVLKAVSQSFAIQLGQQVHSYGIKLGL---QSNEYCESGFINLYCKAGDFDSAH--- 652
Query: 87 SVFSQIPNPHTHFSNQLLRHLSRSSFPEKTIFLYHNLRAINAFALDRFSFPSLLKAVSKV 146
VF + P N L+ LS+ I ++ +++ + F D + S++ A +
Sbjct: 653 KVFDENHEPKLGSWNALISGLSQGGLAMDAIVVFVDMKR-HGFEPDGITMVSVMSACGSI 829
Query: 147 SAFNHGLEIHGLA--SKLGFVDDPFIQTGLIAMYASCRRIMDARLLFDKMCHPDAVAWNM 204
L++H +K + LI MY C R+ A +F M + +W
Sbjct: 830 GDLYLALQLHKYVFQAKTNEWTVILMSNSLIDMYGKCGRMDLAYEVFATMEDRNVSSWTS 1009
Query: 205 IIDGYCQNGHYDDALRLFEDMR 226
+I GY +GH +AL F M+
Sbjct: 1010MIVGYAMHGHAKEALGCFHCMK 1075
Score = 40.0 bits (92), Expect = 0.003
Identities = 29/91 (31%), Positives = 45/91 (48%), Gaps = 2/91 (2%)
Frame = +1
Query: 210 CQNGHYDDALRLFEDMRSSDMKPDSVILCTVLSACGHAGNLSYGRTIHEFVKDNGYAIDS 269
C+ G + L L+E R +KP+ V VLSAC H G + GR + +K N Y I
Sbjct: 1039 CKGGTW--LLSLYEGSRG--VKPNYVTFIGVLSACVHGGTVQEGRFYFDMMK-NIYGITP 1203
Query: 270 HLQ--TALINMYANCGAMDLARKIYDGLSSK 298
LQ ++++ G D AR++ + + K
Sbjct: 1204 QLQHYGCMVDLLGRAGLFDDARRMVEEMPMK 1296
Score = 21.9 bits (45), Expect(3) = 4e-52
Identities = 7/15 (46%), Positives = 10/15 (66%)
Frame = +2
Query: 539 GSLMSACQVHGEAEL 553
G LM AC+ HG ++
Sbjct: 1316 GCLMGACEKHGNVDM 1360
>BG455368 weakly similar to GP|9759524|dbj selenium-binding protein-like
{Arabidopsis thaliana}, partial (33%)
Length = 687
Score = 187 bits (474), Expect = 2e-47
Identities = 91/220 (41%), Positives = 138/220 (62%), Gaps = 1/220 (0%)
Frame = +1
Query: 519 LRKAIELIETMPFAPNVIIWGSLMSACQVHGEAELGEFAAKRLLELEPDHDGALVVLSNI 578
+ KA++L++TMP PN +WG+L+ A +HG ++ E A++ L ELEPD+ G ++LS
Sbjct: 7 VEKALQLVQTMPMEPNGGVWGALLGASHIHGNPDVAEIASRSLFELEPDNLGNYLLLSKT 186
Query: 579 YAKEKRWNDVGLIRKSMSYKGISKEKASSRIEINNQV-HMFMMADRYHKQSDEIYEKLDE 637
YA +W+DV +RK M K + K S +E N + H F D H + +EI + LD+
Sbjct: 187 YALAAKWDDVSRVRKLMREKQLRKNPGCSWVEAKNGIIHEFFAGDVKHPEINEIKKALDD 366
Query: 638 VVSKLKLVGYKPSTSGILIDLEEEDKKELVLWHSEKLAVCYGLISRRNESCIRIVKNLRI 697
++ +LK GY+P + + D+++E K+ L++ HSEKLA+ YGL+S S I+I+KNLRI
Sbjct: 367 LLQRLKCTGYQPKLNSVPYDIDDEGKRCLLVSHSEKLALAYGLLSTDAGSTIKIMKNLRI 546
Query: 698 CEDCHSFMKLVSKVYQIEIVVRDRTRFHHCSGGICSCRDY 737
CEDCH M S + +I+VRD FHH G CSC ++
Sbjct: 547 CEDCHIVMCGASXLTGRKIIVRDNMXFHHFLNGACSCNNF 666
>TC83482 weakly similar to GP|9758453|dbj|BAB08982.1 selenium-binding
protein-like {Arabidopsis thaliana}, partial (10%)
Length = 895
Score = 169 bits (427), Expect = 5e-42
Identities = 84/180 (46%), Positives = 120/180 (66%)
Frame = +2
Query: 431 NVISWSSMINAFAMHGNADSAIKLFRRMKEVNIEPNGVTFIGVLYACGHAGLVEEGEKLF 490
+++SW+++I +A HG A A++ F RMK V P+ VTF+ VL AC HAGLVEEGEK F
Sbjct: 2 DLVSWNAIIGGYASHGLATRALEEFDRMKVVGT-PDEVTFVNVLSACVHAGLVEEGEKHF 178
Query: 491 SSMINEHGISPTREHYGCMVDLYCRANFLRKAIELIETMPFAPNVIIWGSLMSACQVHGE 550
+ M+ ++GI EHY CMVDLY RA +A LI+ MPF P+V++WG+L++AC +H
Sbjct: 179 TDMLTKYGIQAEMEHYSCMVDLYGRAGRFDEAENLIKNMPFEPDVVLWGALLAACGLHSN 358
Query: 551 AELGEFAAKRLLELEPDHDGALVVLSNIYAKEKRWNDVGLIRKSMSYKGISKEKASSRIE 610
ELGE+AA+R+ LE H + VLS I ++ W+ V +R +M +GI K+ A S +E
Sbjct: 359 LELGEYAAERIRRLESSHPVSYSVLSKIQGEKGVWSSVNELRDTMKERGIKKQTAISWVE 538
Score = 71.6 bits (174), Expect = 1e-12
Identities = 37/122 (30%), Positives = 66/122 (53%), Gaps = 2/122 (1%)
Frame = +2
Query: 330 DLVCWSAMISGYAESDQPQEALKLFDEMLQKRSVPDQITMLSVISACSHVGALAQANWIH 389
DLV W+A+I GYA AL+ FD M + PD++T ++V+SAC H G + +
Sbjct: 2 DLVSWNAIIGGYASHGLATRALEEFDRM-KVVGTPDEVTFVNVLSACVHAGLVEEGEKHF 178
Query: 390 T-YVDRSGFGRALSVNNALIDMYAKCGNLVKAREVFENMP-RKNVISWSSMINAFAMHGN 447
T + + G + + ++D+Y + G +A + +NMP +V+ W +++ A +H N
Sbjct: 179 TDMLTKYGIQAEMEHYSCMVDLYGRAGRFDEAENLIKNMPFEPDVVLWGALLAACGLHSN 358
Query: 448 AD 449
+
Sbjct: 359 LE 364
Score = 49.7 bits (117), Expect = 4e-06
Identities = 30/95 (31%), Positives = 45/95 (46%), Gaps = 1/95 (1%)
Frame = +2
Query: 198 DAVAWNMIIDGYCQNGHYDDALRLFEDMRSSDMKPDSVILCTVLSACGHAGNLSYG-RTI 256
D V+WN II GY +G AL F+ M+ PD V VLSAC HAG + G +
Sbjct: 2 DLVSWNAIIGGYASHGLATRALEEFDRMKVVG-TPDEVTFVNVLSACVHAGLVEEGEKHF 178
Query: 257 HEFVKDNGYAIDSHLQTALINMYANCGAMDLARKI 291
+ + G + + ++++Y G D A +
Sbjct: 179 TDMLTKYGIQAEMEHYSCMVDLYGRAGRFDEAENL 283
Score = 33.5 bits (75), Expect = 0.30
Identities = 18/60 (30%), Positives = 33/60 (55%)
Frame = +2
Query: 202 WNMIIDGYCQNGHYDDALRLFEDMRSSDMKPDSVILCTVLSACGHAGNLSYGRTIHEFVK 261
++ ++D Y + G +D+A L ++M +PD V+ +L+ACG NL G E ++
Sbjct: 224 YSCMVDLYGRAGRFDEAENLIKNM---PFEPDVVLWGALLAACGLHSNLELGEYAAERIR 394
>BE941714 weakly similar to GP|11994734|dbj selenium-binding protein-like
{Arabidopsis thaliana}, partial (10%)
Length = 619
Score = 161 bits (408), Expect = 7e-40
Identities = 78/201 (38%), Positives = 127/201 (62%), Gaps = 5/201 (2%)
Frame = +2
Query: 400 ALSVNNALIDMYAKCGNLVKAREVFENMPRKNVISWSSMINAFAMHGNADSAIKLFRRMK 459
++ ++ +L+DMYAKCGNL A+ +F++M ++V+ W++MI+ AMHG+ A+KLF M+
Sbjct: 11 SVRLSTSLLDMYAKCGNLELAKRLFDSMNMRDVVCWNAMISGMAMHGDGKGALKLFYDME 190
Query: 460 EVNIEPNGVTFIGVLYACGHAGLVEEGEKLFSSMINEHGISPTREHYGCMVDLYCRANFL 519
+V ++P+ +TFI V AC ++G+ EG L M + + I P EHYGC+VDL RA
Sbjct: 191 KVGVKPDDITFIAVFTACSYSGMAYEGLMLLDKMCSVYNIVPKSEHYGCLVDLLSRAGLF 370
Query: 520 RKAIELIETMPFAPN----VIIWGSLMSACQVHGEAELGEFAAKRLLELEPD-HDGALVV 574
+A+ +I + + N + W + +SAC HGE +L E AA+++L+L+ H G V+
Sbjct: 371 EEAMVMIRKITNSWNGSEETLAWRAFLSACCNHGETQLAELAAEKVLQLDNHIHSGVYVL 550
Query: 575 LSNIYAKEKRWNDVGLIRKSM 595
LSN+YA + ND ++ M
Sbjct: 551 LSNLYAASGKHNDARRVKDMM 613
Score = 74.3 bits (181), Expect = 2e-13
Identities = 49/162 (30%), Positives = 82/162 (50%), Gaps = 12/162 (7%)
Frame = +2
Query: 302 VSTAMLSGYAKLGMVKDARFIFDQMIERDLVCWSAMISGYAESDQPQEALKLFDEMLQKR 361
+ST++L YAK G ++ A+ +FD M RD+VCW+AMISG A + ALKLF +M +
Sbjct: 20 LSTSLLDMYAKCGNLELAKRLFDSMNMRDVVCWNAMISGMAMHGDGKGALKLFYDMEKVG 199
Query: 362 SVPDQITMLSVISACSHVG-------ALAQANWIHTYVDRSGFGRALSVNNALIDMYAKC 414
PD IT ++V +ACS+ G L + ++ V +S L+D+ ++
Sbjct: 200 VKPDDITFIAVFTACSYSGMAYEGLMLLDKMCSVYNIVPKS------EHYGCLVDLLSRA 361
Query: 415 GNLVKAREVFENMPR-----KNVISWSSMINAFAMHGNADSA 451
G +A + + + ++W + ++A HG A
Sbjct: 362 GLFEEAMVMIRKITNSWNGSEETLAWRAFLSACCNHGETQLA 487
Score = 68.2 bits (165), Expect = 1e-11
Identities = 54/198 (27%), Positives = 89/198 (44%), Gaps = 10/198 (5%)
Frame = +2
Query: 170 IQTGLIAMYASCRRIMDARLLFDKMCHPDAVAWNMIIDGYCQNGHYDDALRLFEDMRSSD 229
+ T L+ MYA C + A+ LFD M D V WN +I G +G AL+LF DM
Sbjct: 20 LSTSLLDMYAKCGNLELAKRLFDSMNMRDVVCWNAMISGMAMHGDGKGALKLFYDMEKVG 199
Query: 230 MKPDSVILCTVLSACGHAGNLSYG-RTIHEFVKDNGYAIDSHLQTALINMYANCG----A 284
+KPD + V +AC ++G G + + S L+++ + G A
Sbjct: 200 VKPDDITFIAVFTACSYSGMAYEGLMLLDKMCSVYNIVPKSEHYGCLVDLLSRAGLFEEA 379
Query: 285 MDLARKIYDGLS-SKHLIVSTAMLSGYAKLGMVKDARFIFDQMIERDLVCWSA----MIS 339
M + RKI + + S+ + A LS G + A +++++ D S + +
Sbjct: 380 MVMIRKITNSWNGSEETLAWRAFLSACCNHGETQLAELAAEKVLQLDNHIHSGVYVLLSN 559
Query: 340 GYAESDQPQEALKLFDEM 357
YA S + +A ++ D M
Sbjct: 560 LYAASGKHNDARRVKDMM 613
>AW776322 weakly similar to PIR|H84859|H84 hypothetical protein At2g42920
[imported] - Arabidopsis thaliana, partial (27%)
Length = 633
Score = 149 bits (377), Expect = 3e-36
Identities = 75/209 (35%), Positives = 121/209 (57%), Gaps = 1/209 (0%)
Frame = +1
Query: 425 ENMPRKNVISWSSMINAFAMHGNADSAIKLFRRMKEVNI-EPNGVTFIGVLYACGHAGLV 483
E PR+ + W+S+I AM+G+ A + F +++ + +P+ V+FIGVL AC H G +
Sbjct: 1 ETCPRRGLSCWNSIIIGLAMNGHEREAFEFFSKLESSKLLKPDSVSFIGVLTACKHLGAI 180
Query: 484 EEGEKLFSSMINEHGISPTREHYGCMVDLYCRANFLRKAIELIETMPFAPNVIIWGSLMS 543
+ F M+N++ I P+ +HY C+VD+ +A L +A ELI+ MP P+ IIWGSL+S
Sbjct: 181 NKARDYFELMMNKYEIEPSIKHYTCIVDVLGQAGLLEEAEELIKGMPLKPDAIIWGSLLS 360
Query: 544 ACQVHGEAELGEFAAKRLLELEPDHDGALVVLSNIYAKEKRWNDVGLIRKSMSYKGISKE 603
+C+ H ++ AA+R+ EL P V++SN++A ++ + R M KE
Sbjct: 361 SCRKHRNVQIARRAAQRVYELNPSDASGYVLMSNVHAASNKFEEAIEQRLLMKENLTEKE 540
Query: 604 KASSRIEINNQVHMFMMADRYHKQSDEIY 632
S IE+ +VH F+ R H ++ EIY
Sbjct: 541 PGCSSIELYGEVHEFIAGGRLHPKTQEIY 627
Score = 69.3 bits (168), Expect = 5e-12
Identities = 36/137 (26%), Positives = 77/137 (55%), Gaps = 3/137 (2%)
Frame = +1
Query: 329 RDLVCWSAMISGYAESDQPQEALKLFDEMLQKRSV-PDQITMLSVISACSHVGALAQA-N 386
R L CW+++I G A + +EA + F ++ + + PD ++ + V++AC H+GA+ +A +
Sbjct: 16 RGLSCWNSIIIGLAMNGHEREAFEFFSKLESSKLLKPDSVSFIGVLTACKHLGAINKARD 195
Query: 387 WIHTYVDRSGFGRALSVNNALIDMYAKCGNLVKAREVFENMPRK-NVISWSSMINAFAMH 445
+ +++ ++ ++D+ + G L +A E+ + MP K + I W S++++ H
Sbjct: 196 YFELMMNKYEIEPSIKHYTCIVDVLGQAGLLEEAEELIKGMPLKPDAIIWGSLLSSCRKH 375
Query: 446 GNADSAIKLFRRMKEVN 462
N A + +R+ E+N
Sbjct: 376 RNVQIARRAAQRVYELN 426
Score = 63.5 bits (153), Expect = 3e-10
Identities = 48/164 (29%), Positives = 79/164 (47%), Gaps = 4/164 (2%)
Frame = +1
Query: 202 WNMIIDGYCQNGHYDDALRLFEDMRSSD-MKPDSVILCTVLSACGHAGNLSYGRTIHEFV 260
WN II G NGH +A F + SS +KPDSV VL+AC H G ++ R E +
Sbjct: 31 WNSIIIGLAMNGHEREAFEFFSKLESSKLLKPDSVSFIGVLTACKHLGAINKARDYFELM 210
Query: 261 KDNGYAIDSHLQ--TALINMYANCGAMDLARKIYDGLSSK-HLIVSTAMLSGYAKLGMVK 317
N Y I+ ++ T ++++ G ++ A ++ G+ K I+ ++LS K V+
Sbjct: 211 M-NKYEIEPSIKHYTCIVDVLGQAGLLEEAEELIKGMPLKPDAIIWGSLLSSCRKHRNVQ 387
Query: 318 DARFIFDQMIERDLVCWSAMISGYAESDQPQEALKLFDEMLQKR 361
AR ++ E + + SGY A F+E +++R
Sbjct: 388 IARRAAQRVYELN----PSDASGYVLMSNVHAASNKFEEAIEQR 507
>BF633384 similar to PIR|T46179|T46 hypothetical protein T8H10.30 -
Arabidopsis thaliana, partial (22%)
Length = 595
Score = 148 bits (373), Expect = 8e-36
Identities = 76/176 (43%), Positives = 107/176 (60%), Gaps = 6/176 (3%)
Frame = +1
Query: 433 ISWSSMINAFAMHGNADSAIKLFRRMKEVN-----IEPNGVTFIGVLYACGHAGLVEEGE 487
I+W+ +I A+ MHG + A+KLFRRM E I PN VT+I + + H+G+V+EG
Sbjct: 1 ITWNVLIMAYGMHGKGEEALKLFRRMVEEGDNNREIRPNEVTYIAIFASLSHSGMVDEGL 180
Query: 488 KLFSSMINEHGISPTREHYGCMVDLYCRANFLRKAIELIETMPF-APNVIIWGSLMSACQ 546
LF +M +HGI PT +HY C+VDL R+ + +A LI+TMP V W SL+ AC+
Sbjct: 181 NLFYTMKAKHGIEPTSDHYACLVDLLGRSGQIEEAYNLIKTMPSNMKKVDAWSSLLGACK 360
Query: 547 VHGEAELGEFAAKRLLELEPDHDGALVVLSNIYAKEKRWNDVGLIRKSMSYKGISK 602
+H E+GE AAK L L+P+ V+LSNIY+ W+ +RK M KG+ K
Sbjct: 361 IHQNLEIGEIAAKNLFVLDPNVASYYVLLSNIYSSAGLWDQAIDVRKKMKEKGVRK 528
Score = 57.0 bits (136), Expect = 3e-08
Identities = 33/126 (26%), Positives = 66/126 (52%), Gaps = 8/126 (6%)
Frame = +1
Query: 332 VCWSAMISGYAESDQPQEALKLFDEMLQK----RSV-PDQITMLSVISACSHVGALAQA- 385
+ W+ +I Y + +EALKLF M+++ R + P+++T +++ ++ SH G + +
Sbjct: 1 ITWNVLIMAYGMHGKGEEALKLFRRMVEEGDNNREIRPNEVTYIAIFASLSHSGMVDEGL 180
Query: 386 NWIHTYVDRSGFGRALSVNNALIDMYAKCGNLVKAREVFENMP--RKNVISWSSMINAFA 443
N +T + G L+D+ + G + +A + + MP K V +WSS++ A
Sbjct: 181 NLFYTMKAKHGIEPTSDHYACLVDLLGRSGQIEEAYNLIKTMPSNMKKVDAWSSLLGACK 360
Query: 444 MHGNAD 449
+H N +
Sbjct: 361 IHQNLE 378
>AL370459 weakly similar to GP|20146256|d hypothetical protein {Oryza sativa
(japonica cultivar-group)}, partial (11%)
Length = 477
Score = 139 bits (351), Expect = 3e-33
Identities = 66/125 (52%), Positives = 85/125 (67%)
Frame = +1
Query: 613 NQVHMFMMADRYHKQSDEIYEKLDEVVSKLKLVGYKPSTSGILIDLEEEDKKELVLWHSE 672
NQVH F + D H ++ +IYEKLDE+ K+K VGY P+T +L D+E+E K++ + HSE
Sbjct: 4 NQVHKFHVGDTLHPKAQQIYEKLDELALKIKNVGYVPNTDFVLHDVEDEQKEQYLFQHSE 183
Query: 673 KLAVCYGLISRRNESCIRIVKNLRICEDCHSFMKLVSKVYQIEIVVRDRTRFHHCSGGIC 732
KLAV + LIS N IR+ KNLR+C DCH+ +K +S V EIVVRD RFHH G C
Sbjct: 184 KLAVAFALISTPNPKPIRVFKNLRVCGDCHTAIKYISMVSGREIVVRDANRFHHMKDGTC 363
Query: 733 SCRDY 737
SC DY
Sbjct: 364 SCNDY 378
>CA921534 similar to PIR|D96516|D96 F16N3.14 [imported] - Arabidopsis
thaliana, partial (18%)
Length = 790
Score = 136 bits (343), Expect = 3e-32
Identities = 80/234 (34%), Positives = 128/234 (54%)
Frame = -1
Query: 504 EHYGCMVDLYCRANFLRKAIELIETMPFAPNVIIWGSLMSACQVHGEAELGEFAAKRLLE 563
E Y +V+++ A L KA E + MP P V +W +L + ++HG+ E + A + L
Sbjct: 784 EPYLGVVNIFGCAVGLMKAHEFMRNMPIEPGVELWETLRNFARIHGDLEREDCADELLTV 605
Query: 564 LEPDHDGALVVLSNIYAKEKRWNDVGLIRKSMSYKGISKEKASSRIEINNQVHMFMMADR 623
L+P A + V L ++ K+ A + +E N+V +
Sbjct: 604 LDPSKAAA--------------DKVPLPQRK-------KQSAINMLEEKNRVSEYRCNMP 488
Query: 624 YHKQSDEIYEKLDEVVSKLKLVGYKPSTSGILIDLEEEDKKELVLWHSEKLAVCYGLISR 683
Y ++ D KL + +++ GY P T +L D++EE+K++ + +HSE+LA+ YGLIS
Sbjct: 487 YKEEGDV---KLRGLTGQMREAGYVPDTRYVLHDIDEEEKEKALQYHSERLAIAYGLIST 317
Query: 684 RNESCIRIVKNLRICEDCHSFMKLVSKVYQIEIVVRDRTRFHHCSGGICSCRDY 737
+ +RI+KNLRIC DCH+ +K++SK+ E++VRD RFHH G CSC DY
Sbjct: 316 PPRTTLRIIKNLRICGDCHNAIKIMSKIVGRELIVRDNKRFHHFKDGKCSCGDY 155
>BE202961 weakly similar to GP|6016735|gb|A hypothetical protein {Arabidopsis
thaliana}, partial (14%)
Length = 613
Score = 127 bits (318), Expect = 2e-29
Identities = 70/235 (29%), Positives = 122/235 (51%)
Frame = +2
Query: 219 LRLFEDMRSSDMKPDSVILCTVLSACGHAGNLSYGRTIHEFVKDNGYAIDSHLQTALINM 278
L L +M +KP+ V + +VL AC NL G IHE + G+ +++ + TAL++M
Sbjct: 2 LDLSSEMLDKRIKPNWVTVVSVLRACACISNLEEGMKIHELAVNYGFEMETTVSTALMDM 181
Query: 279 YANCGAMDLARKIYDGLSSKHLIVSTAMLSGYAKLGMVKDARFIFDQMIERDLVCWSAMI 338
Y K + A IF++M ++D++ W+ +
Sbjct: 182 -------------------------------YMKCFSPEKAVDIFNRMPKKDVIAWAVLF 268
Query: 339 SGYAESDQPQEALKLFDEMLQKRSVPDQITMLSVISACSHVGALAQANWIHTYVDRSGFG 398
SGYA++ E++ +F ML + PD I ++ +++ S +G L QA H +V ++GF
Sbjct: 269 SGYADNGMVHESMWVFRNMLSSGTRPDAIALVKILTTVSELGILQQAVCFHAFVIKNGFE 448
Query: 399 RALSVNNALIDMYAKCGNLVKAREVFENMPRKNVISWSSMINAFAMHGNADSAIK 453
+ +LI++YAKC ++ A +VF+ M K+V++WSS+I A+ HG + A+K
Sbjct: 449 NNQFIGASLIEVYAKCSSIEDANKVFKGMTYKDVVTWSSIIAAYGFHGQGEEALK 613
Score = 107 bits (266), Expect = 2e-23
Identities = 53/182 (29%), Positives = 99/182 (54%)
Frame = +2
Query: 138 SLLKAVSKVSAFNHGLEIHGLASKLGFVDDPFIQTGLIAMYASCRRIMDARLLFDKMCHP 197
S+L+A + +S G++IH LA GF + + T L+ MY C A +F++M
Sbjct: 62 SVLRACACISNLEEGMKIHELAVNYGFEMETTVSTALMDMYMKCFSPEKAVDIFNRMPKK 241
Query: 198 DAVAWNMIIDGYCQNGHYDDALRLFEDMRSSDMKPDSVILCTVLSACGHAGNLSYGRTIH 257
D +AW ++ GY NG +++ +F +M SS +PD++ L +L+ G L H
Sbjct: 242 DVIAWAVLFSGYADNGMVHESMWVFRNMLSSGTRPDAIALVKILTTVSELGILQQAVCFH 421
Query: 258 EFVKDNGYAIDSHLQTALINMYANCGAMDLARKIYDGLSSKHLIVSTAMLSGYAKLGMVK 317
FV NG+ + + +LI +YA C +++ A K++ G++ K ++ +++++ Y G +
Sbjct: 422 AFVIKNGFENNQFIGASLIEVYAKCSSIEDANKVFKGMTYKDVVTWSSIIAAYGFHGQGE 601
Query: 318 DA 319
+A
Sbjct: 602 EA 607
Score = 107 bits (266), Expect = 2e-23
Identities = 55/202 (27%), Positives = 110/202 (54%)
Frame = +2
Query: 351 LKLFDEMLQKRSVPDQITMLSVISACSHVGALAQANWIHTYVDRSGFGRALSVNNALIDM 410
L L EML KR P+ +T++SV+ AC+ + L + IH GF +V+ AL+DM
Sbjct: 2 LDLSSEMLDKRIKPNWVTVVSVLRACACISNLEEGMKIHELAVNYGFEMETTVSTALMDM 181
Query: 411 YAKCGNLVKAREVFENMPRKNVISWSSMINAFAMHGNADSAIKLFRRMKEVNIEPNGVTF 470
Y KC + KA ++F MP+K+VI+W+ + + +A +G ++ +FR M P+ +
Sbjct: 182 YMKCFSPEKAVDIFNRMPKKDVIAWAVLFSGYADNGMVHESMWVFRNMLSSGTRPDAIAL 361
Query: 471 IGVLYACGHAGLVEEGEKLFSSMINEHGISPTREHYGCMVDLYCRANFLRKAIELIETMP 530
+ +L G++++ F + + ++G + ++++Y + + + A ++ + M
Sbjct: 362 VKILTTVSELGILQQA-VCFHAFVIKNGFENNQFIGASLIEVYAKCSSIEDANKVFKGMT 538
Query: 531 FAPNVIIWGSLMSACQVHGEAE 552
+ +V+ W S+++A HG+ E
Sbjct: 539 Y-KDVVTWSSIIAAYGFHGQGE 601
Score = 58.2 bits (139), Expect = 1e-08
Identities = 34/136 (25%), Positives = 65/136 (47%)
Frame = +2
Query: 85 ALSVFSQIPNPHTHFSNQLLRHLSRSSFPEKTIFLYHNLRAINAFALDRFSFPSLLKAVS 144
A+ +F+++P L + + ++++++ N+ + + D + +L VS
Sbjct: 209 AVDIFNRMPKKDVIAWAVLFSGYADNGMVHESMWVFRNMLS-SGTRPDAIALVKILTTVS 385
Query: 145 KVSAFNHGLEIHGLASKLGFVDDPFIQTGLIAMYASCRRIMDARLLFDKMCHPDAVAWNM 204
++ + H K GF ++ FI LI +YA C I DA +F M + D V W+
Sbjct: 386 ELGILQQAVCFHAFVIKNGFENNQFIGASLIEVYAKCSSIEDANKVFKGMTYKDVVTWSS 565
Query: 205 IIDGYCQNGHYDDALR 220
II Y +G ++AL+
Sbjct: 566 IIAAYGFHGQGEEALK 613
>AW774626 similar to PIR|T47909|T47 hypothetical protein T20K12.70 -
Arabidopsis thaliana, partial (12%)
Length = 703
Score = 126 bits (316), Expect = 3e-29
Identities = 78/266 (29%), Positives = 134/266 (50%), Gaps = 2/266 (0%)
Frame = +3
Query: 174 LIAMYASCRRIMDARLLFD--KMCHPDAVAWNMIIDGYCQNGHYDDALRLFEDMRSSDMK 231
L+ MYA C+ + +A LF + + V W ++ GY QNG A+ F M + ++
Sbjct: 3 LVDMYAKCKCVSEAEFLFKGLEFDRKNHVLWTAMVTGYAQNGDGYKAVEFFRYMHAQGVE 182
Query: 232 PDSVILCTVLSACGHAGNLSYGRTIHEFVKDNGYAIDSHLQTALINMYANCGAMDLARKI 291
+ T+L+AC +G +H F+ +G+ + ++Q+AL++MYA
Sbjct: 183 CNQYTFPTILTACSSVLARCFGEQVHGFIVKSGFGSNVYVQSALVDMYA----------- 329
Query: 292 YDGLSSKHLIVSTAMLSGYAKLGMVKDARFIFDQMIERDLVCWSAMISGYAESDQPQEAL 351
K G +K+A+ + + M + D+V W++++ G+ +EAL
Sbjct: 330 --------------------KCGDLKNAKNMLETMEDDDVVSWNSLMVGFVRHGLEEEAL 449
Query: 352 KLFDEMLQKRSVPDQITMLSVISACSHVGALAQANWIHTYVDRSGFGRALSVNNALIDMY 411
+LF M + D T SV++ C VG++ + +H + ++GF V+NAL+DMY
Sbjct: 450 RLFKNMHGRNMKIDDYTFPSVLNCCV-VGSINPKS-VHGLIIKTGFENYKLVSNALVDMY 623
Query: 412 AKCGNLVKAREVFENMPRKNVISWSS 437
AK G++ A VFE M K+VISW+S
Sbjct: 624 AKTGDMDCAYTVFEKMLEKDVISWTS 701
Score = 122 bits (306), Expect = 5e-28
Identities = 73/233 (31%), Positives = 124/233 (52%), Gaps = 2/233 (0%)
Frame = +3
Query: 310 YAKLGMVKDARFIFD--QMIERDLVCWSAMISGYAESDQPQEALKLFDEMLQKRSVPDQI 367
YAK V +A F+F + ++ V W+AM++GYA++ +A++ F M + +Q
Sbjct: 15 YAKCKCVSEAEFLFKGLEFDRKNHVLWTAMVTGYAQNGDGYKAVEFFRYMHAQGVECNQY 194
Query: 368 TMLSVISACSHVGALAQANWIHTYVDRSGFGRALSVNNALIDMYAKCGNLVKAREVFENM 427
T ++++ACS V A +H ++ +SGFG + V +AL+DMYAKCG+L A+ + E M
Sbjct: 195 TFPTILTACSSVLARCFGEQVHGFIVKSGFGSNVYVQSALVDMYAKCGDLKNAKNMLETM 374
Query: 428 PRKNVISWSSMINAFAMHGNADSAIKLFRRMKEVNIEPNGVTFIGVLYACGHAGLVEEGE 487
+V+SW+S++ F HG + A++LF+ M N++ + TF VL C +
Sbjct: 375 EDDDVVSWNSLMVGFVRHGLEEEALRLFKNMHGRNMKIDDYTFPSVLNCCVVGSI---NP 545
Query: 488 KLFSSMINEHGISPTREHYGCMVDLYCRANFLRKAIELIETMPFAPNVIIWGS 540
K +I + G + +VD+Y + + A + E M +VI W S
Sbjct: 546 KSVHGLIIKTGFENYKLVSNALVDMYAKTGDMDCAYTVFEKM-LEKDVISWTS 701
Score = 102 bits (255), Expect = 4e-22
Identities = 54/174 (31%), Positives = 102/174 (58%)
Frame = +3
Query: 132 DRFSFPSLLKAVSKVSAFNHGLEIHGLASKLGFVDDPFIQTGLIAMYASCRRIMDARLLF 191
++++FP++L A S V A G ++HG K GF + ++Q+ L+ MYA C + +A+ +
Sbjct: 186 NQYTFPTILTACSSVLARCFGEQVHGFIVKSGFGSNVYVQSALVDMYAKCGDLKNAKNML 365
Query: 192 DKMCHPDAVAWNMIIDGYCQNGHYDDALRLFEDMRSSDMKPDSVILCTVLSACGHAGNLS 251
+ M D V+WN ++ G+ ++G ++ALRLF++M +MK D +VL+ C G+++
Sbjct: 366 ETMEDDDVVSWNSLMVGFVRHGLEEEALRLFKNMHGRNMKIDDYTFPSVLNCC-VVGSIN 542
Query: 252 YGRTIHEFVKDNGYAIDSHLQTALINMYANCGAMDLARKIYDGLSSKHLIVSTA 305
+++H + G+ + AL++MYA G MD A +++ + K +I T+
Sbjct: 543 -PKSVHGLIIKTGFENYKLVSNALVDMYAKTGDMDCAYTVFEKMLEKDVISWTS 701
Score = 73.9 bits (180), Expect = 2e-13
Identities = 46/148 (31%), Positives = 76/148 (51%), Gaps = 2/148 (1%)
Frame = +3
Query: 407 LIDMYAKCGNLVKAREVFENM--PRKNVISWSSMINAFAMHGNADSAIKLFRRMKEVNIE 464
L+DMYAKC + +A +F+ + RKN + W++M+ +A +G+ A++ FR M +E
Sbjct: 3 LVDMYAKCKCVSEAEFLFKGLEFDRKNHVLWTAMVTGYAQNGDGYKAVEFFRYMHAQGVE 182
Query: 465 PNGVTFIGVLYACGHAGLVEEGEKLFSSMINEHGISPTREHYGCMVDLYCRANFLRKAIE 524
N TF +L AC GE++ ++ + G +VD+Y + L+ A
Sbjct: 183 CNQYTFPTILTACSSVLARCFGEQVHGFIV-KSGFGSNVYVQSALVDMYAKCGDLKNAKN 359
Query: 525 LIETMPFAPNVIIWGSLMSACQVHGEAE 552
++ETM +V+ W SLM HG E
Sbjct: 360 MLETME-DDDVVSWNSLMVGFVRHGLEE 440
>BG645370 weakly similar to PIR|T00405|T004 hypothetical protein At2g44880
[imported] - Arabidopsis thaliana, partial (11%)
Length = 784
Score = 124 bits (311), Expect = 1e-28
Identities = 55/129 (42%), Positives = 85/129 (65%)
Frame = -2
Query: 470 FIGVLYACGHAGLVEEGEKLFSSMINEHGISPTREHYGCMVDLYCRANFLRKAIELIETM 529
F VL AC HAG +++GE+ F+ MI +G+SP + Y C++DLY R LRKA +L+E M
Sbjct: 783 FTAVLTACNHAGFIDKGEEYFNKMITNYGLSPDIDIYACLIDLYARNGNLRKARDLMEEM 604
Query: 530 PFAPNVIIWGSLMSACQVHGEAELGEFAAKRLLELEPDHDGALVVLSNIYAKEKRWNDVG 589
P+ PN IIW S +SAC+++G+ ELG AA +L+++EP + + L++IY + WN+
Sbjct: 603 PYDPNCIIWSSFLSACKIYGDVELGREAAIQLIKMEPCNAAPYLTLAHIYTTKGLWNEAS 424
Query: 590 LIRKSMSYK 598
+R M +
Sbjct: 423 EVRSLMQQR 397
Score = 55.8 bits (133), Expect = 6e-08
Identities = 38/130 (29%), Positives = 69/130 (52%), Gaps = 7/130 (5%)
Frame = -2
Query: 371 SVISACSHVGALAQAN-WIHTYVDRSGFGRALSVNNALIDMYAKCGNLVKAREVFENMP- 428
+V++AC+H G + + + + + G + + LID+YA+ GNL KAR++ E MP
Sbjct: 777 AVLTACNHAGFIDKGEEYFNKMITNYGLSPDIDIYACLIDLYARNGNLRKARDLMEEMPY 598
Query: 429 RKNVISWSSMINAFAMHGNA----DSAIKLFRRMKEVNIEP-NGVTFIGVLYACGHAGLV 483
N I WSS ++A ++G+ ++AI+L + +EP N ++ + + GL
Sbjct: 597 DPNCIIWSSFLSACKIYGDVELGREAAIQL------IKMEPCNAAPYLTLAHIYTTKGLW 436
Query: 484 EEGEKLFSSM 493
E ++ S M
Sbjct: 435 NEASEVRSLM 406
Score = 34.7 bits (78), Expect = 0.14
Identities = 25/93 (26%), Positives = 47/93 (49%), Gaps = 2/93 (2%)
Frame = -2
Query: 240 VLSACGHAGNLSYGRT-IHEFVKDNGYAIDSHLQTALINMYANCGAMDLARKIYDGLS-S 297
VL+AC HAG + G ++ + + G + D + LI++YA G + AR + + +
Sbjct: 774 VLTACNHAGFIDKGEEYFNKMITNYGLSPDIDIYACLIDLYARNGNLRKARDLMEEMPYD 595
Query: 298 KHLIVSTAMLSGYAKLGMVKDARFIFDQMIERD 330
+ I+ ++ LS G V+ R Q+I+ +
Sbjct: 594 PNCIIWSSFLSACKIYGDVELGREAAIQLIKME 496
Score = 28.5 bits (62), Expect = 9.7
Identities = 17/65 (26%), Positives = 36/65 (55%), Gaps = 3/65 (4%)
Frame = -3
Query: 577 NIYAKEKRWNDVGLIRKSMSYKGISKEK---ASSRIEINNQVHMFMMADRYHKQSDEIYE 633
+I ++K + L +++ KG+ + A S +E++ H+F + D H+QS+EIY
Sbjct: 464 HISIQQKDYGTKLLKLEALCNKGLKENLQGGAWSWVEVDKHNHVFAVDDVTHQQSNEIYA 285
Query: 634 KLDEV 638
+ + +
Sbjct: 284 E*ESI 270
>BF640739 weakly similar to GP|7363286|dbj Similar to Arabidopsis thaliana
DNA chromosome 4 BAC clone F4I10; vegetative storage
protein., partial (14%)
Length = 444
Score = 120 bits (300), Expect = 2e-27
Identities = 61/146 (41%), Positives = 89/146 (60%), Gaps = 1/146 (0%)
Frame = +1
Query: 349 EALKLFDEMLQKRSVPDQITMLSVISACSHVGALAQANWIHTYVDRSGFGRALSVNNALI 408
EAL LF M + PD T +SVI+A + + QA WIH R+ + V AL+
Sbjct: 7 EALNLFCTMQSQGIKPDSFTFVSVITALADLSVTRQAKWIHGLAIRTNMDTNVFVATALV 186
Query: 409 DMYAKCGNLVKAREVFENMPRKNVISWSSMINAFAMHGNADSAIKLFRRMK-EVNIEPNG 467
DMYAKCG + ARE+F+ M ++VI+W++MI+ + HG +A+ LF M+ E +++PN
Sbjct: 187 DMYAKCGAIETARELFDMMQERHVITWNAMIDGYGTHGLGKAALDLFDDMQNEASLKPND 366
Query: 468 VTFIGVLYACGHAGLVEEGEKLFSSM 493
+TF+ V+ AC H+G VEEG F M
Sbjct: 367 ITFLSVISACSHSGFVEEGLYYFKIM 444
Score = 89.4 bits (220), Expect = 5e-18
Identities = 47/123 (38%), Positives = 71/123 (57%), Gaps = 1/123 (0%)
Frame = +1
Query: 132 DRFSFPSLLKAVSKVSAFNHGLEIHGLASKLGFVDDPFIQTGLIAMYASCRRIMDARLLF 191
D F+F S++ A++ +S IHGLA + + F+ T L+ MYA C I AR LF
Sbjct: 55 DSFTFVSVITALADLSVTRQAKWIHGLAIRTNMDTNVFVATALVDMYAKCGAIETARELF 234
Query: 192 DKMCHPDAVAWNMIIDGYCQNGHYDDALRLFEDMRS-SDMKPDSVILCTVLSACGHAGNL 250
D M + WN +IDGY +G AL LF+DM++ + +KP+ + +V+SAC H+G +
Sbjct: 235 DMMQERHVITWNAMIDGYGTHGLGKAALDLFDDMQNEASLKPNDITFLSVISACSHSGFV 414
Query: 251 SYG 253
G
Sbjct: 415 EEG 423
Score = 88.6 bits (218), Expect = 8e-18
Identities = 58/166 (34%), Positives = 85/166 (50%), Gaps = 1/166 (0%)
Frame = +1
Query: 216 DDALRLFEDMRSSDMKPDSVILCTVLSACGHAGNLSYGRTIHEFVKDNGYAIDSHLQTAL 275
++AL LF M+S +KPDS +V++A + IH G AI +++ T
Sbjct: 4 NEALNLFCTMQSQGIKPDSFTFVSVITALADLSVTRQAKWIH------GLAIRTNMDT-- 159
Query: 276 INMYANCGAMDLARKIYDGLSSKHLIVSTAMLSGYAKLGMVKDARFIFDQMIERDLVCWS 335
++ V+TA++ YAK G ++ AR +FD M ER ++ W+
Sbjct: 160 -----------------------NVFVATALVDMYAKCGAIETARELFDMMQERHVITWN 270
Query: 336 AMISGYAESDQPQEALKLFDEMLQKRSV-PDQITMLSVISACSHVG 380
AMI GY + AL LFD+M + S+ P+ IT LSVISACSH G
Sbjct: 271 AMIDGYGTHGLGKAALDLFDDMQNEASLKPNDITFLSVISACSHSG 408
>BF642497 weakly similar to PIR|H96713|H96 hypothetical protein T6L1.11
[imported] - Arabidopsis thaliana, partial (20%)
Length = 461
Score = 110 bits (275), Expect = 2e-24
Identities = 51/148 (34%), Positives = 90/148 (60%)
Frame = +3
Query: 332 VCWSAMISGYAESDQPQEALKLFDEMLQKRSVPDQITMLSVISACSHVGALAQANWIHTY 391
+ W++MI+G+ ++ ++A+ +F EM + DQ T SV++AC V AL + +H Y
Sbjct: 6 ISWTSMITGFTQNGLDRDAIDIFREMKLENLQMDQYTFGSVLTACGGVMALQEGKQVHAY 185
Query: 392 VDRSGFGRALSVNNALIDMYAKCGNLVKAREVFENMPRKNVISWSSMINAFAMHGNADSA 451
+ R+ + + V +AL+DMY KC N+ A VF+ M KNV+SW++M+ + +G ++ A
Sbjct: 186 IIRTDYKDNIFVASALVDMYCKCKNIKSAEAVFKKMTCKNVVSWTAMLVGYGQNGYSEEA 365
Query: 452 IKLFRRMKEVNIEPNGVTFIGVLYACGH 479
+K F M++ IEP+ T V+ +C +
Sbjct: 366 VKTFSDMQKYGIEPDDFTLGSVISSCAN 449
Score = 87.0 bits (214), Expect = 2e-17
Identities = 40/148 (27%), Positives = 81/148 (54%)
Frame = +3
Query: 103 LLRHLSRSSFPEKTIFLYHNLRAINAFALDRFSFPSLLKAVSKVSAFNHGLEIHGLASKL 162
++ +++ I ++ ++ N +D+++F S+L A V A G ++H +
Sbjct: 21 MITGFTQNGLDRDAIDIFREMKLEN-LQMDQYTFGSVLTACGGVMALQEGKQVHAYIIRT 197
Query: 163 GFVDDPFIQTGLIAMYASCRRIMDARLLFDKMCHPDAVAWNMIIDGYCQNGHYDDALRLF 222
+ D+ F+ + L+ MY C+ I A +F KM + V+W ++ GY QNG+ ++A++ F
Sbjct: 198 DYKDNIFVASALVDMYCKCKNIKSAEAVFKKMTCKNVVSWTAMLVGYGQNGYSEEAVKTF 377
Query: 223 EDMRSSDMKPDSVILCTVLSACGHAGNL 250
DM+ ++PD L +V+S+C + +L
Sbjct: 378 SDMQKYGIEPDDFTLGSVISSCANLASL 461
Score = 86.3 bits (212), Expect = 4e-17
Identities = 50/184 (27%), Positives = 93/184 (50%)
Frame = +3
Query: 199 AVAWNMIIDGYCQNGHYDDALRLFEDMRSSDMKPDSVILCTVLSACGHAGNLSYGRTIHE 258
+++W +I G+ QNG DA+ +F +M+ +++ D +VL+ACG L G+ +H
Sbjct: 3 SISWTSMITGFTQNGLDRDAIDIFREMKLENLQMDQYTFGSVLTACGGVMALQEGKQVHA 182
Query: 259 FVKDNGYAIDSHLQTALINMYANCGAMDLARKIYDGLSSKHLIVSTAMLSGYAKLGMVKD 318
++ Y + + +AL++MY C + A ++ ++ K+++ TAML GY +
Sbjct: 183 YIIRTDYKDNIFVASALVDMYCKCKNIKSAEAVFKKMTCKNVVSWTAMLVGYGQ------ 344
Query: 319 ARFIFDQMIERDLVCWSAMISGYAESDQPQEALKLFDEMLQKRSVPDQITMLSVISACSH 378
+GY+E EA+K F +M + PD T+ SVIS+C++
Sbjct: 345 --------------------NGYSE-----EAVKTFSDMQKYGIEPDDFTLGSVISSCAN 449
Query: 379 VGAL 382
+ +L
Sbjct: 450 LASL 461
Score = 61.2 bits (147), Expect = 1e-09
Identities = 40/137 (29%), Positives = 70/137 (50%), Gaps = 2/137 (1%)
Frame = +3
Query: 433 ISWSSMINAFAMHGNADSAIKLFRRMKEVNIEPNGVTFIGVLYACGHAGLVEEGEKLFSS 492
ISW+SMI F +G AI +FR MK N++ + TF VL ACG ++EG+++ +
Sbjct: 6 ISWTSMITGFTQNGLDRDAIDIFREMKLENLQMDQYTFGSVLTACGGVMALQEGKQVHAY 185
Query: 493 MINEHGISPTREHYGCMVDLYCRANFLRKAIELIETMPFAPNVIIWGSLMSACQVHGEAE 552
+I +VD+YC+ ++ A + + M NV+ W +++ +G +E
Sbjct: 186 IIRT-DYKDNIFVASALVDMYCKCKNIKSAEAVFKKMT-CKNVVSWTAMLVGYGQNGYSE 359
Query: 553 --LGEFAAKRLLELEPD 567
+ F+ + +EPD
Sbjct: 360 EAVKTFSDMQKYGIEPD 410
>AW257410 weakly similar to GP|10177533|db selenium-binding protein-like
{Arabidopsis thaliana}, partial (21%)
Length = 724
Score = 110 bits (275), Expect = 2e-24
Identities = 56/199 (28%), Positives = 111/199 (55%), Gaps = 2/199 (1%)
Frame = +1
Query: 298 KHLIVSTAMLSGYAKLGM--VKDARFIFDQMIERDLVCWSAMISGYAESDQPQEALKLFD 355
+H + + +L+ YA + + AR +FD++ + V W+ MI Y+ S+ P+EAL L+
Sbjct: 67 RHKLTVSRLLTTYASMEFSNLTYARMVFDRISSPNTVMWNTMIRAYSNSNDPEEALLLYH 246
Query: 356 EMLQKRSVPDQITMLSVISACSHVGALAQANWIHTYVDRSGFGRALSVNNALIDMYAKCG 415
+ML + T ++ ACS + ALA+ + IH + + GFG + N+L+ +YA G
Sbjct: 247 QMLHHSIPHNAYTFPFLLKACSALSALAETHQIHVQIIKRGFGSEVYATNSLLRVYAISG 426
Query: 416 NLVKAREVFENMPRKNVISWSSMINAFAMHGNADSAIKLFRRMKEVNIEPNGVTFIGVLY 475
++ A +F+ +P ++++SW++MI+ + GN + A K+F+ M E N+ +++ ++
Sbjct: 427 SIKSAHVLFDLLPSRDIVSWNTMIDGYIKCGNVEMAYKIFQAMPEKNV----ISWTSMIV 594
Query: 476 ACGHAGLVEEGEKLFSSMI 494
G+ ++ L M+
Sbjct: 595 GFVRTGMHKKALSLLQQML 651
Score = 101 bits (252), Expect = 9e-22
Identities = 58/208 (27%), Positives = 105/208 (49%), Gaps = 2/208 (0%)
Frame = +1
Query: 161 KLGFVDDPFIQTGLIAMYASCR--RIMDARLLFDKMCHPDAVAWNMIIDGYCQNGHYDDA 218
K G + + L+ YAS + AR++FD++ P+ V WN +I Y + ++A
Sbjct: 52 KKGTIRHKLTVSRLLTTYASMEFSNLTYARMVFDRISSPNTVMWNTMIRAYSNSNDPEEA 231
Query: 219 LRLFEDMRSSDMKPDSVILCTVLSACGHAGNLSYGRTIHEFVKDNGYAIDSHLQTALINM 278
L L+ M + ++ +L AC L+ IH + G+ + + +L+ +
Sbjct: 232 LLLYHQMLHHSIPHNAYTFPFLLKACSALSALAETHQIHVQIIKRGFGSEVYATNSLLRV 411
Query: 279 YANCGAMDLARKIYDGLSSKHLIVSTAMLSGYAKLGMVKDARFIFDQMIERDLVCWSAMI 338
YA G++ A ++D L S+ ++ M+ GY K G V+ A IF M E++++ W++MI
Sbjct: 412 YAISGSIKSAHVLFDLLPSRDIVSWNTMIDGYIKCGNVEMAYKIFQAMPEKNVISWTSMI 591
Query: 339 SGYAESDQPQEALKLFDEMLQKRSVPDQ 366
G+ + ++AL L +ML R PD+
Sbjct: 592 VGFVRTGMHKKALSLLQQMLVARIKPDK 675
Score = 97.8 bits (242), Expect = 1e-20
Identities = 61/187 (32%), Positives = 100/187 (52%)
Frame = +1
Query: 39 TTLSHLKQIHAQILHSNTTPENTNTLLSKLALSICTLSSSSSSLHYALSVFSQIPNPHTH 98
+ + LKQI+ Q L T +S+L + ++ S+ L YA VF +I +P+T
Sbjct: 10 SNIGELKQIYVQFLKKGTIRHKLT--VSRLLTTYASMEFSN--LTYARMVFDRISSPNTV 177
Query: 99 FSNQLLRHLSRSSFPEKTIFLYHNLRAINAFALDRFSFPSLLKAVSKVSAFNHGLEIHGL 158
N ++R S S+ PE+ + LYH + ++ + ++FP LLKA S +SA +IH
Sbjct: 178 MWNTMIRAYSNSNDPEEALLLYHQMLH-HSIPHNAYTFPFLLKACSALSALAETHQIHVQ 354
Query: 159 ASKLGFVDDPFIQTGLIAMYASCRRIMDARLLFDKMCHPDAVAWNMIIDGYCQNGHYDDA 218
K GF + + L+ +YA I A +LFD + D V+WN +IDGY + G+ + A
Sbjct: 355 IIKRGFGSEVYATNSLLRVYAISGSIKSAHVLFDLLPSRDIVSWNTMIDGYIKCGNVEMA 534
Query: 219 LRLFEDM 225
++F+ M
Sbjct: 535 YKIFQAM 555
>BF635667 weakly similar to PIR|C85359|C853 hypothetical protein AT4g30700
[imported] - Arabidopsis thaliana, partial (6%)
Length = 603
Score = 106 bits (264), Expect = 4e-23
Identities = 57/163 (34%), Positives = 88/163 (53%), Gaps = 1/163 (0%)
Frame = +3
Query: 481 GLVEEGEKLFSSMINEHGISPTREHYGCMVDLYCRANFLRKAIELIETMPFAP-NVIIWG 539
GL+++G+K+F SMI+++G+ PT EHY CMVDL+ R L +A++L+E M + +WG
Sbjct: 30 GLLDQGKKIFGSMISDYGLEPTAEHYACMVDLFSRCGRLEEALQLLERMKSSSVTGSMWG 209
Query: 540 SLMSACQVHGEAELGEFAAKRLLELEPDHDGALVVLSNIYAKEKRWNDVGLIRKSMSYKG 599
+L++ C +H ++GE AA L +LEP++ V L IY V I M G
Sbjct: 210 ALLAGCVMHQNVKIGEVAAHHLFQLEPNNTSNYVALWGIYQSRGMVLGVSTIXGKMRDLG 389
Query: 600 ISKEKASSRIEINNQVHMFMMADRYHKQSDEIYEKLDEVVSKL 642
+ K I I + H F D H S I +++ E+ L
Sbjct: 390 LVKTPGCXWINIAGRAHKFYQGDLSHPLSHIICKRVYEIXXTL 518
>TC84835 weakly similar to PIR|T02328|T02328 hypothetical protein At2g34400
[imported] - Arabidopsis thaliana, partial (5%)
Length = 647
Score = 103 bits (258), Expect = 2e-22
Identities = 55/185 (29%), Positives = 103/185 (54%), Gaps = 1/185 (0%)
Frame = +1
Query: 77 SSSSSLHYALSVFSQIPNPHTHFSNQLLRHLSRSSFPEKTIFLYHNLRAINAFALDRFSF 136
++++ HY+LS+F+ +P N L++ + + + I L++ LR +N D +++
Sbjct: 40 NNNNYFHYSLSIFNHTLHPSLFLYNLLIKSFFKRNSFQTLISLFNQLR-LNGLYPDNYTY 216
Query: 137 PSLLKAVSKVSAFNHGLEIHGLASKLGFVDDPFIQTGLIAMYASCRRIMDARLLFDKMCH 196
P +LKAV+ ++ F G +IH K G D ++ + MYA RI R LFD++
Sbjct: 217 PFVLKAVAFIADFRQGTKIHAFVFKTGLDSDYYVSNSFMDMYAELGRIDFVRKLFDEISE 396
Query: 197 PDAVAWNMIIDGYCQNGHYDDALRLFEDMR-SSDMKPDSVILCTVLSACGHAGNLSYGRT 255
D+V+WN++I G + +++A+ +F+ MR S+ K + + L+AC + N+ G+
Sbjct: 397 RDSVSWNVMISGCVKCRRFEEAVEVFQRMRVDSNEKISEATVVSSLTACAASRNVEVGKE 576
Query: 256 IHEFV 260
IH F+
Sbjct: 577 IHGFI 591
Score = 78.2 bits (191), Expect = 1e-14
Identities = 43/176 (24%), Positives = 91/176 (51%), Gaps = 1/176 (0%)
Frame = +1
Query: 322 IFDQMIERDLVCWSAMISGYAESDQPQEALKLFDEMLQKRSVPDQITMLSVISACSHVGA 381
IF+ + L ++ +I + + + Q + LF+++ PD T V+ A + +
Sbjct: 73 IFNHTLHPSLFLYNLLIKSFFKRNSFQTLISLFNQLRLNGLYPDNYTYPFVLKAVAFIAD 252
Query: 382 LAQANWIHTYVDRSGFGRALSVNNALIDMYAKCGNLVKAREVFENMPRKNVISWSSMINA 441
Q IH +V ++G V+N+ +DMYA+ G + R++F+ + ++ +SW+ MI+
Sbjct: 253 FRQGTKIHAFVFKTGLDSDYYVSNSFMDMYAELGRIDFVRKLFDEISERDSVSWNVMISG 432
Query: 442 FAMHGNADSAIKLFRRMK-EVNIEPNGVTFIGVLYACGHAGLVEEGEKLFSSMINE 496
+ A+++F+RM+ + N + + T + L AC + VE G+++ +I +
Sbjct: 433 CVKCRRFEEAVEVFQRMRVDSNEKISEATVVSSLTACAASRNVEVGKEIHGFIIEK 600
Score = 72.8 bits (177), Expect = 4e-13
Identities = 47/204 (23%), Positives = 88/204 (43%), Gaps = 1/204 (0%)
Frame = +1
Query: 190 LFDKMCHPDAVAWNMIIDGYCQNGHYDDALRLFEDMRSSDMKPDSVILCTVLSACGHAGN 249
+F+ HP +N++I + + + + LF +R + + PD+ VL A +
Sbjct: 73 IFNHTLHPSLFLYNLLIKSFFKRNSFQTLISLFNQLRLNGLYPDNYTYPFVLKAVAFIAD 252
Query: 250 LSYGRTIHEFVKDNGYAIDSHLQTALINMYANCGAMDLARKIYDGLSSKHLIVSTAMLSG 309
G IH FV G D ++ + ++MYA
Sbjct: 253 FRQGTKIHAFVFKTGLDSDYYVSNSFMDMYA----------------------------- 345
Query: 310 YAKLGMVKDARFIFDQMIERDLVCWSAMISGYAESDQPQEALKLFDEM-LQKRSVPDQIT 368
+LG + R +FD++ ERD V W+ MISG + + +EA+++F M + + T
Sbjct: 346 --ELGRIDFVRKLFDEISERDSVSWNVMISGCVKCRRFEEAVEVFQRMRVDSNEKISEAT 519
Query: 369 MLSVISACSHVGALAQANWIHTYV 392
++S ++AC+ + IH ++
Sbjct: 520 VVSSLTACAASRNVEVGKEIHGFI 591
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.323 0.135 0.403
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 24,294,156
Number of Sequences: 36976
Number of extensions: 369185
Number of successful extensions: 2631
Number of sequences better than 10.0: 174
Number of HSP's better than 10.0 without gapping: 2262
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2523
length of query: 737
length of database: 9,014,727
effective HSP length: 103
effective length of query: 634
effective length of database: 5,206,199
effective search space: 3300730166
effective search space used: 3300730166
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 62 (28.5 bits)
Medicago: description of AC146852.5


