

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146819.12 + phase: 0 /pseudo
(1226 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
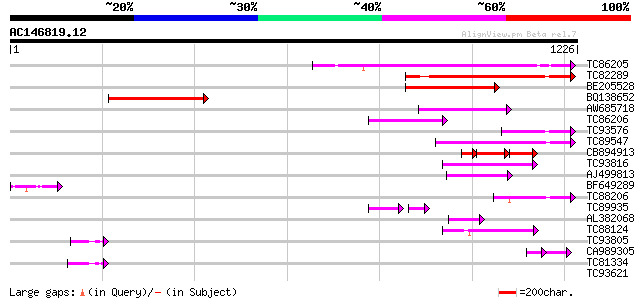
Score E
Sequences producing significant alignments: (bits) Value
TC86205 similar to PIR|A86289|A86289 probable ABC transporter [i... 308 8e-84
TC82289 similar to GP|12382007|dbj|BAB21276. putative ABC transp... 300 2e-81
BE205528 similar to GP|12382006|d putative ABC transporter prote... 224 1e-58
BQ138652 similar to GP|20522008|d pleiotropic drug resistance li... 189 6e-48
AW685718 homologue to GP|20522008|d pleiotropic drug resistance ... 139 7e-33
TC86206 similar to GP|20522008|dbj|BAB92011. pleiotropic drug re... 136 6e-32
TC93576 similar to GP|12382007|dbj|BAB21276. putative ABC transp... 108 1e-23
TC89547 similar to GP|12382007|dbj|BAB21276. putative ABC transp... 102 9e-22
CB894913 similar to GP|20522008|d pleiotropic drug resistance li... 55 4e-19
TC93816 similar to GP|9279716|dbj|BAB01273.1 ABC transporter {Ar... 93 7e-19
AJ499813 similar to GP|9279716|db ABC transporter {Arabidopsis t... 77 3e-14
BF649289 weakly similar to GP|16604683|gb putative ABC transport... 77 5e-14
TC88206 similar to PIR|A86289|A86289 probable ABC transporter [i... 74 3e-13
TC89935 similar to GP|16604683|gb|AAL24134.1 putative ABC transp... 44 3e-10
AL382068 homologue to GP|9279716|dbj ABC transporter {Arabidopsi... 59 9e-09
TC88124 weakly similar to PIR|T45888|T45888 ABC transporter-like... 54 3e-07
TC93805 weakly similar to GP|11994269|dbj|BAB01452. ABC transpor... 47 6e-05
CA989305 similar to GP|20522008|d pleiotropic drug resistance li... 36 1e-04
TC81334 weakly similar to GP|19423992|gb|AAL87274.1 putative ABC... 45 1e-04
TC93621 similar to GP|12382007|dbj|BAB21276. putative ABC transp... 35 0.003
>TC86205 similar to PIR|A86289|A86289 probable ABC transporter [imported] -
Arabidopsis thaliana, partial (52%)
Length = 2721
Score = 308 bits (789), Expect = 8e-84
Identities = 221/582 (37%), Positives = 311/582 (52%), Gaps = 14/582 (2%)
Frame = +3
Query: 655 LKRQYKEVVVMGILDITCNVWTECLVE**VPREDMETC-CTELNRTTRSSSFEISWILYS 713
LK+ Y+ +V +G+LD+T +VW EC *+ + C C R +R +L +
Sbjct: 252 LKKGYQRLVDLGLLDLTFDVWAECFDGQ*ISWAQLAQCYC*FGKRLSRYPR-----VLPT 416
Query: 714 VQLVLDRFWCLDWIHITIHHWLHSCSYFLEPN--GLCST-*RTSGR*I----RSIA---- 762
LVLDR W + WI + + S P C+ *R R* RS
Sbjct: 417 CILVLDRRWGIGWICVPFQRGIWCGSRCTRPI**A*CNNN*RRFRR*FIYRTRS*ITTY* 596
Query: 763 -IQEEKICNGK*TLWEERNDSFF*TTLYHL**SNIFS*YATGNEESTSCRGKVEPIEWCQ 821
+ ++ C E+ N S *TT YH ** I *+A+GNE + RG++ E C
Sbjct: 597 KFR*KRFCYRIQPWKEKGNGSSI*TTFYHF**YCILC*HASGNEGARCYRGQISAFEGC* 776
Query: 822 WFIQTGCSHSSDGCDWCWQNNSDGCTSWKKNQRIYWWDHHNFWLFKEAGNLCKSLWIL*T 881
W IQ C++ DGC W W NNSDGC+ W +N+ IY W H FW+ +A N+C +L +L*
Sbjct: 777 WRIQAWCTYGFDGCKWSW*NNSDGCSGW*ENRWIY*WRHQGFWVS*KARNIC*NLRLL*A 956
Query: 882 KLYPFSLCYCI*IIALFGMAPIVCRNQC*NQEDVHRGSYGACRTDPTEGYNC-CAWCYWS 940
+ YPF+ CY +*+ AL +A R+ +Q+DV+ S+G+ + E + AWC WS
Sbjct: 957 E*YPFTSCYSL*VFALLSVASFTFRS*FQHQKDVY*RSHGSSGAELVEELSSWFAWCEWS 1136
Query: 941 LYTTAKKVDYCGRVSGKSFNNVHG*ANFRARCKICRYCYESH*KYSGEWKNCCLCHSSIK 1000
L TA++ DYC R G S N HG*A F RC CY + ++SG KN C+ H S
Sbjct: 1137 LN*TAQETDYCSRTCG*SIYNFHG*AYFWFRC*SGCNCYANCKEHSGHGKNSCVYHPST* 1316
Query: 1001 Y*YI*IF**AVIDEARRASDICGANRTSFFPFNKLF*GD*RCQ*D*RWL*PSSLDVGNYF 1060
+*+I* **A+ EA R +IC +SF+ F+++F* *R +*+ R +*PS++DVG+Y
Sbjct: 1317 H*HI*SI**AIPYEAWRTRNICWTTGSSFYSFDQVF*EH*RSE*NQRRI*PSNMDVGSYN 1496
Query: 1061 FRKRNAIGD*FF*SVQKFRIIQEKQSSYCRIEHSSS*FCESSFSFKILKTPFRSIQGLLM 1120
+R R G *F * VQKF I EKQ++Y R + S F SSF + IL + GLLM
Sbjct: 1497 YRTRT*FGR*FH*LVQKF*SI*EKQATYSRT*RTCSWFKGSSFPYSILAVFLGPMPGLLM 1676
Query: 1121 ETTLVLLAESAIQCIKISLHSRCLYLFRERVLWPWLQNVHFNQLQ*KETRSP*LHWLNVD 1180
ETTLV+LA S I C ++ LH Y+ R V W + + L+* TR W NV
Sbjct: 1677 ETTLVILA*STIHCCEVFLH----YIHRLDV-WNNVLGPRWKTLK*--TRPSERCWFNVY 1835
Query: 1181 YYSTYWH*ECWFCASSGDCRESSFL*GKCCQNVFSTGICFWS 1222
W + +FCA+ G ++ L* K C NV +C ++
Sbjct: 1836 RCPLPWSTKFFFCAARGCSGKNCIL*RKSCWNVLCLTLCIFT 1961
Score = 40.4 bits (93), Expect = 0.004
Identities = 39/147 (26%), Positives = 66/147 (44%), Gaps = 17/147 (11%)
Frame = +1
Query: 83 VDIPTIEVRFEHLNVQAQVHVGKRAL------HTITN----YMLDLVEYILKR--RKQQL 130
V++P IE +V H K+ + H+IT Y +D+ + ++ + +L
Sbjct: 577 VELPRIESSGRRDSVTESSHGKKKGMVLPFEPHSITFDDIVYSVDMPAEMKEQGVTEDRL 756
Query: 131 NILQDVSGILKHSRLTLLLGPPNSGKTILLLALAGK-----LDPNLKIANEVQFYEQFAG 185
+L+ VSG + LT L+G +GKT L+ LAG+ +D ++K++ + E FA
Sbjct: 757 VLLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGYIDGDIKVSGYPKKQETFA- 933
Query: 186 KVSYNGHEMNEFVPQRTAAYVSQNDTH 212
R + Y QND H
Sbjct: 934 ---------------RISGYCEQNDIH 969
>TC82289 similar to GP|12382007|dbj|BAB21276. putative ABC transporter protein
{Oryza sativa (japonica cultivar-group)}, partial (25%)
Length = 1108
Score = 300 bits (769), Expect = 2e-81
Identities = 187/369 (50%), Positives = 227/369 (60%), Gaps = 1/369 (0%)
Frame = +2
Query: 856 YWWDHHNFWLFKEAGNLCKSLWIL*TKLYPFSLCYCI*IIALFGMAPIVCRNQC*NQEDV 915
Y W+HHNF L KEA NLCK+ W+L* K + R+QC*N +DV
Sbjct: 2 YLWEHHNFRLSKEARNLCKNFWVL*AK**ALA------------------RHQC*N*KDV 127
Query: 916 HRGSYGACRTDPT-EGYNCCAWCYWSLYTTAKKVDYCGRVSGKSFNNVHG*ANFRARCKI 974
GS+G C T+ T + + AWC WSL A+ VD C V SFNN+HG* NF ARCK
Sbjct: 128 R*GSHGTCGTETTAKRISRTAWC*WSLDGAAQTVDNCS*VGS*SFNNIHG*TNFWARCKG 307
Query: 975 CRYCYESH*KYSGEWKNCCLCHSSIKY*YI*IF**AVIDEARRASDICGANRTSFFPFNK 1034
C YC E+ *++S WKN CL + +*YI*IF**A ARRA DICGA T FF FN+
Sbjct: 308 CSYCDENS*EHSKHWKNSCLYYPPA*H*YI*IF**AFAT*ARRARDICGATWT*FFQFNQ 487
Query: 1035 LF*GD*RCQ*D*RWL*PSSLDVGNYFFRKRNAIGD*FF*SVQKFRIIQEKQSSYCRIEHS 1094
L *GD C D*R L* S++DVG++ F KR IGD*F V KFR+IQEKQS+Y RIE+S
Sbjct: 488 LL*GDPWC*QD*RRL*SSNMDVGSHNFIKRKGIGD*FCRGVPKFRVIQEKQSTYQRIEYS 667
Query: 1095 SS*FCESSFSFKILKTPFRSIQGLLMETTLVLLAESAIQCIKISLHSRCLYLFRERVLWP 1154
S+ F S F F +L+ ++ GLLMETTLVLLA+S I C IS+ +R + VL P
Sbjct: 668 STLFERSLFCFTVLEILLDTMHGLLMETTLVLLAQS*I*CH*ISVLNRSCCFVWKHVLGP 847
Query: 1155 WLQNVHFNQLQ*KETRSP*LHWLNVDYYSTYWH*ECWFCASSGDCRESSFL*GKCCQNVF 1214
WLQN *K TRS * H ++V WH EC F A+SG CR++ L* K +NVF
Sbjct: 848 WLQN-------*KGTRSF*CHGVHVFCCDRNWHQEC*FSAASGCCRKNGLL*RKSSRNVF 1006
Query: 1215 STGICFWSG 1223
S ICF SG
Sbjct: 1007 SFSICFCSG 1033
>BE205528 similar to GP|12382006|d putative ABC transporter protein {Oryza
sativa (japonica cultivar-group)}, partial (14%)
Length = 617
Score = 224 bits (572), Expect = 1e-58
Identities = 120/205 (58%), Positives = 143/205 (69%), Gaps = 1/205 (0%)
Frame = +3
Query: 856 YWWDHHNFWLFKEAGNLCKSLWIL*TKLYPFSLCYCI*IIALFGMAPIVCRNQC*NQEDV 915
YWW+HHNF L KEA NLCK+ W+L* K YP SLCYC+*I ALF MAP+V R+QC*N +DV
Sbjct: 3 YWWEHHNFRLSKEARNLCKNFWVL*AK*YPLSLCYCL*IFALFSMAPVVPRHQC*N*KDV 182
Query: 916 HRGSYGACRTDPT-EGYNCCAWCYWSLYTTAKKVDYCGRVSGKSFNNVHG*ANFRARCKI 974
GS+G C T+ T + + AWC WSL A+ VD C V SFNN+HG* NF ARCK
Sbjct: 183 R*GSHGTCGTETTAKRISRTAWC*WSLDGAAQTVDNCS*VGS*SFNNIHG*TNFWARCKG 362
Query: 975 CRYCYESH*KYSGEWKNCCLCHSSIKY*YI*IF**AVIDEARRASDICGANRTSFFPFNK 1034
C YC E+ *++S WKN CL + +*YI*IF**A ARRA DICGA T FF FN+
Sbjct: 363 CSYCDENS*EHSKHWKNSCLYYPPA*H*YI*IF**AFAT*ARRARDICGATWT*FFQFNQ 542
Query: 1035 LF*GD*RCQ*D*RWL*PSSLDVGNY 1059
L *GD C D*R L* S++DVG++
Sbjct: 543 LL*GDPWC*QD*RRL*SSNMDVGSH 617
>BQ138652 similar to GP|20522008|d pleiotropic drug resistance like protein
{Nicotiana tabacum}, partial (15%)
Length = 670
Score = 189 bits (480), Expect = 6e-48
Identities = 123/218 (56%), Positives = 143/218 (65%)
Frame = +3
Query: 213 LGN*LSEKPWPFQQEFKELDLDMIC*KRCVEERWRKTLFLIRISMSI*RL*QLRTREQML 272
L *L EKPW QQ KEL L M C*+ C EE+ + LI+ M *RL*QL+ R Q+
Sbjct: 12 LEK*LLEKPWLSQQGSKELGLVMTC*QSCPEEKNMQISCLIQTLMCT*RL*QLKARRQI* 191
Query: 273 *QIIF*RFWDWTYVKILW*EMRS*KASLKDKGNVLQ*GRR*SDH*NLYSWMIYLLVWMTQ 332
*QI+ * FWD YV IL *EM+ + SL DK N LQ GR D LYSWM YLLVW+ +
Sbjct: 192 *QIMS*GFWD*RYVLILL*EMQCYEVSLVDKRNALQLGRCWLDRRKLYSWMKYLLVWIAR 371
Query: 333 QPFKL*NR*NNLSTFSKELL*SHYNSHH*RLTIFLMTLFYSPMDTLCTKVLVYKCLIFLL 392
Q F+L* +*+NLSTFSKEL S SHH RLTIFL TLFYS + TL TKV V CL F
Sbjct: 372 QLFRL*IQ*SNLSTFSKELRSSRSFSHHQRLTIFLTTLFYSLIVTLYTKVPVSTCLNFSN 551
Query: 393 Q*VLCVLRGNQW*TFYKK*HQ*KIRNNIGHIKKSLIYL 430
Q VL VL G +W F KK*HQ K +++IG+ K SLI L
Sbjct: 552 QLVLNVLIGKEWQIFCKK*HQGKTKSSIGNTKTSLIXL 665
>AW685718 homologue to GP|20522008|d pleiotropic drug resistance like protein
{Nicotiana tabacum}, partial (12%)
Length = 613
Score = 139 bits (350), Expect = 7e-33
Identities = 84/203 (41%), Positives = 116/203 (56%), Gaps = 2/203 (0%)
Frame = +1
Query: 885 PFSLCYCI*IIALFGMAPIVCRNQC*NQEDVHRGSYGACRTDPTEG-YNCCAWCYWSLYT 943
PFS CYC+*+ L G+A R Q+DVH GS+G P E + AWC WS
Sbjct: 1 PFSSCYCL*VTGLLGLASFTSRR*FKYQKDVH*GSHGTSGAQPIEKLFGWFAWCEWSFNR 180
Query: 944 TAKKVDYCGRVSGKSFNNVHG*ANFRARCKICRYCYESH*KYSGEWKNCCLCHSSIKY*Y 1003
T ++ +YC +SG+S +N HG*A RC C CY + *++ KN CL HSS KY*+
Sbjct: 181 TTQEANYCS*ISGQSIHNFHG*AYIWIRC*SCCNCYANR*EHG*HRKNSCLHHSSTKY*H 360
Query: 1004 I*IF**AVIDEARRASDICGANRTSFFPFNKLF*GD*RCQ*D*RWL*PSSLDVGNYFFRK 1063
I F** + E RR +IC +SF +++F* *R *+ RWL* S+LDVG++ F
Sbjct: 361 IRSF**VIPHETRRRRNICWTAGSSF*SIDQVF*EH*RG**NQRWL*SSNLDVGSFEFST 540
Query: 1064 RNAIGD*FF*SVQK-FRIIQEKQ 1085
+N + + F K F ++QE +
Sbjct: 541 KNLL*ELIFHHAYKNF*VVQENK 609
>TC86206 similar to GP|20522008|dbj|BAB92011. pleiotropic drug resistance
like protein {Nicotiana tabacum}, partial (11%)
Length = 757
Score = 136 bits (342), Expect = 6e-32
Identities = 77/174 (44%), Positives = 101/174 (57%), Gaps = 2/174 (1%)
Frame = +2
Query: 776 WEER-NDSFF*TTLYHL**SNIFS*YATGNEESTSCRGKVEPIEWCQWFIQTGCSHSSDG 834
W+E+ N S *TT YH ** I *+A+ NE + RG+ E C IQ CSHS DG
Sbjct: 209 WKEKGNGSSI*TTFYHF**YCILC*HAS*NEGARCYRGQTSAFEGC*RCIQARCSHSFDG 388
Query: 835 CDWCWQNNSDGCTSWKKNQRIYWWDHHNFWLFKEAGNLCKSLWIL*TKLYPFSLCYCI*I 894
C W W NNSDGC+ W +N+ IY W H FW+ +A N+C ++W+L*T+ Y F+ CY +*I
Sbjct: 389 CKWSW*NNSDGCSGW*ENRWIY*WRHQGFWVP*KARNIC*NIWLL*TE*YSFTSCYSL*I 568
Query: 895 IALFGMAPIVCRNQC*NQEDVHRGSYGACRTDPTEGY-NCCAWCYWSLYTTAKK 947
+L MA + +DV+ GS G+C + EG AWC WSL T K
Sbjct: 569 PSLLSMASFTFWS*FQY*KDVY*GSNGSCGAELVEGLPRWFAWC*WSLNGTNVK 730
Score = 40.0 bits (92), Expect = 0.006
Identities = 39/147 (26%), Positives = 66/147 (44%), Gaps = 17/147 (11%)
Frame = +3
Query: 83 VDIPTIEVRFEHLNVQAQVHVGKRAL------HTITN----YMLDLVEYILKR--RKQQL 130
V++P IE +V H K+ + H+IT Y +D+ + ++ + +L
Sbjct: 150 VELPRIESSGRGDSVTVSSHGKKKGMVLPFEPHSITFDDIVYSVDMPAEMKEQGVTEDRL 329
Query: 131 NILQDVSGILKHSRLTLLLGPPNSGKTILLLALAGK-----LDPNLKIANEVQFYEQFAG 185
+L+ VSG + LT L+G +GKT L+ LAG+ +D ++K++ + E FA
Sbjct: 330 VLLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGYIDGDIKVSGYPKKQETFA- 506
Query: 186 KVSYNGHEMNEFVPQRTAAYVSQNDTH 212
R + Y QND H
Sbjct: 507 ---------------RISGYCEQNDIH 542
>TC93576 similar to GP|12382007|dbj|BAB21276. putative ABC transporter protein
{Oryza sativa (japonica cultivar-group)}, partial (18%)
Length = 814
Score = 108 bits (271), Expect = 1e-23
Identities = 74/160 (46%), Positives = 96/160 (59%)
Frame = +2
Query: 1064 RNAIGD*FF*SVQKFRIIQEKQSSYCRIEHSSS*FCESSFSFKILKTPFRSIQGLLMETT 1123
R+ GD*F VQKFR+IQ+KQS+Y I SS+ F S FS+ +L+ F ++ GLLMETT
Sbjct: 11 RSGTGD*FCRVVQKFRVIQDKQSTYQGIG*SSTLFKRSLFSYSVLEILFYTMYGLLMETT 190
Query: 1124 LVLLAESAIQCIKISLHSRCLYLFRERVLWPWLQNVHFNQLQ*KETRSP*LHWLNVDYYS 1183
LVLLA+S I C KIS+ +R R+ VL P LQN *K RS * H ++V
Sbjct: 191 LVLLAQS*I*CHKISVLNRSGCFARKHVLGP*LQN-------*KGARSF*CHGVHVCCCY 349
Query: 1184 TYWH*ECWFCASSGDCRESSFL*GKCCQNVFSTGICFWSG 1223
W E F A+ G C ++ L* K +NVF+ IC W+G
Sbjct: 350 PNWRHEW*FSAAGGCC*KNGLL*RKSSRNVFNFSICVWAG 469
>TC89547 similar to GP|12382007|dbj|BAB21276. putative ABC transporter protein
{Oryza sativa (japonica cultivar-group)}, partial (28%)
Length = 1283
Score = 102 bits (254), Expect = 9e-22
Identities = 94/305 (30%), Positives = 142/305 (45%), Gaps = 1/305 (0%)
Frame = +2
Query: 920 YGACRTDPTEGYNC-CAWCYWSLYTTAKKVDYCGRVSGKSFNNVHG*ANFRARCKICRYC 978
YGACR + C A W+ T ++ + +S +N+HG*AN C C
Sbjct: 5 YGACRA*FIKRSTCWIARRDWTFDRTTQETNDRS*TCSESIDNIHG*ANLWP*C*SSCNC 184
Query: 979 YESH*KYSGEWKNCCLCHSSIKY*YI*IF**AVIDEARRASDICGANRTSFFPFNKLF*G 1038
E+ ++S CCL + S KY*+I **A E R ++I + R F+ L *G
Sbjct: 185 NENSEEHS*HRTYCCLHNPSAKY*HIRRL**ASTYEIGR*ANIFWSIRPPLRSFDSLL*G 364
Query: 1039 D*RCQ*D*RWL*PSSLDVGNYFFRKRNAIGD*FF*SVQKFRIIQEKQSSYCRIEHSSS*F 1098
*R *D*RWL* ++DVG+Y R R+ * +QK I EKQ+ I+++SS F
Sbjct: 365 Y*RSS*D*RWL*SCNMDVGSYISRIRSKS*GQLH*RLQKLGTIPEKQTIDPGIKYTSSRF 544
Query: 1099 CESSFSFKILKTPFRSIQGLLMETTLVLLAESAIQCIKISLHSRCLYLFRERVLWPWLQN 1158
F + + +L+ETT ++LA+ I +++ +L +L WL+
Sbjct: 545 KGIIL*FSVYTNYVVPM*SMLVETTFIVLAKHVIYSS*TVIYNTDSFLVWNHILEHWLEK 724
Query: 1159 VHFNQLQ*KETRSP*LHWLNVDYYSTYWH*ECWFCASSGDCRESSFL*GKCCQNVFSTGI 1218
K RS + ++V + YW + A+ E+S L K C NVF +
Sbjct: 725 E-------KGARSFQCNGIDVCFGYIYWSTKWCLSAACNSS*ENSLLQRKSCWNVFRFAL 883
Query: 1219 CFWSG 1223
C +G
Sbjct: 884 CSSTG 898
>CB894913 similar to GP|20522008|d pleiotropic drug resistance like protein
{Nicotiana tabacum}, partial (12%)
Length = 765
Score = 54.7 bits (130), Expect(3) = 4e-19
Identities = 30/60 (50%), Positives = 39/60 (65%)
Frame = +1
Query: 1081 IQEKQSSYCRIEHSSS*FCESSFSFKILKTPFRSIQGLLMETTLVLLAESAIQCIKISLH 1140
+QEKQ++Y R + S F SSF + IL S+ GLLMETTLV+LA S I C ++ LH
Sbjct: 541 LQEKQATYSRT*RACSWFKGSSFPYSILAVFLGSMPGLLMETTLVILA*STIHCCEVFLH 720
Score = 53.5 bits (127), Expect(3) = 4e-19
Identities = 31/72 (43%), Positives = 45/72 (62%)
Frame = +2
Query: 1010 AVIDEARRASDICGANRTSFFPFNKLF*GD*RCQ*D*RWL*PSSLDVGNYFFRKRNAIGD 1069
A+ EA R +IC +SF+ F+++F *R *+ RW+*PS++DVG+Y + R G
Sbjct: 215 AIPYEAWRTRNICWTTGSSFYSFDQVFREH*RS**NQRWI*PSNMDVGSYNYSTRT*FGC 394
Query: 1070 *FF*SVQKFRII 1081
* + VQKFR I
Sbjct: 395 *LYRFVQKFRFI 430
Score = 25.8 bits (55), Expect(3) = 4e-19
Identities = 14/34 (41%), Positives = 21/34 (61%)
Frame = +1
Query: 978 CYESH*KYSGEWKNCCLCHSSIKY*YI*IF**AV 1011
CY + ++SG KN C+ H S +*+I* ** +
Sbjct: 10 CYANCKEHSGHGKNSCVYHPST*H*HI*SI**GM 111
>TC93816 similar to GP|9279716|dbj|BAB01273.1 ABC transporter {Arabidopsis
thaliana}, partial (16%)
Length = 743
Score = 92.8 bits (229), Expect = 7e-19
Identities = 71/204 (34%), Positives = 101/204 (48%)
Frame = +3
Query: 937 CYWSLYTTAKKVDYCGRVSGKSFNNVHG*ANFRARCKICRYCYESH*KYSGEWKNCCLCH 996
CY + T KK + G SFN+ HG* NFR+ CK C CYE K+ WKN CL +
Sbjct: 18 CYRVVNRTTKKANNSG*AYC*SFNHFHG*TNFRS*CKSCSNCYEDSEKHC*HWKNSCLHN 197
Query: 997 SSIKY*YI*IF**AVIDEARRASDICGANRTSFFPFNKLF*GD*RCQ*D*RWL*PSSLDV 1056
S Y Y+* A DE RRASD+ R F ++F*G+ R *+* + S++DV
Sbjct: 198 SPA*YRYL*SLRRAPFDEERRASDLLRTIRKEFTQDYRIF*GNSRSP*N*GEVQSSNMDV 377
Query: 1057 GNYFFRKRNAIGD*FF*SVQKFRIIQEKQSSYCRIEHSSS*FCESSFSFKILKTPFRSIQ 1116
+ + * +Q I EKQSS ++++++ F + F +I
Sbjct: 378 RGKLNSS*S*AWNGLC*ILQNIDIASEKQSSCK*VKYTTTRSKRCIFLHSVFSVNFWTI* 557
Query: 1117 GLLMETTLVLLAESAIQCIKISLH 1140
+ ME + LL ES +Q +I LH
Sbjct: 558 IMSMEAMVDLLEESRLQPCQIFLH 629
>AJ499813 similar to GP|9279716|db ABC transporter {Arabidopsis thaliana},
partial (13%)
Length = 573
Score = 77.4 bits (189), Expect = 3e-14
Identities = 55/144 (38%), Positives = 78/144 (53%)
Frame = -2
Query: 944 TAKKVDYCGRVSGKSFNNVHG*ANFRARCKICRYCYESH*KYSGEWKNCCLCHSSIKY*Y 1003
T K+ D CGR SFN++HG*ANF +RCK YCYE K+ WKN C+ +SS +*Y
Sbjct: 446 TKKEADNCGRACC*SFNHIHG*ANFWSRCKSSCYCYEDCKKHC*YWKNSCVHNSST*H*Y 267
Query: 1004 I*IF**AVIDEARRASDICGANRTSFFPFNKLF*GD*RCQ*D*RWL*PSSLDVGNYFFRK 1063
+* **A IDE RRA+ + R+ F + *G+ C + + + D + +
Sbjct: 266 L*SL**AHIDEKRRATCLWRTFRSEFTQDY*IL*GNSWCPKNQGNVQSCNTDARS*LRSR 87
Query: 1064 RNAIGD*FF*SVQKFRIIQEKQSS 1087
+ + F * +Q + EKQSS
Sbjct: 86 *SPSWNGFC*ILQVISFVSEKQSS 15
>BF649289 weakly similar to GP|16604683|gb putative ABC transporter protein
{Arabidopsis thaliana}, partial (12%)
Length = 654
Score = 76.6 bits (187), Expect = 5e-14
Identities = 46/123 (37%), Positives = 73/123 (58%), Gaps = 10/123 (8%)
Frame = +3
Query: 1 MESDEISLMKWDSIQRLPTVARLRRGLLTT-PEGDS---------NEIDVHKIGLQERTY 50
++ DE +L KW +I++LPT RLR ++ T EGD E+DV K+ + ER
Sbjct: 297 VDEDEEAL-KWAAIEKLPTYDRLRTSIMQTFTEGDQPQPGNRQQHKEVDVTKLDMNERQQ 473
Query: 51 LLQRLLRNNTVEVDNDHSFLLKLMRDRIDRAGVDIPTIEVRFEHLNVQAQVHVGKRALHT 110
++ ++ + E DN+ L+ R+RID+ G+ +PT+EVRF++L V+A VG AL T
Sbjct: 474 IIDKIFK--VAEEDNEK--YLRKFRNRIDKVGIRLPTVEVRFKNLTVEADSFVGSXALPT 641
Query: 111 ITN 113
+ N
Sbjct: 642 LPN 650
>TC88206 similar to PIR|A86289|A86289 probable ABC transporter [imported] -
Arabidopsis thaliana, partial (21%)
Length = 1483
Score = 74.3 bits (181), Expect = 3e-13
Identities = 67/208 (32%), Positives = 91/208 (43%), Gaps = 31/208 (14%)
Frame = +3
Query: 1047 RWL*PSSLDVGNYFFRKRNAIGD*FF*SVQKF---------------------------- 1078
RWL* S+LDVG++ F R +*F +QKF
Sbjct: 123 RWL*SSNLDVGSFEFSTRTYSRN*FSSCIQKF*VVQVSKFI*KYGFII*YQL*SYWKKTS 302
Query: 1079 ---RIIQEKQSSYCRIEHSSS*FCESSFSFKILKTPFRSIQGLLMETTLVLLAESAIQCI 1135
I+QEKQ++Y RI + + F S F IL F + GLLMETTLV+LA+ AI
Sbjct: 303 LL*TIVQEKQATYRRIRQTCTWFK*SLFLCPILTIIFSPMFGLLMETTLVILAQPAIHFC 482
Query: 1136 KISLHSRCLYLFRERVLWPWLQNVHFNQLQ*KETRSP*LHWLNVDYYSTYWH*ECWFCAS 1195
++ LH +L W + + KETR * +V S W E C +
Sbjct: 483 EVFLHCFHRLDVWNNILGSWKEIL-------KETRPV*CPRFDVYC*SLPWGPEFISCTT 641
Query: 1196 SGDCRESSFL*GKCCQNVFSTGICFWSG 1223
SG + L* K C +VF +C +G
Sbjct: 642 SGSS*KICVL*RKSCWDVFCLTLCICTG 725
>TC89935 similar to GP|16604683|gb|AAL24134.1 putative ABC transporter protein
{Arabidopsis thaliana}, partial (34%)
Length = 1244
Score = 43.9 bits (102), Expect(2) = 3e-10
Identities = 21/47 (44%), Positives = 27/47 (56%)
Frame = +1
Query: 862 NFWLFKEAGNLCKSLWIL*TKLYPFSLCYCI*IIALFGMAPIVCRNQ 908
N WL E+ N+CK+ WIL T YPF+ YC I L G + R+Q
Sbjct: 1096 NLWLP*ESRNICKNFWILRTNRYPFTTSYCQRICDLLGFP*VT*RSQ 1236
Score = 40.0 bits (92), Expect(2) = 3e-10
Identities = 28/76 (36%), Positives = 37/76 (47%), Gaps = 1/76 (1%)
Frame = +2
Query: 777 EERNDSFF*TTLYHL**SNIFS*YATGNEESTSCRGKVEPIEWCQWFIQTGCSHSSDGCD 836
+ERN S F TT ** + YA+ NE + S R W IQ S+ +G
Sbjct: 839 KERNGSSFSTTCNVF**C*LLCRYAS*NERARSNR**ATIA*RSNWCIQAWSSNGFNGS* 1018
Query: 837 WCWQNNSDGCT-SWKK 851
W W++N DGC WK+
Sbjct: 1019WSWKDNFDGCF*PWKE 1066
>AL382068 homologue to GP|9279716|dbj ABC transporter {Arabidopsis thaliana},
partial (6%)
Length = 274
Score = 59.3 bits (142), Expect = 9e-09
Identities = 35/78 (44%), Positives = 46/78 (58%)
Frame = +2
Query: 950 YCGRVSGKSFNNVHG*ANFRARCKICRYCYESH*KYSGEWKNCCLCHSSIKY*YI*IF** 1009
YC R + FN++HG* N R RC Y + KYS +NCCL +S KY + * F**
Sbjct: 14 YCCRACCQPFNHLHG*TNIRPRC*SRCNRYANREKYSRYRENCCLHNS*TKYRHF*SF** 193
Query: 1010 AVIDEARRASDICGANRT 1027
A++DE RR S I +R+
Sbjct: 194 AIVDEKRRTSYIRWTSRS 247
>TC88124 weakly similar to PIR|T45888|T45888 ABC transporter-like protein -
Arabidopsis thaliana, partial (27%)
Length = 1641
Score = 54.3 bits (129), Expect = 3e-07
Identities = 59/215 (27%), Positives = 90/215 (41%), Gaps = 7/215 (3%)
Frame = +2
Query: 936 WCYWSLYTTAKKVDYCGRVSGKSFNNVHG*ANFRARCKICRYCYESH*KYSGEWKNCC-- 993
W W +Y K ++C +S +N +G* R C C Y + CC
Sbjct: 44 WPKWLIYRAT*KTNHCSGACIESIDNFYG*TYIRFGCSSCCSSYACS-------QECCYH 202
Query: 994 -----LCHSSIKY*YI*IF**AVIDEARRASDICGANRTSFFPFNKLF*GD*RCQ*D*RW 1048
L H+S KY YI* F** +E R + + R+SF + +F D
Sbjct: 203 R*DHSLHHTSTKYRYI*DF**VDSNEIWRKDHL*WSVRSSFK*TD*IFSEYIGSSKDQGQ 382
Query: 1049 L*PSSLDVGNYFFRKRNAIGD*FF*SVQKFRIIQEKQSSYCRIEHSSS*FCESSFSFKIL 1108
L* ++DVG+Y R R + F +Q+ +E + + F +F +
Sbjct: 383 L*SCNMDVGSYICRCRR*T*NRFCKYIQRVTPSSGYP*VGETVE*TRAKFKRLAFLNSLP 562
Query: 1109 KTPFRSIQGLLMETTLVLLAESAIQCIKISLHSRC 1143
++ G+ ME TLVLL +S +Q I +H C
Sbjct: 563 TE*LGTVYGVPMEATLVLLEKSRVQLNTICVHDCC 667
>TC93805 weakly similar to GP|11994269|dbj|BAB01452. ABC transporter-like
protein {Arabidopsis thaliana}, partial (15%)
Length = 649
Score = 46.6 bits (109), Expect = 6e-05
Identities = 29/83 (34%), Positives = 45/83 (53%)
Frame = +2
Query: 131 NILQDVSGILKHSRLTLLLGPPNSGKTILLLALAGKLDPNLKIANEVQFYEQFAGKVSYN 190
+ILQ ++G K +L ++GP GK+ LL LAG+L N + G++ N
Sbjct: 263 SILQGLTGYAKPGQLLAIMGPSGCGKSTLLDTLAGRLSSNTR----------QTGEILIN 412
Query: 191 GHEMNEFVPQRTAAYVSQNDTHL 213
GH+ + T+AYV+Q+DT L
Sbjct: 413 GHKQE--LSYGTSAYVTQDDTLL 475
>CA989305 similar to GP|20522008|d pleiotropic drug resistance like protein
{Nicotiana tabacum}, partial (7%)
Length = 655
Score = 35.8 bits (81), Expect(2) = 1e-04
Identities = 20/43 (46%), Positives = 25/43 (57%)
Frame = -2
Query: 1117 GLLMETTLVLLAESAIQCIKISLHSRCLYLFRERVLWPWLQNV 1159
G METTLV+LA+SAI C ++ LH VL PW + V
Sbjct: 615 GCPMETTLVILAQSAIHCCEVFLHYFHSIDVWNNVLEPWKEIV 487
Score = 28.9 bits (63), Expect(2) = 1e-04
Identities = 22/64 (34%), Positives = 27/64 (41%)
Frame = -1
Query: 1151 VLWPWLQNVHFNQLQ*KETRSP*LHWLNVDYYSTYWH*ECWFCASSGDCRESSFL*GKCC 1210
+ +P F Q K TRS * +V S W E C + G C + L* K C
Sbjct: 439 IFFPT*MLTSFVDFQLK*TRSV*CPRFDVYCCSLPWGPEFIVCTTRGSC*KICLL*RKSC 260
Query: 1211 QNVF 1214
NVF
Sbjct: 259 WNVF 248
>TC81334 weakly similar to GP|19423992|gb|AAL87274.1 putative ABC
transporter protein {Arabidopsis thaliana}, partial
(29%)
Length = 977
Score = 45.4 bits (106), Expect = 1e-04
Identities = 29/89 (32%), Positives = 50/89 (55%)
Frame = +2
Query: 125 RRKQQLNILQDVSGILKHSRLTLLLGPPNSGKTILLLALAGKLDPNLKIANEVQFYEQFA 184
+ + +ILQ ++G K ++L ++GP GK+ LL ALAG+L N + +
Sbjct: 113 KTNESKSILQGLTGYAKPAQLLAIMGPSGCGKSTLLDALAGRLGSNTR----------QS 262
Query: 185 GKVSYNGHEMNEFVPQRTAAYVSQNDTHL 213
G + NG++ + + T+AYV+Q+DT L
Sbjct: 263 GDILINGNK--QALAYGTSAYVTQDDTLL 343
>TC93621 similar to GP|12382007|dbj|BAB21276. putative ABC transporter protein
{Oryza sativa (japonica cultivar-group)}, partial (16%)
Length = 839
Score = 34.7 bits (78), Expect(2) = 0.003
Identities = 16/31 (51%), Positives = 20/31 (63%)
Frame = +1
Query: 1193 CASSGDCRESSFL*GKCCQNVFSTGICFWSG 1223
CA+SG C ++ L K +NVFS ICF SG
Sbjct: 157 CAASGSC*KNGLLSRKSSRNVFSFSICFCSG 249
Score = 25.0 bits (53), Expect(2) = 0.003
Identities = 18/39 (46%), Positives = 20/39 (51%)
Frame = +3
Query: 1151 VLWPWLQNVHFNQLQ*KETRSP*LHWLNVDYYSTYWH*E 1189
VL PWLQ *K TRS * H ++V S WH E
Sbjct: 51 VLEPWLQ-------Y*KGTRSF*CHGVHVFSRSPNWHQE 146
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.352 0.155 0.576
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 47,436,288
Number of Sequences: 36976
Number of extensions: 849085
Number of successful extensions: 12758
Number of sequences better than 10.0: 75
Number of HSP's better than 10.0 without gapping: 4093
Number of HSP's successfully gapped in prelim test: 645
Number of HSP's that attempted gapping in prelim test: 8078
Number of HSP's gapped (non-prelim): 5791
length of query: 1226
length of database: 9,014,727
effective HSP length: 107
effective length of query: 1119
effective length of database: 5,058,295
effective search space: 5660232105
effective search space used: 5660232105
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.5 bits)
S2: 64 (29.3 bits)
Medicago: description of AC146819.12


