

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146818.6 + phase: 0 /pseudo
(642 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
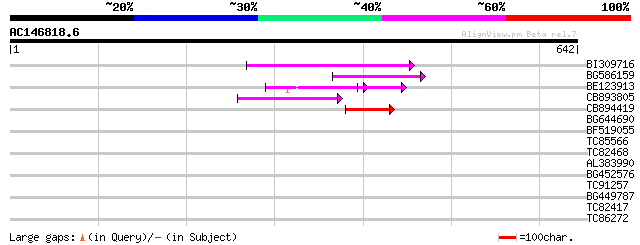
Score E
Sequences producing significant alignments: (bits) Value
BI309716 weakly similar to PIR|G96722|G9 hypothetical protein F2... 82 5e-16
BG586159 weakly similar to PIR|T47841|T4 hypothetical protein T2... 74 1e-13
BE123913 weakly similar to GP|22093573|d polyprotein {Oryza sati... 64 2e-10
CB893805 similar to GP|10177935|d copia-type polyprotein {Arabid... 60 3e-09
CB894419 weakly similar to GP|10177935|db copia-type polyprotein... 46 4e-05
BG644690 weakly similar to GP|18542179|gb putative pol protein {... 42 0.001
BF519055 similar to GP|21592754|gb| unknown {Arabidopsis thalian... 37 0.018
TC85566 homologue to SP|P27880|HS12_MEDSA 18.2 kDa class I heat ... 34 0.20
TC82468 similar to GP|8920639|gb|AAF81361.1| Identical to wall-a... 31 1.3
AL383990 31 1.3
BG452576 weakly similar to GP|12005223|gb|A reverse transcriptas... 30 2.9
TC91257 similar to GP|21434|emb|CAA36616.1|| ORF4 {Solanum tuber... 30 2.9
BG449787 similar to SP|Q9SZR0|CHMO Probable choline monooxygenas... 29 6.4
TC82417 similar to GP|15293221|gb|AAK93721.1 unknown protein {Ar... 28 8.3
TC86272 similar to GP|4204761|gb|AAD11482.1| peroxidase precurso... 28 8.3
>BI309716 weakly similar to PIR|G96722|G9 hypothetical protein F20P5.25
[imported] - Arabidopsis thaliana, partial (10%)
Length = 744
Score = 82.4 bits (202), Expect = 5e-16
Identities = 54/190 (28%), Positives = 92/190 (48%)
Frame = +2
Query: 269 NKKVLSTLKNLLQLQDWKQSGYFYPMQ*ILIKNDFKRGQVDTTLFRGTLKKYILVVQIYV 328
N KV K++ L+ + Y + LI + + D +LF + +YV
Sbjct: 11 NTKVCELQKSIYGLKQASRQWYS-KLSESLISFGYLQSSSDFSLFTKFKDSSFTTLLVYV 187
Query: 329 DDIIFGSTNASLCKEFSK*MQDEFEKSMMGELKFFLGIQINQRKDGVYVHQTKYTKELLK 388
DDI+ + S + + D F+ +G L++FLG+++ + K G+ ++Q KYT ELL+
Sbjct: 188 DDIVLAGNDISEIQHVKCFLIDRFKIKDLGSLRYFLGLEVARSKQGILLNQRKYTLELLE 367
Query: 389 KFKLEDCKVMNTPMHPTCT*SKEDIGTKVDQKLYIGMIGSLLYLTASRPDILFSVCLCAR 448
K TP + D D+ Y +IG L+YLT +RPDI F+V ++
Sbjct: 368 DSGNLAVKSTLTPYDISLKLHNSDSPLYNDETQYRRLIGKLIYLTTTRPDISFAVQQLSQ 547
Query: 449 FQSNPKRISF 458
F S P+++ +
Sbjct: 548 FVSKPQQVHY 577
>BG586159 weakly similar to PIR|T47841|T4 hypothetical protein T2O9.150 -
Arabidopsis thaliana, partial (11%)
Length = 732
Score = 74.3 bits (181), Expect = 1e-13
Identities = 33/105 (31%), Positives = 60/105 (56%)
Frame = +1
Query: 366 IQINQRKDGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCT*SKEDIGTKVDQKLYIGM 425
+++ Q ++G+Y+ Q KY +LL++F +E + P+ P C K++ G KVD Y +
Sbjct: 1 VEVIQNEEGIYICQRKYVTDLLERFGMEKSNLSRNPIAPRCKLIKDENGVKVDATKYKQI 180
Query: 426 IGSLLYLTASRPDILFSVCLCARFQSNPKRISFNCC*ENLHVFEG 470
+G L+YL A+RPD+++ + L +RF + P + + L G
Sbjct: 181 VGCLMYLAATRPDLMYVLSLISRFMNCPTELHMHAVKRVLRYLNG 315
>BE123913 weakly similar to GP|22093573|d polyprotein {Oryza sativa (japonica
cultivar-group)}, partial (8%)
Length = 503
Score = 63.5 bits (153), Expect = 2e-10
Identities = 39/120 (32%), Positives = 62/120 (51%), Gaps = 3/120 (2%)
Frame = +1
Query: 290 YFYPMQ*ILIKNDFKRGQVDTTLF---RGTLKKYILVVQIYVDDIIFGSTNASLCKEFSK 346
+F *++ K + + Q D +F T+KK IL+V YVDDI + K
Sbjct: 16 WFDRFT*VVKKFGYIQCQTDHAMFIKHSSTVKKAILIV--YVDDIFLTGDHGK*IKRLKN 189
Query: 347 *MQDEFEKSMMGELKFFLGIQINQRKDGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTC 406
+ +EFE +G LK+FLG+++ + K G + Q KY +LLK+ ++ CK + P C
Sbjct: 190 LLAEEFEIKDLGNLKYFLGMEVARWKKGSSISQRKYVLDLLKETRMIGCKTIRDPYGCNC 369
Score = 46.6 bits (109), Expect = 3e-05
Identities = 24/56 (42%), Positives = 32/56 (56%)
Frame = +2
Query: 394 DCKVMNTPMHPTCT*SKEDIGTKVDQKLYIGMIGSLLYLTASRPDILFSVCLCARF 449
D K TPM T D GT VD+ Y ++G L+YL+ +RPDI F VC ++F
Sbjct: 332 DVKPSETPMDATVKLGTLDNGTLVDKGRYQRLVGKLIYLSHTRPDISFVVCTMSQF 499
>CB893805 similar to GP|10177935|d copia-type polyprotein {Arabidopsis
thaliana}, partial (14%)
Length = 778
Score = 59.7 bits (143), Expect = 3e-09
Identities = 34/121 (28%), Positives = 68/121 (56%), Gaps = 2/121 (1%)
Frame = +3
Query: 259 KLYLLNKVTVNKKVLS-TLKNLLQLQDWKQSGYFYPMQ*ILIKNDFKRGQVDTTLF-RGT 316
K L+N + ++V+S +K L ++ ++ K F++ + TLF + +
Sbjct: 411 KFLLINHRVM*RRVIS*RVKRALYGLKQAPRAWYSRIEAYFTKEGFEKCPYEHTLFVKLS 590
Query: 317 LKKYILVVQIYVDDIIFGSTNASLCKEFSK*MQDEFEKSMMGELKFFLGIQINQRKDGVY 376
IL++ +YVDD+IF + ++ +EF K M+ EF S +G++ +FLG+++ Q + G+Y
Sbjct: 591 EGGKILIISLYVDDLIFIGNDENMFEEFKKSMKKEFNMSDLGKMHYFLGVEVTQNEKGIY 770
Query: 377 V 377
+
Sbjct: 771 I 773
>CB894419 weakly similar to GP|10177935|db copia-type polyprotein
{Arabidopsis thaliana}, partial (2%)
Length = 170
Score = 46.2 bits (108), Expect = 4e-05
Identities = 23/55 (41%), Positives = 34/55 (61%)
Frame = +3
Query: 381 KYTKELLKKFKLEDCKVMNTPMHPTCT*SKEDIGTKVDQKLYIGMIGSLLYLTAS 435
K ++LKKFK+E K ++TP+ ++E G +VD Y +IGSL YLTA+
Sbjct: 6 KCASDILKKFKMEHSKPISTPVEEKLKLTRESDGKRVDSTHYKSLIGSLRYLTAT 170
>BG644690 weakly similar to GP|18542179|gb putative pol protein {Zea mays},
partial (22%)
Length = 629
Score = 41.6 bits (96), Expect = 0.001
Identities = 18/34 (52%), Positives = 26/34 (75%)
Frame = -2
Query: 298 LIKNDFKRGQVDTTLFRGTLKKYILVVQIYVDDI 331
L+KN FKRG++D TLF + +L++Q+YVDDI
Sbjct: 103 LLKNGFKRGKIDNTLFLLKRE*ELLIIQVYVDDI 2
Score = 39.3 bits (90), Expect = 0.005
Identities = 22/56 (39%), Positives = 30/56 (53%)
Frame = -1
Query: 193 YQ*LNLPQMMKHFQMMDGY*LCKKS*ISFKEMMCEIWYPNLSRRTLLEQNGYSETS 248
Y L+ + KH M G LCKK+ IS KE+ W+ +L + LE G+ ETS
Sbjct: 599 YHLLSPRMLKKHCVMQTGSILCKKNSISLKEVRYGTWFLDLKAKQ*LELGGFLETS 432
>BF519055 similar to GP|21592754|gb| unknown {Arabidopsis thaliana}, partial
(30%)
Length = 675
Score = 37.4 bits (85), Expect = 0.018
Identities = 16/41 (39%), Positives = 27/41 (65%)
Frame = -3
Query: 418 DQKLYIGMIGSLLYLTASRPDILFSVCLCARFQSNPKRISF 458
D KL ++G+LLY+T + PD+ FS+ ++F P +I+F
Sbjct: 646 DSKLVHSIVGALLYITVTCPDLSFSINKPSQFMHKPTQINF 524
>TC85566 homologue to SP|P27880|HS12_MEDSA 18.2 kDa class I heat shock
protein. [Alfalfa] {Medicago sativa}, complete
Length = 946
Score = 33.9 bits (76), Expect = 0.20
Identities = 16/40 (40%), Positives = 26/40 (65%)
Frame = -1
Query: 364 LGIQINQRKDGVYVHQTKYTKELLKKFKLEDCKVMNTPMH 403
LG+++ Q V + Q KYT+E++K DCK +NTP++
Sbjct: 811 LGMEVRQDNK*VLICQMKYTREIMK-----DCKRINTPVN 707
>TC82468 similar to GP|8920639|gb|AAF81361.1| Identical to wall-associated
kinase 4 from Arabidopsis thaliana gb|AJ009695 and
contains Eukaryotic, partial (22%)
Length = 937
Score = 31.2 bits (69), Expect = 1.3
Identities = 16/41 (39%), Positives = 25/41 (60%), Gaps = 1/41 (2%)
Frame = +3
Query: 220 SFKEMMCEIWYPN-LSRRTLLEQNGYSETS*MNRVK*SETK 259
SFK+M+ WY N L +TLL+Q+ YS+ + + + TK
Sbjct: 261 SFKKMVVPFWYSNYLQEKTLLKQHKYSQKRNLRKPPKNTTK 383
>AL383990
Length = 465
Score = 31.2 bits (69), Expect = 1.3
Identities = 15/37 (40%), Positives = 24/37 (64%), Gaps = 2/37 (5%)
Frame = -3
Query: 101 IYRSLKTFQNLIRK--LNLKKVQKHNPLQNLNEPQKH 135
+Y+ L FQ+++ + LN +K H+PL NL E +KH
Sbjct: 427 VYQGLYVFQDVLLQ**LNQQKKYNHHPLNNLYELKKH 317
>BG452576 weakly similar to GP|12005223|gb|A reverse transcriptase-like
protein {Spiranthes hongkongensis}, partial (25%)
Length = 676
Score = 30.0 bits (66), Expect = 2.9
Identities = 23/92 (25%), Positives = 45/92 (48%)
Frame = +2
Query: 298 LIKNDFKRGQVDTTLFRGTLKKYILVVQIYVDDIIFGSTNASLCKEFSK*MQDEFEKSMM 357
LI +K+ D ++F + + + +Y+DDI+F + S + + F +
Sbjct: 266 LISLGYKQSPNDHSIF--SFGRRFTIFLVYLDDIVFARDDHSETQLVKSHLDKNFIIIGL 439
Query: 358 GELKFFLGIQINQRKDGVYVHQTKYTKELLKK 389
G L + +G++I + + Q KYT ELL++
Sbjct: 440 GTLHY-VGVKIA*SES*IIDDQCKYTIELLEE 532
>TC91257 similar to GP|21434|emb|CAA36616.1|| ORF4 {Solanum tuberosum},
partial (8%)
Length = 854
Score = 30.0 bits (66), Expect = 2.9
Identities = 13/32 (40%), Positives = 21/32 (65%)
Frame = +3
Query: 418 DQKLYIGMIGSLLYLTASRPDILFSVCLCARF 449
D Y ++G L YLT +RPDI ++V + ++F
Sbjct: 750 DPGRYRRLVGKLNYLTMTRPDISYAVSVVSQF 845
>BG449787 similar to SP|Q9SZR0|CHMO Probable choline monooxygenase
chloroplast precursor (EC 1.14.15.7). [Mouse-ear cress],
partial (32%)
Length = 663
Score = 28.9 bits (63), Expect = 6.4
Identities = 17/52 (32%), Positives = 26/52 (49%), Gaps = 7/52 (13%)
Frame = +3
Query: 112 IRKLNLKKVQKHNPLQNLNEPQKHKMKMLLKKLTMT-------LSK*FNPKV 156
+ N+ +QK P QNLN P KH K + L+ + L++ FNP +
Sbjct: 12 VTPFNIPNLQKRQP-QNLNFPNKHSSKPICCSLSSSDMVVSQKLAQQFNPNI 164
>TC82417 similar to GP|15293221|gb|AAK93721.1 unknown protein {Arabidopsis
thaliana}, partial (29%)
Length = 1151
Score = 28.5 bits (62), Expect = 8.3
Identities = 15/48 (31%), Positives = 26/48 (53%), Gaps = 1/48 (2%)
Frame = -2
Query: 104 SLKTFQNLIRKLNLKKVQKHNPLQNLNEPQK-HKMKMLLKKLTMTLSK 150
++ TFQ + R+ N ++ + P QNL QK + + K+T +SK
Sbjct: 298 TITTFQQIPRRWNCFRIHRSKPTQNLPTIQKLQPIASIFNKMTNQVSK 155
>TC86272 similar to GP|4204761|gb|AAD11482.1| peroxidase precursor {Glycine
max}, partial (86%)
Length = 1350
Score = 28.5 bits (62), Expect = 8.3
Identities = 10/26 (38%), Positives = 17/26 (64%)
Frame = +3
Query: 569 YYCYLFVKESNSTFKSQAYINQTPLY 594
Y+CY+F+ + N++F S +Y LY
Sbjct: 84 YHCYVFINKRNTSFSSISYSCSCQLY 161
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.352 0.155 0.517
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 19,709,005
Number of Sequences: 36976
Number of extensions: 280334
Number of successful extensions: 2346
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 1325
Number of HSP's successfully gapped in prelim test: 116
Number of HSP's that attempted gapping in prelim test: 991
Number of HSP's gapped (non-prelim): 1485
length of query: 642
length of database: 9,014,727
effective HSP length: 102
effective length of query: 540
effective length of database: 5,243,175
effective search space: 2831314500
effective search space used: 2831314500
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (22.0 bits)
S2: 61 (28.1 bits)
Medicago: description of AC146818.6


