

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
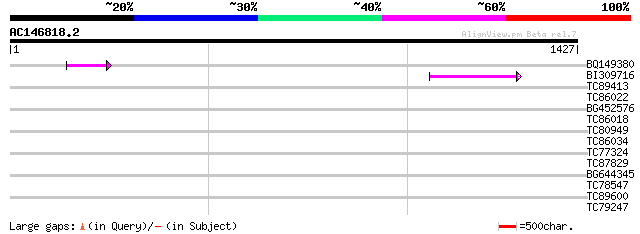
Query= AC146818.2 - phase: 0 /pseudo
(1427 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BQ149380 60 5e-09
BI309716 weakly similar to PIR|G96722|G9 hypothetical protein F2... 47 5e-05
TC89413 weakly similar to GP|8778473|gb|AAF79481.1| F1L3.23 {Ara... 32 1.4
TC86022 homologue to GP|19423939|gb|AAL87312.1 putative RNA heli... 32 2.3
BG452576 weakly similar to GP|12005223|gb|A reverse transcriptas... 32 2.3
TC86018 similar to GP|19423939|gb|AAL87312.1 putative RNA helica... 31 3.0
TC80949 similar to PIR|T52422|T52422 alternative oxidase-related... 31 3.9
TC86034 similar to GP|6815067|dbj|BAA90354.1 ESTs AU082210(C5365... 31 3.9
TC77324 homologue to GP|537313|gb|AAB41813.1|| unknown protein {... 30 5.1
TC87829 similar to GP|21592510|gb|AAM64460.1 ATP-binding protein... 30 5.1
BG644345 weakly similar to GP|19697333|gb putative protein poten... 30 6.7
TC78547 similar to GP|12003386|gb|AAG43550.1 Avr9/Cf-9 rapidly e... 30 6.7
TC89600 similar to GP|16323167|gb|AAL15318.1 At1g30760/T5I8_22 {... 30 8.8
TC79247 weakly similar to GP|15294288|gb|AAK95321.1 AT5g60490/mu... 30 8.8
>BQ149380
Length = 419
Score = 60.5 bits (145), Expect = 5e-09
Identities = 46/114 (40%), Positives = 60/114 (52%), Gaps = 1/114 (0%)
Frame = +2
Query: 144 EGSMGRIGAI*ANASV*LSYYLYLCCYEVC*RV*IGR*NYTVLNWIE*GISRSGFSGFTY 203
E +G IG+I A ASV LS +YL CYE R*NY V N IE* +SR
Sbjct: 74 ERFVG*IGSIQA*ASVYLSNSMYLFCYEK*QSSYSRR*NYQVFNGIE**LSRGDIKSVVD 253
Query: 204 GSNATNQ*GFFNGYATGKENAIWCYQCSEYTIRGYYKY-CKCSEWSKTVWQR*R 256
S +N+ GF G A K++A+ C Y+ R Y + C C WS+ +W+ *R
Sbjct: 254 VSFTSNKHGFLYGNAARKKDAVKC-SIFTYSYR*YNLWSC*CHGWSEAIWKG*R 412
>BI309716 weakly similar to PIR|G96722|G9 hypothetical protein F20P5.25
[imported] - Arabidopsis thaliana, partial (10%)
Length = 744
Score = 47.0 bits (110), Expect = 5e-05
Identities = 65/232 (28%), Positives = 103/232 (44%)
Frame = +3
Query: 1056 KFASSRNPFMVLSKQVGNGLRSLLSFFMLRDSLKLMQITHYLLKSLLLPTLWFWFMWMTL 1115
++ S +N +M S+QV NG+ S L+ + I +L +L L +M MTL
Sbjct: 18 RYVSFKNLYMASSRQVDNGILSYLNLSFPLVIYNPLLIFPFLQNLKILHLLHC*YMLMTL 197
Query: 1116 F*QEHVCRRLIH*NKLLTMLFASRI*GNLNSS*DWK*HVPQKAFLSVNANTVWSFLMMLA 1175
F* E + + N L ++ S+ + * + KAF + N + +F ++
Sbjct: 198 F*LEMIYLKSNMLNVFLLIVLKSKTLVPYDIF*VLRLLEANKAFYLIKENILLNF*RIVV 377
Query: 1176 *LVANQFLLLLILLYVYLKILGLFMVMSLVIDV*LEDCYISPPHALI*LLQHNN*VNSWL 1235
L+ N LLL+I L ++ F +M L I+ + +I LI L NN* N +
Sbjct: 378 ILL*NPLLLLMIFL*NSTILIRPFTMMKLNIEDS*ANLFI*LLLVLIYPLLFNN*ANLFR 557
Query: 1236 LQLNCITKQLLEF*DI*SALLDVVYFSLNPLSFNCLVSVMQTGEGV*TLVVP 1287
+ LLEF +I* L Y + L N LV + G+ LV+P
Sbjct: 558 NLSKFTIRLLLEFYNI*KLPLPKDYSTQQLLI*NFLVLQILIGQ----LVLP 701
>TC89413 weakly similar to GP|8778473|gb|AAF79481.1| F1L3.23 {Arabidopsis
thaliana}, partial (47%)
Length = 749
Score = 32.3 bits (72), Expect = 1.4
Identities = 25/66 (37%), Positives = 38/66 (56%), Gaps = 6/66 (9%)
Frame = +2
Query: 800 LIWNFLSIP---LVLLQFPLHIFTLIVKLLTLCLLMTLL*ILLKIMMRFLMKIHLLILKK 856
++ FLS P L+LL L + L++KLL LLM L LL +++ L K++ ++LK
Sbjct: 236 ILVKFLSFPSANLILLLLFLLLVLLLIKLLLFLLLMLLTMFLLLLLL--LFKLNKMLLKT 409
Query: 857 ---GMM 859
GMM
Sbjct: 410 THLGMM 427
>TC86022 homologue to GP|19423939|gb|AAL87312.1 putative RNA helicase
{Arabidopsis thaliana}, partial (89%)
Length = 1982
Score = 31.6 bits (70), Expect = 2.3
Identities = 16/46 (34%), Positives = 30/46 (64%)
Frame = -2
Query: 821 LIVKLLTLCLLMTLL*ILLKIMMRFLMKIHLLILKKGMMKWSL*PP 866
L++ LL L L++ L+ I+L +++ + +H+L+L +M WSL P
Sbjct: 313 LLLLLLILLLILLLILIVLLMLILHHVPLHILLLLLILMMWSLLKP 176
>BG452576 weakly similar to GP|12005223|gb|A reverse transcriptase-like protein
{Spiranthes hongkongensis}, partial (25%)
Length = 676
Score = 31.6 bits (70), Expect = 2.3
Identities = 17/45 (37%), Positives = 24/45 (52%)
Frame = +3
Query: 1072 GNGLRSLLSFFMLRDSLKLMQITHYLLKSLLLPTLWFWFMWMTLF 1116
GN + S L D L IT +L+ L + L+FW++WM LF
Sbjct: 237 GNDIESCYQL*FL*DINNLQMITPFLV--LAVGLLFFWYIWMILF 365
>TC86018 similar to GP|19423939|gb|AAL87312.1 putative RNA helicase
{Arabidopsis thaliana}, partial (12%)
Length = 885
Score = 31.2 bits (69), Expect = 3.0
Identities = 19/59 (32%), Positives = 36/59 (60%)
Frame = -1
Query: 808 PLVLLQFPLHIFTLIVKLLTLCLLMTLL*ILLKIMMRFLMKIHLLILKKGMMKWSL*PP 866
PL+LL + +++ +L L LLM L+ +L+ I++ + +H+L+L ++ WSL P
Sbjct: 702 PLLLL-----LLLILLLILLLILLMMLIVLLMLILILHQVPLHILLL---LIMWSLLKP 550
>TC80949 similar to PIR|T52422|T52422 alternative oxidase-related protein
IMMUTANS [validated] - Arabidopsis thaliana, partial
(69%)
Length = 1369
Score = 30.8 bits (68), Expect = 3.9
Identities = 26/62 (41%), Positives = 33/62 (52%), Gaps = 11/62 (17%)
Frame = +3
Query: 810 VLLQFPLHIFTLI--------VKLL--TLCLLMTLL*ILLKIMMRFLMKIHLLIL-KKGM 858
VL F LH + +KLL TL LL LL*I L + M +HLL+L KKG+
Sbjct: 144 VLFHFVLHFSEFVRLCYKIKKIKLLHKTLFLLKPLL*IPLLKIQLMTMILHLLVLGKKGL 323
Query: 859 MK 860
+K
Sbjct: 324 LK 329
>TC86034 similar to GP|6815067|dbj|BAA90354.1 ESTs AU082210(C53655)
AU056546(S20671) AU056545(S20671) correspond to a
region of the predicted, partial (42%)
Length = 2412
Score = 30.8 bits (68), Expect = 3.9
Identities = 13/48 (27%), Positives = 32/48 (66%)
Frame = -1
Query: 805 LSIPLVLLQFPLHIFTLIVKLLTLCLLMTLL*ILLKIMMRFLMKIHLL 852
L +PL+L+ LH+ + L ++C L+ +L +LL++++ ++++ L+
Sbjct: 288 LLLPLLLVLLHLHLRFEVDLLQSVCFLLLVLLVLLRLLLLVVVRVQLV 145
>TC77324 homologue to GP|537313|gb|AAB41813.1|| unknown protein {Medicago
sativa}, complete
Length = 1698
Score = 30.4 bits (67), Expect = 5.1
Identities = 16/38 (42%), Positives = 24/38 (63%)
Frame = +3
Query: 798 NFLIWNFLSIPLVLLQFPLHIFTLIVKLLTLCLLMTLL 835
N ++W +L I LVL + L + LI+ LL L +L+ LL
Sbjct: 555 NMMLWKWLFIILVLKKILLVVLLLILLLLGLSILLKLL 668
>TC87829 similar to GP|21592510|gb|AAM64460.1 ATP-binding protein-like
protein {Arabidopsis thaliana}, partial (98%)
Length = 1127
Score = 30.4 bits (67), Expect = 5.1
Identities = 14/46 (30%), Positives = 26/46 (56%)
Frame = +2
Query: 807 IPLVLLQFPLHIFTLIVKLLTLCLLMTLL*ILLKIMMRFLMKIHLL 852
+ VL+Q+PL I+ + CLL+T L+ + RF +K++ +
Sbjct: 887 VEFVLVQYPLGFLACIIAVTFSCLLITYKGFLMYYIPRFTIKMYCI 1024
>BG644345 weakly similar to GP|19697333|gb putative protein potential
transcriptional repressor Not4hp - Mus musculus, partial
(7%)
Length = 635
Score = 30.0 bits (66), Expect = 6.7
Identities = 20/49 (40%), Positives = 30/49 (60%)
Frame = -3
Query: 809 LVLLQFPLHIFTLIVKLLTLCLLMTLL*ILLKIMMRFLMKIHLLILKKG 857
L+LLQ L + ++ LL L LL+ LL +LLK+ + LL+L+KG
Sbjct: 432 LLLLQLQLLLLLKLLLLLLLKLLLLLLLLLLKLKL-------LLLLRKG 307
>TC78547 similar to GP|12003386|gb|AAG43550.1 Avr9/Cf-9 rapidly elicited
protein 132 {Nicotiana tabacum}, partial (41%)
Length = 1648
Score = 30.0 bits (66), Expect = 6.7
Identities = 49/194 (25%), Positives = 88/194 (45%), Gaps = 8/194 (4%)
Frame = +3
Query: 818 IFTLIVKLLTLCLLMTLL*ILLKIMMRFLMKIHLLILKKGMMKWSL*PPENPPEVLNHQF 877
+F L+VK+L LC+ +T L + LK + L + + K M W N F
Sbjct: 501 LFLLLVKIL-LCMEVTKLDLKLKFLSLCLFWFSRMRISK--MVW------------NVLF 635
Query: 878 TY------KIMSVIQQVVITL*QILLVIQLYLLNTVLMLTL*IL--RLSPLVIFQLVKIL 929
Y K + Q ++ I+L+ L+ VL + +* L ++ +V+ Q+VK
Sbjct: 636 VYVMLLKEKKLGFYQNAIMGFI*IVLICGFNLILLVLFVGI*FLLNHVNQIVLLQMVKR* 815
Query: 930 GG*LLCRQR*RLLMQIILGNLLIFLQMLYP*VVNGYIRSKDMQMVP*NVSKHVWLLKVLI 989
* C Q+ ++ + ++ +L IFLQM + G ++ K + ++ V+L K +
Sbjct: 816 MC*--CHQKVKI*VM*MV*SLQIFLQMFW---FGGILKDKLVLLL-------VFLWKKEV 959
Query: 990 KLKGLIILKHSPLL 1003
+ L+ H LL
Sbjct: 960 LINNLVQHHHHHLL 1001
>TC89600 similar to GP|16323167|gb|AAL15318.1 At1g30760/T5I8_22 {Arabidopsis
thaliana}, partial (29%)
Length = 700
Score = 29.6 bits (65), Expect = 8.8
Identities = 15/27 (55%), Positives = 16/27 (58%), Gaps = 4/27 (14%)
Frame = +1
Query: 281 SGNLLQEAWIPSKLGKRW----RKFLC 303
S NLLQE W PSK K W R F+C
Sbjct: 292 SRNLLQETWNPSKSKKWWP*L*RTFIC 372
>TC79247 weakly similar to GP|15294288|gb|AAK95321.1 AT5g60490/muf9_140
{Arabidopsis thaliana}, partial (47%)
Length = 864
Score = 29.6 bits (65), Expect = 8.8
Identities = 17/62 (27%), Positives = 35/62 (56%)
Frame = -2
Query: 800 LIWNFLSIPLVLLQFPLHIFTLIVKLLTLCLLMTLL*ILLKIMMRFLMKIHLLILKKGMM 859
L+W + LVLL+ L++ LL L L+M L+ +L+ ++ +M + + +++ G+
Sbjct: 326 LLWWVMVHQLVLLEQVELCLILVMALLLLLLVMVLMLLLMVRVVMEMMLLGIELVRVGLK 147
Query: 860 KW 861
W
Sbjct: 146 WW 141
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.370 0.167 0.644
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 53,155,779
Number of Sequences: 36976
Number of extensions: 900887
Number of successful extensions: 13915
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 3204
Number of HSP's successfully gapped in prelim test: 647
Number of HSP's that attempted gapping in prelim test: 10304
Number of HSP's gapped (non-prelim): 4780
length of query: 1427
length of database: 9,014,727
effective HSP length: 108
effective length of query: 1319
effective length of database: 5,021,319
effective search space: 6623119761
effective search space used: 6623119761
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.8 bits)
S2: 65 (29.6 bits)
Medicago: description of AC146818.2


