

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146817.2 + phase: 0
(145 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
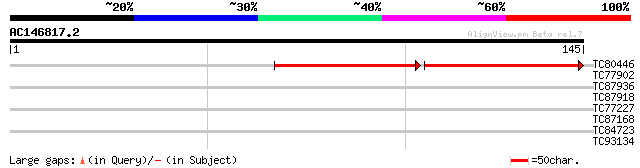
TC80446 similar to GP|21740740|emb|CAD40549. OSJNBa0072K14.5 {Or... 75 9e-15
TC77902 similar to PIR|E96789|E96789 protein T23E18.10 [imported... 28 0.99
TC87936 similar to GP|15010674|gb|AAK73996.1 AT3g16810/K20I9_3 {... 27 3.8
TC87918 weakly similar to GP|15810397|gb|AAL07086.1 unknown prot... 26 6.4
TC77227 similar to SP|Q9LDU6|ST7R_ARATH 7-dehydrocholesterol red... 26 6.4
TC87168 similar to PIR|T47595|T47595 RING finger protein T12E18.... 26 6.4
TC84723 similar to GP|16945387|emb|CAD11800. conserved hypotheti... 25 8.4
TC93134 similar to GP|5669638|gb|AAD46404.1| ethylene-responsive... 25 8.4
>TC80446 similar to GP|21740740|emb|CAD40549. OSJNBa0072K14.5 {Oryza
sativa}, partial (11%)
Length = 798
Score = 75.1 bits (183), Expect = 9e-15
Identities = 31/40 (77%), Positives = 35/40 (87%)
Frame = +2
Query: 106 KQGCPHIAKLGKGRRIKPLKHSNFNERNCRDKFRQDPCRF 145
KQGCPHIAKL +G RI+P KHS+FN NCRDKFRQ+PCRF
Sbjct: 506 KQGCPHIAKLSRGGRIRPFKHSSFNH*NCRDKFRQEPCRF 625
Score = 60.1 bits (144), Expect = 3e-10
Identities = 29/37 (78%), Positives = 33/37 (88%)
Frame = +3
Query: 68 MIIQPKEKGKSNSGKGEHITSRPLQAAHDEYEEEATL 104
+I Q KEKGKS SGKGE+ITS+ L+AAHDEYEEEATL
Sbjct: 117 VIAQQKEKGKSKSGKGENITSQHLKAAHDEYEEEATL 227
>TC77902 similar to PIR|E96789|E96789 protein T23E18.10 [imported] -
Arabidopsis thaliana, partial (95%)
Length = 2016
Score = 28.5 bits (62), Expect = 0.99
Identities = 12/22 (54%), Positives = 13/22 (58%)
Frame = +3
Query: 117 KGRRIKPLKHSNFNERNCRDKF 138
K RI PL+H NE NC KF
Sbjct: 1173 KSARIIPLRHDKLNEDNCTIKF 1238
>TC87936 similar to GP|15010674|gb|AAK73996.1 AT3g16810/K20I9_3 {Arabidopsis
thaliana}, partial (36%)
Length = 982
Score = 26.6 bits (57), Expect = 3.8
Identities = 17/45 (37%), Positives = 21/45 (45%), Gaps = 2/45 (4%)
Frame = +1
Query: 73 KEKGKSNSGKGEHITSRPLQAA--HDEYEEEATLAKQGCPHIAKL 115
K KGK + G HITSR LQ H E + ++ PH L
Sbjct: 391 KMKGKIHEIAGSHITSRVLQTCVKHCSQAERDQVFEELRPHFLNL 525
>TC87918 weakly similar to GP|15810397|gb|AAL07086.1 unknown protein
{Arabidopsis thaliana}, partial (67%)
Length = 1746
Score = 25.8 bits (55), Expect = 6.4
Identities = 17/71 (23%), Positives = 36/71 (49%)
Frame = +2
Query: 36 NNQIFEDSNSLKVKEKDNMKHHCDEKREVYEYMIIQPKEKGKSNSGKGEHITSRPLQAAH 95
NN + E + + +EK+ K+ E++E + +EKG+ N+G GE + ++
Sbjct: 929 NNGVNEGKEA-EGEEKEQEKNGVKEEKET------EVEEKGQENNGAGEGKVTEGVEKGQ 1087
Query: 96 DEYEEEATLAK 106
+ ++ A + K
Sbjct: 1088ETGKKTAVIFK 1120
>TC77227 similar to SP|Q9LDU6|ST7R_ARATH 7-dehydrocholesterol reductase (EC
1.3.1.21) (7-DHC reductase) (Sterol delta-7-reductase),
partial (71%)
Length = 1248
Score = 25.8 bits (55), Expect = 6.4
Identities = 11/27 (40%), Positives = 18/27 (65%)
Frame = -3
Query: 48 VKEKDNMKHHCDEKREVYEYMIIQPKE 74
+K+KDN K+H + ++EV E P+E
Sbjct: 928 IKKKDNKKYHVEIRQEVIEKSWDCPEE 848
>TC87168 similar to PIR|T47595|T47595 RING finger protein T12E18.50 -
Arabidopsis thaliana, partial (73%)
Length = 1723
Score = 25.8 bits (55), Expect = 6.4
Identities = 14/49 (28%), Positives = 26/49 (52%), Gaps = 4/49 (8%)
Frame = +2
Query: 29 IRVPRGVNNQIFEDSNSLKVKEKDNMKHHCDE----KREVYEYMIIQPK 73
+++PR +DSN+ K +++ KHH + K+EV + + PK
Sbjct: 152 VKIPRP------DDSNASKKSNENSNKHHVEHDSKVKKEVKDSASVSPK 280
>TC84723 similar to GP|16945387|emb|CAD11800. conserved hypothetical protein
{Neurospora crassa}, partial (5%)
Length = 999
Score = 25.4 bits (54), Expect = 8.4
Identities = 20/95 (21%), Positives = 34/95 (35%), Gaps = 16/95 (16%)
Frame = +3
Query: 28 PIRVPRGVNNQIFEDSNSLKVKEKDNMKHHCDEKREVYEYMIIQPKEKGKSNSGKGEHIT 87
P P +NN+I+ D +S + ++ HH + + + S + E+
Sbjct: 324 PAPTPVIINNRIYNDHSSDEEDDRHMQVHHSRRRSHSRGSLYSHSRSPSSSYMTREEYEA 503
Query: 88 SRPLQAAHD----------------EYEEEATLAK 106
R LQ + EY EEA L +
Sbjct: 504 ERALQELRELKISKAREREELRVQKEYREEAELQR 608
>TC93134 similar to GP|5669638|gb|AAD46404.1| ethylene-responsive RNA
helicase {Lycopersicon esculentum}, partial (38%)
Length = 684
Score = 25.4 bits (54), Expect = 8.4
Identities = 8/14 (57%), Positives = 11/14 (78%)
Frame = +2
Query: 130 NERNCRDKFRQDPC 143
+ RNCRD+ R+D C
Sbjct: 437 SNRNCRDRIRKDTC 478
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.315 0.133 0.393
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,520,698
Number of Sequences: 36976
Number of extensions: 53573
Number of successful extensions: 191
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 191
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 191
length of query: 145
length of database: 9,014,727
effective HSP length: 87
effective length of query: 58
effective length of database: 5,797,815
effective search space: 336273270
effective search space used: 336273270
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 53 (25.0 bits)
Medicago: description of AC146817.2


