

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146794.6 + phase: 0
(1018 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
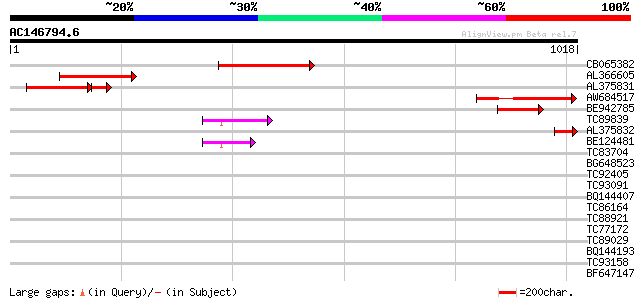
Score E
Sequences producing significant alignments: (bits) Value
CB065382 similar to PIR|T09671|T096 RPE15 protein - alfalfa (fra... 301 8e-82
AL366605 289 4e-78
AL375831 212 3e-71
AW684517 similar to GP|9927273|dbj Similar to Arabidopsis thalia... 250 2e-66
BE942785 157 1e-38
TC89839 homologue to SP|P23466|CYAA_SACKL Adenylate cyclase (EC ... 89 7e-18
AL375832 76 6e-14
BE124481 70 3e-12
TC83704 similar to PIR|T05005|T05005 hypothetical protein T19P19... 39 0.010
BG648523 similar to GP|9955540|emb| putative protein {Arabidopsi... 38 0.017
TC92405 similar to GP|11994325|dbj|BAB02284. gene_id:MLM24.13~un... 38 0.017
TC93091 similar to GP|4115538|dbj|BAA36412.1 UDP-glycose:flavono... 34 0.25
BQ144407 weakly similar to GP|6523547|emb| hydroxyproline-rich g... 34 0.25
TC86164 weakly similar to GP|20514261|gb|AAM22959.1 tubulin fold... 34 0.33
TC88921 similar to SP|P92966|RS41_ARATH Arginine/serine-rich spl... 34 0.33
TC77172 similar to GP|20466472|gb|AAM20553.1 putative protein {A... 34 0.33
TC89029 weakly similar to GP|15450467|gb|AAK96527.1 At1g12810/F1... 33 0.43
BQ144193 weakly similar to GP|14026007|dbj putative transporter ... 33 0.43
TC93158 similar to PIR|T01123|T01123 hypothetical protein At2g32... 33 0.43
BF647147 weakly similar to GP|7303636|gb|A CG13214-PA {Drosophil... 33 0.43
>CB065382 similar to PIR|T09671|T096 RPE15 protein - alfalfa (fragment),
partial (9%)
Length = 624
Score = 301 bits (771), Expect = 8e-82
Identities = 146/172 (84%), Positives = 152/172 (87%)
Frame = -2
Query: 375 PVQQQQQHQQARPTFPPIPMLYAELLPTLLHRGHCTTRQGKPPPDPLPPRFRSDLKCDFH 434
PV QQ+Q QQARPTFP IPMLYAE LPTLL RGHCT RQGKPPPDPLPPRFRSDLKCDFH
Sbjct: 518 PVHQQRQ*QQARPTFPLIPMLYAE*LPTLLLRGHCTIRQGKPPPDPLPPRFRSDLKCDFH 339
Query: 435 QGALGHDVEGCYALKYIVKKLIDQGKLTFENNVPHVLDNPLPNHATVNMIEVCEEALRLD 494
QGALGHDVEGCYALK+IVKKLI+QGKLTFENNVPHVLDNPLPNHA VNMIEV EEA LD
Sbjct: 338 QGALGHDVEGCYALKHIVKKLINQGKLTFENNVPHVLDNPLPNHAAVNMIEVYEEAPGLD 159
Query: 495 VRNVATPLVPLHIKLCKASLFSHDHAKCLGCLRNPLGCFTVQDDIQSLMNDN 546
VRNV TPLVPLHIKLC+ASLF HDHA C NPLGC VQ+DIQSLMN+N
Sbjct: 158 VRNVTTPLVPLHIKLCQASLFDHDHANCQE*FYNPLGCCVVQNDIQSLMNNN 3
>AL366605
Length = 422
Score = 289 bits (739), Expect = 4e-78
Identities = 137/140 (97%), Positives = 138/140 (97%)
Frame = -3
Query: 89 HEIKANQGNADSFKTQDLCLVPKVDVPKKFKIPDFDRYNGLTCPQNHIIKYVRKMGNYKD 148
HEIKAN+GNADSFKTQDLCLV KVDVPKKFKIPDFDRYNGLTCPQNHIIKYVRKMGNYKD
Sbjct: 420 HEIKANRGNADSFKTQDLCLVSKVDVPKKFKIPDFDRYNGLTCPQNHIIKYVRKMGNYKD 241
Query: 149 NDSLMIHCFQDSLMEDATEWYTSLSKNDIHTFDELAAAFKSHYGFNTRLKPNREFLRSLS 208
NDSLMIHCFQDSLMEDA EWYTSLSKNDIHTFDELAAAFKSHYGFNTRLKPNREFLRSLS
Sbjct: 240 NDSLMIHCFQDSLMEDAAEWYTSLSKNDIHTFDELAAAFKSHYGFNTRLKPNREFLRSLS 61
Query: 209 QKKEESFREYAQRWRGAAAR 228
QKKEESFREYAQRWRGAAAR
Sbjct: 60 QKKEESFREYAQRWRGAAAR 1
>AL375831
Length = 467
Score = 212 bits (540), Expect(2) = 3e-71
Identities = 102/119 (85%), Positives = 107/119 (89%)
Frame = +2
Query: 30 IPATNVVSINTTLPQTTAAVTEPLVHAIPQSVNINTHHGNIPVIKTMEERMEELAKELRH 89
IPATN SI TLPQTTAAVTEPLVH +PQ +NINT H +IPV KTMEE MEELAKELRH
Sbjct: 8 IPATNAASIIATLPQTTAAVTEPLVHTLPQGININTQHRSIPVTKTMEEMMEELAKELRH 187
Query: 90 EIKANQGNADSFKTQDLCLVPKVDVPKKFKIPDFDRYNGLTCPQNHIIKYVRKMGNYKD 148
EI+AN+GNADS KTQDLCLV KVDVPKKFKIPDFDRYNGLTCPQNHIIKYVRKMGNYKD
Sbjct: 188 EIQANRGNADSVKTQDLCLVSKVDVPKKFKIPDFDRYNGLTCPQNHIIKYVRKMGNYKD 364
Score = 76.3 bits (186), Expect(2) = 3e-71
Identities = 34/36 (94%), Positives = 34/36 (94%)
Frame = +3
Query: 147 KDNDSLMIHCFQDSLMEDATEWYTSLSKNDIHTFDE 182
K NDSLMIHCFQDSLMEDA EWYTSLSKNDIHTFDE
Sbjct: 360 KTNDSLMIHCFQDSLMEDAAEWYTSLSKNDIHTFDE 467
>AW684517 similar to GP|9927273|dbj Similar to Arabidopsis thaliana chromosome
II BAC F26H6; putative retroelement pol polyprotein,
partial (1%)
Length = 488
Score = 250 bits (638), Expect = 2e-66
Identities = 127/180 (70%), Positives = 139/180 (76%)
Frame = +1
Query: 838 ITFQVMDINASYSCLLGRPWIHDAGAVTSTLHQKLKFAKSGKLVTIHGEEAYLVSQLSSF 897
ITFQVMDINASYSCLLGRPWIHDAGAVTSTLHQKLKF K+G
Sbjct: 1 ITFQVMDINASYSCLLGRPWIHDAGAVTSTLHQKLKFVKNG------------------- 123
Query: 898 SCIEAGSAEGTAFQGLTVEGTEPKRDGTAMASLKDAQRAVQEGQAAGWGRLIQLRENKHK 957
SAEGTAFQGL++EG EPK+ G AMASLKDAQ+AVQEGQAA WG+LIQL ENK K
Sbjct: 124 ------SAEGTAFQGLSMEGAEPKKVGAAMASLKDAQKAVQEGQAADWGKLIQLCENKRK 285
Query: 958 EGLGFSPTSGVSTGAFYSAGFVNAITEEATGFGPRPVFVTPGGIARDWDAIDIPSIMHVS 1017
EGL FSPTSGVSTG F+SAGFVN + EE F PRP+FV PGGIA+DWDA+D+PSIMHVS
Sbjct: 286 EGLRFSPTSGVSTGTFHSAGFVNTLAEEVARFVPRPLFVIPGGIAKDWDAVDVPSIMHVS 465
>BE942785
Length = 460
Score = 157 bits (398), Expect = 1e-38
Identities = 78/82 (95%), Positives = 80/82 (97%)
Frame = -2
Query: 877 SGKLVTIHGEEAYLVSQLSSFSCIEAGSAEGTAFQGLTVEGTEPKRDGTAMASLKDAQRA 936
SG LVTIHGEEAYL+SQLSSFSCIEAGSAEGTAFQGLTVEGTEPKRDGTAMASLKDAQRA
Sbjct: 378 SGXLVTIHGEEAYLISQLSSFSCIEAGSAEGTAFQGLTVEGTEPKRDGTAMASLKDAQRA 199
Query: 937 VQEGQAAGWGRLIQLRENKHKE 958
VQE QAAGWGRLIQLRENKHK+
Sbjct: 198 VQESQAAGWGRLIQLRENKHKD 133
>TC89839 homologue to SP|P23466|CYAA_SACKL Adenylate cyclase (EC 4.6.1.1)
(ATP pyrophosphate-lyase) (Adenylyl cyclase). [Yeast],
partial (0%)
Length = 665
Score = 89.4 bits (220), Expect = 7e-18
Identities = 55/133 (41%), Positives = 68/133 (50%), Gaps = 7/133 (5%)
Frame = +3
Query: 346 PPFYHQYPLPPGQPQVPVNAIAQQMKQQLPVQQ-------QQQHQQARPTFPPIPMLYAE 398
PP Q P P QP P Q +Q +P Q Q Q Q+A PIP+ YA+
Sbjct: 228 PPLVQQRPQSPYQPMCPQKPRQQAPRQSIPQNQVPQKFIPQNQVQKASQC-DPIPVKYAD 404
Query: 399 LLPTLLHRGHCTTRQGKPPPDPLPPRFRSDLKCDFHQGALGHDVEGCYALKYIVKKLIDQ 458
LLP LL + T P+ LPP +R DL C FHQGA GHD E CY LK V+KLI+
Sbjct: 405 LLPILLKKNLIQTLPLPRVPNSLPPWYRPDLNCVFHQGAPGHDTEQCYPLKEEVQKLIEN 584
Query: 459 GKLTFENNVPHVL 471
+F++ VL
Sbjct: 585 NVWSFDDQDIKVL 623
>AL375832
Length = 535
Score = 76.3 bits (186), Expect = 6e-14
Identities = 36/41 (87%), Positives = 37/41 (89%)
Frame = +3
Query: 978 FVNAITEEATGFGPRPVFVTPGGIARDWDAIDIPSIMHVSE 1018
FVNAI+EEATG G RP FVTPGGIA DWDAIDIPSIMHVSE
Sbjct: 3 FVNAISEEATGSGLRPAFVTPGGIASDWDAIDIPSIMHVSE 125
>BE124481
Length = 538
Score = 70.5 bits (171), Expect = 3e-12
Identities = 43/103 (41%), Positives = 51/103 (48%), Gaps = 7/103 (6%)
Frame = +3
Query: 346 PPFYHQYPLPPGQPQVPVNAIAQQMKQQLPVQQ-------QQQHQQARPTFPPIPMLYAE 398
PP Q P P QP P Q +Q +P Q Q Q Q+A PIP+ YA+
Sbjct: 231 PPLVQQRPQSPYQPMCPQKPRQQAPRQSIPQNQVPQKFIPQNQVQKASQC-DPIPVKYAD 407
Query: 399 LLPTLLHRGHCTTRQGKPPPDPLPPRFRSDLKCDFHQGALGHD 441
LLP LL + T P+ LPP +R DL C FHQGA GHD
Sbjct: 408 LLPILLKKNLIQTLPLPRVPNSLPPWYRPDLNCVFHQGAPGHD 536
>TC83704 similar to PIR|T05005|T05005 hypothetical protein T19P19.70 -
Arabidopsis thaliana, partial (15%)
Length = 716
Score = 38.9 bits (89), Expect = 0.010
Identities = 32/111 (28%), Positives = 46/111 (40%), Gaps = 6/111 (5%)
Frame = +1
Query: 320 ATVAPINAAQMPPSYPYAPYSQHPFFPPFYHQYPLPPGQPQVPVNAIAQQMKQQLPVQQQ 379
+TV P+ +P PY QH PP P PP ++P A +Q+ P+ +
Sbjct: 262 STVPPVATTSLPEPLPY----QHKEQPPIPVTLPPPPPLSKLPP-ATREQLPPPPPLSKL 426
Query: 380 QQHQQARPTFPP----IPMLYAELLPTL--LHRGHCTTRQGKPPPDPLPPR 424
+ P PP +P E LP L + R+ PPP PLP +
Sbjct: 427 PPAAREHPPPPPPSAKLPPAARERLPLPPPLSKLPPAARERLPPPSPLPEK 579
>BG648523 similar to GP|9955540|emb| putative protein {Arabidopsis thaliana},
partial (9%)
Length = 770
Score = 38.1 bits (87), Expect = 0.017
Identities = 30/94 (31%), Positives = 38/94 (39%), Gaps = 2/94 (2%)
Frame = +3
Query: 332 PSYPYAPYSQHPFFPPFYHQYPLPPGQPQVPVNAIAQQMKQQLPVQQQQQHQQARPTFPP 391
PS+ + Q P PLPPG +N QQ QQ P Q Q QQ + P
Sbjct: 348 PSFDGSQGEQKQPHPGSIGAPPLPPGPHPSLLNPNQQQPFQQNPQQIPQHQQQLQQHMGP 527
Query: 392 IPMLYAELLPTLLHRGHCT--TRQGKPPPDPLPP 423
+PM +P + H H + Q P P P P
Sbjct: 528 LPM--PPNMPQIQHSSHSSMLPHQHLPRPPPQMP 623
>TC92405 similar to GP|11994325|dbj|BAB02284. gene_id:MLM24.13~unknown
protein {Arabidopsis thaliana}, partial (69%)
Length = 864
Score = 38.1 bits (87), Expect = 0.017
Identities = 44/156 (28%), Positives = 64/156 (40%), Gaps = 7/156 (4%)
Frame = +2
Query: 735 SIVGNITACSNLWFSEDELP------EAGKHHNLALHISVNCKSDMLSNVLVDTGSSLNV 788
SI G+I SN + S L +G + LA K+D + L D G +++
Sbjct: 152 SISGSIPIPSNPFDSTSPLKFNLLSAVSGLNRGLAASEEDLQKADAAAKELEDAGGLVDL 331
Query: 789 MPKSTLDQLSYRGTPLRRSTFLVKAFDGTRKSV-LGEIDLPITIGPETFLITFQVMDINA 847
LD+L R + S F + G+R +G + LPIT+G I D +
Sbjct: 332 T--DNLDRLQGRWKLIYSSAFSSRTLGGSRPGPPIGRL-LPITLGQVFQRIDILSKDFDN 502
Query: 848 SYSCLLGRPWIHDAGAVTSTLHQKLKFAKSGKLVTI 883
LG PW VT+TL K + S K+ I
Sbjct: 503 IVDLQLGAPWPLPPLEVTATLAHKFELVGSSKIKII 610
>TC93091 similar to GP|4115538|dbj|BAA36412.1 UDP-glycose:flavonoid
glycosyltransferase {Vigna mungo}, partial (37%)
Length = 845
Score = 34.3 bits (77), Expect = 0.25
Identities = 24/94 (25%), Positives = 38/94 (39%), Gaps = 12/94 (12%)
Frame = -3
Query: 318 SMATVAPINAAQMPPSYPYAPYSQHPFFPPFYHQY-----------PLPPGQPQVPVNAI 366
+++ +P+ P P+AP Q P +PP H Y L QP +P +
Sbjct: 486 TLSPSSPLPLLSQTPHLPFAPPQQPPSYPPPSHSY*HAALHAMAMQSLALQQPPLPTPSS 307
Query: 367 AQQMKQQLPVQQQQQHQQARPT-FPPIPMLYAEL 399
+ Q + +H+ PT PP P L+ L
Sbjct: 306 SHSDSQTHQPIRDSEHRLVGPTSSPPGPSLWFSL 205
>BQ144407 weakly similar to GP|6523547|emb| hydroxyproline-rich glycoprotein
DZ-HRGP {Volvox carteri f. nagariensis}, partial (23%)
Length = 1217
Score = 34.3 bits (77), Expect = 0.25
Identities = 30/108 (27%), Positives = 44/108 (39%), Gaps = 7/108 (6%)
Frame = -2
Query: 324 PINAAQMPPSYPYAPYSQHPFFPP---FYHQYPLPPGQPQV-PVNAIAQQMKQQLPVQQQ 379
PI+ + +P YP +P FPP +PL P P ++ +LP+
Sbjct: 601 PIHPSVLPHPYPRLTTPSYPPFPPPLVVLSPFPLTPSPTLASPFQHPPPHLQFRLPILFT 422
Query: 380 QQHQQARPTFPPIPMLYAELLPTLLHRGHCTTRQGKPPPDP---LPPR 424
+ + PT PP P ++ LLP + PPP P PPR
Sbjct: 421 R*TALSSPTPPPRPAAHS-LLPPPFRCPPASCPLSAPPPLPPSAFPPR 281
>TC86164 weakly similar to GP|20514261|gb|AAM22959.1 tubulin folding cofactor
C {Arabidopsis thaliana}, partial (8%)
Length = 1369
Score = 33.9 bits (76), Expect = 0.33
Identities = 29/102 (28%), Positives = 34/102 (32%)
Frame = -1
Query: 323 APINAAQMPPSYPYAPYSQHPFFPPFYHQYPLPPGQPQVPVNAIAQQMKQQLPVQQQQQH 382
API P YAP H PP QYP PG P +
Sbjct: 1000 APIRQRYCPSRP*YAPVPPHSRPPPSSRQYPSAPG-----------SQTTSSPTARATCP 854
Query: 383 QQARPTFPPIPMLYAELLPTLLHRGHCTTRQGKPPPDPLPPR 424
Q +R + P P + T G C TR G+ P P R
Sbjct: 853 QYSRASPPASPTTWR---ATPTSAG*CATRAGRKCKSPWPSR 737
>TC88921 similar to SP|P92966|RS41_ARATH Arginine/serine-rich splicing
factor RSP41. [Mouse-ear cress] {Arabidopsis thaliana},
partial (33%)
Length = 1419
Score = 33.9 bits (76), Expect = 0.33
Identities = 20/57 (35%), Positives = 29/57 (50%), Gaps = 8/57 (14%)
Frame = -1
Query: 10 PPFFTPSTAAGTSGTANNGPIPATNV-----VSINTTLPQ---TTAAVTEPLVHAIP 58
PP F TAAG + + + P+P+ +S T+LP +T AV P+V A P
Sbjct: 615 PPLFMIRTAAGVANSLSTIPLPSVRTRVSTPISRTTSLPSMGASTTAVATPIVRAAP 445
>TC77172 similar to GP|20466472|gb|AAM20553.1 putative protein {Arabidopsis
thaliana}, partial (78%)
Length = 4130
Score = 33.9 bits (76), Expect = 0.33
Identities = 38/177 (21%), Positives = 56/177 (31%), Gaps = 32/177 (18%)
Frame = +1
Query: 264 EMVTMGTRLEEAVREGIIVFEKAESSVNASKRYGNGHHKKKETEVGMVSAGAGQSMATVA 323
+ + + T E+ ++ FE ++S + YG + G VS +
Sbjct: 2431 DRIALSTETEKDLKT--TAFENSQS--HGGSFYGADNSNYVNHYQGSVSTHVPPGVHVPP 2598
Query: 324 PINAAQMPPSY----------------PYAPYSQHPFFPPFYHQYPLPPG---------- 357
++ Q P SY P+ P Q F P P PP
Sbjct: 2599 GVSGGQYPDSYQPPYDNRYVPGYGAPVPHQPPQQPNIFVPSQTTQPQPPQSNFPNTSGAQ 2778
Query: 358 ------QPQVPVNAIAQQMKQQLPVQQQQQHQQARPTFPPIPMLYAELLPTLLHRGH 408
+PQ P + QQ + Q + PTFPP Y P H GH
Sbjct: 2779 PPVRVFEPQTPALIRNPEQYQQPTLGSQLYNTNNNPTFPPTNQPYQPTPPAPSHIGH 2949
>TC89029 weakly similar to GP|15450467|gb|AAK96527.1 At1g12810/F13K23_4
{Arabidopsis thaliana}, partial (71%)
Length = 643
Score = 33.5 bits (75), Expect = 0.43
Identities = 33/122 (27%), Positives = 38/122 (31%), Gaps = 14/122 (11%)
Frame = +3
Query: 338 PYSQHPFFPPFYHQYPLPPGQPQVPVNAIAQQMKQQLPVQQQQQHQQARPT------FPP 391
P S +P P F YP PPG P QH + P FPP
Sbjct: 51 PMSNYPP-PGFGSPYPPPPGPPP-------------------HQHHEGYPPPGYPGGFPP 170
Query: 392 IPMLYAELLPTLLHRGHCTTRQ--------GKPPPDPLPPRFRSDLKCDFHQGALGHDVE 443
P P H H G PPP P PP++ + D H H
Sbjct: 171 PPP------PPHHHHHHGPPHDSYQGYFDNGYPPPPPAPPQYHNYQHVDHHHHHGHHGDP 332
Query: 444 GC 445
GC
Sbjct: 333 GC 338
>BQ144193 weakly similar to GP|14026007|dbj putative transporter
{Mesorhizobium loti}, partial (4%)
Length = 1037
Score = 33.5 bits (75), Expect = 0.43
Identities = 21/67 (31%), Positives = 28/67 (41%), Gaps = 1/67 (1%)
Frame = +2
Query: 293 SKRYGNGHHKK-KETEVGMVSAGAGQSMATVAPINAAQMPPSYPYAPYSQHPFFPPFYHQ 351
+K+ NG K ET G + +S + +MPP YP HP PP H
Sbjct: 47 TKKRANGKQAKINETNAGKLKV---KSNTPAYLVGYKEMPPVYPRKSPKTHPRPPPNSHN 217
Query: 352 YPLPPGQ 358
P PP +
Sbjct: 218 RPEPPAK 238
>TC93158 similar to PIR|T01123|T01123 hypothetical protein At2g32840
[imported] - Arabidopsis thaliana, partial (9%)
Length = 673
Score = 33.5 bits (75), Expect = 0.43
Identities = 20/51 (39%), Positives = 21/51 (40%), Gaps = 2/51 (3%)
Frame = +2
Query: 375 PVQQQQQHQQARPTFPPIPMLYAELLPTLLHRGHCTTRQGKPPP--DPLPP 423
P QQQQ QQA P P LY P+ H PPP P PP
Sbjct: 134 PQQQQQPQQQALHVRSPNPFLYPFASPSRASANHAVGGYPPPPPPSQPQPP 286
>BF647147 weakly similar to GP|7303636|gb|A CG13214-PA {Drosophila
melanogaster}, partial (8%)
Length = 426
Score = 33.5 bits (75), Expect = 0.43
Identities = 33/122 (27%), Positives = 38/122 (31%), Gaps = 14/122 (11%)
Frame = +3
Query: 338 PYSQHPFFPPFYHQYPLPPGQPQVPVNAIAQQMKQQLPVQQQQQHQQARPT------FPP 391
P S +P P F YP PPG P QH + P FPP
Sbjct: 24 PMSNYPP-PGFGSPYPPPPGPPP-------------------HQHHEGYPPPGYPGGFPP 143
Query: 392 IPMLYAELLPTLLHRGHCTTRQ--------GKPPPDPLPPRFRSDLKCDFHQGALGHDVE 443
P P H H G PPP P PP++ + D H H
Sbjct: 144 PPP------PPHHHHHHGPPHDSYQGYFDNGYPPPPPAPPQYHNYQHVDHHHHHGHHGDP 305
Query: 444 GC 445
GC
Sbjct: 306 GC 311
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.317 0.134 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 30,799,533
Number of Sequences: 36976
Number of extensions: 465403
Number of successful extensions: 3505
Number of sequences better than 10.0: 90
Number of HSP's better than 10.0 without gapping: 3165
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3422
length of query: 1018
length of database: 9,014,727
effective HSP length: 106
effective length of query: 912
effective length of database: 5,095,271
effective search space: 4646887152
effective search space used: 4646887152
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 63 (28.9 bits)
Medicago: description of AC146794.6


