

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146794.13 + phase: 0 /pseudo
(818 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
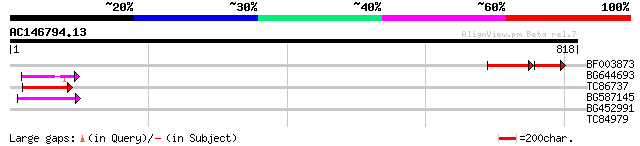
BF003873 similar to GP|14715222|em putative polyprotein {Cicer a... 106 6e-29
BG644693 weakly similar to GP|18767374|g Putative 22 kDa kafirin... 62 1e-09
TC86737 weakly similar to GP|6683624|dbj|BAA89272.1 Pol {Alterna... 55 1e-07
BG587145 similar to PIR|H86337|H8 protein F5M15.26 [imported] - ... 49 1e-05
BG452991 PIR|A25875|A25 histone H4 - Tetrahymena thermophila, pa... 42 0.001
TC84979 31 2.2
>BF003873 similar to GP|14715222|em putative polyprotein {Cicer arietinum},
partial (82%)
Length = 559
Score = 106 bits (264), Expect(2) = 6e-29
Identities = 47/66 (71%), Positives = 55/66 (83%)
Frame = +1
Query: 690 WSCLEIEEVDAEVYWPVSDIRKSWNSGI*SGFATTSFEFA*CVSCVATSEVCIGSIACDS 749
W+C E++EVD E++W VSDIRKSWN G+ SG T SFEFA*C+SCVATSEVC GSI+CD
Sbjct: 7 WTCFEVKEVDCEIHWSVSDIRKSWNGGVSSGITTASFEFA*CLSCVATSEVCSGSISCDP 186
Query: 750 EG*CAS 755
E *CAS
Sbjct: 187 E**CAS 204
Score = 40.4 bits (93), Expect(2) = 6e-29
Identities = 26/45 (57%), Positives = 31/45 (68%)
Frame = +3
Query: 757 EITLWLRPDH*GLKVAK*RS*EARRYL**K*FGEEQLVRV*RGGL 801
E TL R *GL + K*R *EARRYL *+ FG E++V+ *RG L
Sbjct: 207 ETTLR*RLYR*GLMIEK*RH*EARRYLL*ESFGTERMVKA*RGSL 341
>BG644693 weakly similar to GP|18767374|g Putative 22 kDa kafirin cluster;
Ty3-Gypsy type {Oryza sativa}, partial (15%)
Length = 716
Score = 61.6 bits (148), Expect = 1e-09
Identities = 36/88 (40%), Positives = 52/88 (58%), Gaps = 5/88 (5%)
Frame = +2
Query: 18 AFRCYMNKIFHAYLDWFVVVFIDDILIYSRSEEEHAEHLKIMLQVLKEKKMYAKLSKCEF 77
AF MN++F YLD V+VF +DILIYS++E EH HL++ L+VLK+ + C+
Sbjct: 467 AFMELMNRVFQDYLDSLVIVFSNDILIYSKNENEHENHLRLALKVLKD------IGLCQI 628
Query: 78 ----WLSEVSFLG-HIISDSGIVVTHQK 100
L EV F H+IS G+ V ++
Sbjct: 629 SYV*ILVEVGFFSLHVISGEGLKVDSKR 712
>TC86737 weakly similar to GP|6683624|dbj|BAA89272.1 Pol {Alternaria
alternata}, partial (21%)
Length = 1540
Score = 54.7 bits (130), Expect = 1e-07
Identities = 26/73 (35%), Positives = 47/73 (63%), Gaps = 1/73 (1%)
Frame = +1
Query: 19 FRCYMNKIFHAYLDWFVVVFIDDILIYSR-SEEEHAEHLKIMLQVLKEKKMYAKLSKCEF 77
F+ Y+NK H +LD FV +IDD+LIY+ S+++H ++ +L+ L + + KCEF
Sbjct: 862 FQRYINKTLHEFLDDFVTAYIDDVLIYTTGSKKDHEAQVRRVLRRLADAGLSLDPKKCEF 1041
Query: 78 WLSEVSFLGHIIS 90
++ V ++G I++
Sbjct: 1042SVTTVKYVGFILT 1080
>BG587145 similar to PIR|H86337|H8 protein F5M15.26 [imported] - Arabidopsis
thaliana, partial (13%)
Length = 763
Score = 48.5 bits (114), Expect = 1e-05
Identities = 27/90 (30%), Positives = 48/90 (53%)
Frame = +2
Query: 12 LRIFSDAFRCYMNKIFHAYLDWFVVVFIDDILIYSRSEEEHAEHLKIMLQVLKEKKMYAK 71
L+ ++ +N++F L + V+IDD+L+ S +H HLK + L E M
Sbjct: 80 LKNAGSTYQRLVNRMFADKLGNTMEVYIDDMLVKSLRATDHLNHLKE*FKTLDEYIMKLN 259
Query: 72 LSKCEFWLSEVSFLGHIISDSGIVVTHQKL 101
+KC F ++ FLG+I++ GI V +++
Sbjct: 260 PAKCTFGVTSGEFLGYIVTQQGIEVNPKQI 349
>BG452991 PIR|A25875|A25 histone H4 - Tetrahymena thermophila, partial (33%)
Length = 560
Score = 41.6 bits (96), Expect = 0.001
Identities = 35/72 (48%), Positives = 41/72 (56%)
Frame = +2
Query: 312 *TV*RLESSL*VVASECEVVYVEDQQ*ILEQY*RSAES*C*VCGLVGC**SN*RW*LQG* 371
*TV RLE L* VA+ECE+ VEDQQ I Y R +E C V *S+*RW* *
Sbjct: 71 *TVQRLEFGL*SVATECEIGDVEDQQRIFG*YQRGSEGGCEVGRFNVWE*SD*RW*F*S* 250
Query: 372 **RCVEIPRQNL 383
* R V + N+
Sbjct: 251 *SRSVAVSG*NM 286
>TC84979
Length = 641
Score = 30.8 bits (68), Expect = 2.2
Identities = 14/16 (87%), Positives = 15/16 (93%)
Frame = +2
Query: 801 LMSHLLLIFICFLV*F 816
LMSHLL IFIC+LV*F
Sbjct: 494 LMSHLLSIFICYLV*F 541
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.372 0.167 0.652
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 25,785,925
Number of Sequences: 36976
Number of extensions: 353803
Number of successful extensions: 4623
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 956
Number of HSP's successfully gapped in prelim test: 138
Number of HSP's that attempted gapping in prelim test: 3605
Number of HSP's gapped (non-prelim): 1211
length of query: 818
length of database: 9,014,727
effective HSP length: 104
effective length of query: 714
effective length of database: 5,169,223
effective search space: 3690825222
effective search space used: 3690825222
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.9 bits)
S2: 62 (28.5 bits)
Medicago: description of AC146794.13


