

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146792.8 - phase: 0
(266 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
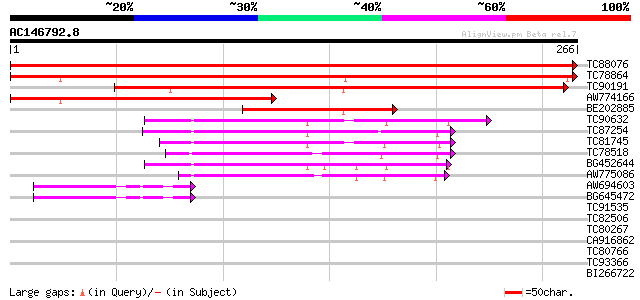
Sequences producing significant alignments: (bits) Value
TC88076 similar to GP|17979373|gb|AAL49912.1 putative membrane a... 533 e-152
TC78864 similar to GP|17979373|gb|AAL49912.1 putative membrane a... 371 e-103
TC90191 similar to GP|8809582|dbj|BAA97133.1 membrane associated... 317 4e-87
AW774166 similar to GP|17979373|gb| putative membrane associated... 130 4e-31
BE202885 similar to GP|8809582|dbj| membrane associated protein ... 115 1e-26
TC90632 similar to GP|21554053|gb|AAM63134.1 putative VAMP-assoc... 62 2e-10
TC87254 similar to PIR|JC7234|JC7234 27k vesicle-associated memb... 61 5e-10
TC81745 similar to PIR|JC7234|JC7234 27k vesicle-associated memb... 60 9e-10
TC78518 similar to PIR|JC7234|JC7234 27k vesicle-associated memb... 57 7e-09
BG452644 similar to GP|21554053|gb putative VAMP-associated prot... 51 4e-07
AW775086 similar to PIR|JC7234|JC7 27k vesicle-associated membra... 45 4e-05
AW694603 homologue to GP|8489871|gb|A DEAD box protein P68 {Pisu... 43 1e-04
BG645472 homologue to GP|8489871|gb| DEAD box protein P68 {Pisum... 43 1e-04
TC91535 similar to PIR|T06521|T06521 pitrilysin (EC 3.4.24.55) -... 40 0.001
TC82506 39 0.002
TC80267 similar to GP|6692128|gb|AAF24593.1| T19E23.14 {Arabidop... 38 0.005
CA916862 GP|12805231|gb Similar to mesenchyme homeobox 2 {Mus mu... 38 0.005
TC80766 similar to GP|15292967|gb|AAK93594.1 unknown protein {Ar... 37 0.006
TC93366 similar to PIR|E96745|E96745 hypothetical protein T9N14.... 37 0.008
BI266722 similar to GP|17473763|gb| putative protein {Arabidopsi... 36 0.018
>TC88076 similar to GP|17979373|gb|AAL49912.1 putative membrane associated
protein {Arabidopsis thaliana}, partial (65%)
Length = 1165
Score = 533 bits (1373), Expect = e-152
Identities = 266/266 (100%), Positives = 266/266 (100%)
Frame = +1
Query: 1 MAVAEPKLHSEPKVWNFFKLPFRNSNTSSTNTTSSSVNNNLHHHHHHNPNSNTPLEGSTS 60
MAVAEPKLHSEPKVWNFFKLPFRNSNTSSTNTTSSSVNNNLHHHHHHNPNSNTPLEGSTS
Sbjct: 103 MAVAEPKLHSEPKVWNFFKLPFRNSNTSSTNTTSSSVNNNLHHHHHHNPNSNTPLEGSTS 282
Query: 61 HTSNSVSSVARSLLPTRRRLKLDPSNKLYFPYEPGKQVRSAIRIKNTSKSNVAFKFQTTA 120
HTSNSVSSVARSLLPTRRRLKLDPSNKLYFPYEPGKQVRSAIRIKNTSKSNVAFKFQTTA
Sbjct: 283 HTSNSVSSVARSLLPTRRRLKLDPSNKLYFPYEPGKQVRSAIRIKNTSKSNVAFKFQTTA 462
Query: 121 PKSCFMRPPGAILAPGESIIATVFKFVEQPENNEKPEKSGLKFKIMSLKVKGSIDYVPEL 180
PKSCFMRPPGAILAPGESIIATVFKFVEQPENNEKPEKSGLKFKIMSLKVKGSIDYVPEL
Sbjct: 463 PKSCFMRPPGAILAPGESIIATVFKFVEQPENNEKPEKSGLKFKIMSLKVKGSIDYVPEL 642
Query: 181 FDEQKDQVAVEQILRVVFLDPERPSPILEKLKRQLADADAALEERKKPAEDAGPKIIGEG 240
FDEQKDQVAVEQILRVVFLDPERPSPILEKLKRQLADADAALEERKKPAEDAGPKIIGEG
Sbjct: 643 FDEQKDQVAVEQILRVVFLDPERPSPILEKLKRQLADADAALEERKKPAEDAGPKIIGEG 822
Query: 241 LVIDEWKERRERYLAKQQGDVVVDSV 266
LVIDEWKERRERYLAKQQGDVVVDSV
Sbjct: 823 LVIDEWKERRERYLAKQQGDVVVDSV 900
>TC78864 similar to GP|17979373|gb|AAL49912.1 putative membrane associated
protein {Arabidopsis thaliana}, partial (71%)
Length = 1303
Score = 371 bits (953), Expect = e-103
Identities = 193/271 (71%), Positives = 226/271 (83%), Gaps = 5/271 (1%)
Frame = +2
Query: 1 MAVAEPKLHSEPKVWNFFKLPF--RNSNTSSTNTTSSSVNNNLHHHHHHNPNSNTPLEGS 58
M V K S+ KVWNF ++PF ++N SS++TT+SS + + H+ HH S ++ S
Sbjct: 179 MEVESEKPGSDGKVWNFCRMPFWQTSNNPSSSSTTTSSSSTSYMHNVHHQSQSIHSVDRS 358
Query: 59 TSHTSNSVSSVARSLLPTRRRLKLDPSNKLYFPYEPGKQVRSAIRIKNTSKSNVAFKFQT 118
+S +VSSVA+SLLPTRRRL+LDP NKLYFPYEPGKQVRSAI IKNT KS+VAFKFQT
Sbjct: 359 VPQSSATVSSVAKSLLPTRRRLRLDPPNKLYFPYEPGKQVRSAITIKNTCKSHVAFKFQT 538
Query: 119 TAPKSCFMRPPGAILAPGESIIATVFKFVEQPENNEKP--EKSGLKFKIMSLKVKGSIDY 176
TAPKSC+MRPPG ILAPGESIIATVFKFVE PENNEKP +KS +KFKIMSLKV+G +DY
Sbjct: 539 TAPKSCYMRPPGGILAPGESIIATVFKFVEPPENNEKPTDQKSRVKFKIMSLKVQGEMDY 718
Query: 177 VPELFDEQKDQVAVEQILRVVFLDPERPSPILEKLKRQLADADAALEERKKPAEDAGPKI 236
VPELFDEQ+DQVAVEQILRVVFLDP+R SP ++KLKRQLA+A+AALE RKKP E+ GP++
Sbjct: 719 VPELFDEQRDQVAVEQILRVVFLDPDRNSPAMDKLKRQLAEAEAALEARKKPPEETGPRV 898
Query: 237 IGEGLVIDEWKERRERYLAKQQGD-VVVDSV 266
GEGLVIDEWKERRERYLAKQQ + VVVDSV
Sbjct: 899 AGEGLVIDEWKERRERYLAKQQVEGVVVDSV 991
>TC90191 similar to GP|8809582|dbj|BAA97133.1 membrane associated protein
{Arabidopsis thaliana}, partial (64%)
Length = 839
Score = 317 bits (811), Expect = 4e-87
Identities = 160/216 (74%), Positives = 183/216 (84%), Gaps = 3/216 (1%)
Frame = +1
Query: 50 NSNTPLEGSTSHTSNSVSSVARSLL-PTRRRLKLDPSNKLYFPYEPGKQVRSAIRIKNTS 108
N +G+ + +VSSVARSLL P RRRL+LDPSN LYFPYEPGKQVRSA+R+KNTS
Sbjct: 169 NVQNQTQGNAQKSPKTVSSVARSLLLPPRRRLRLDPSNHLYFPYEPGKQVRSAVRLKNTS 348
Query: 109 KSNVAFKFQTTAPKSCFMRPPGAILAPGESIIATVFKFVEQPENNEK--PEKSGLKFKIM 166
KS+VAFKFQTTAPKSC+MRPPG ILAPGES+IA VFKFVEQPENNEK +K+ +KFKIM
Sbjct: 349 KSHVAFKFQTTAPKSCYMRPPGGILAPGESVIAAVFKFVEQPENNEKLSNQKNKVKFKIM 528
Query: 167 SLKVKGSIDYVPELFDEQKDQVAVEQILRVVFLDPERPSPILEKLKRQLADADAALEERK 226
SLKVK +DYVPELFDEQ+D V VE+IL VVFLDPERPSP LEKLKRQLA+ADAA+E RK
Sbjct: 529 SLKVKLGVDYVPELFDEQRDLVTVERILGVVFLDPERPSPALEKLKRQLAEADAAVEARK 708
Query: 227 KPAEDAGPKIIGEGLVIDEWKERRERYLAKQQGDVV 262
KPA + GP++ EGLVIDEWKERRE+YLA+QQ V
Sbjct: 709 KPAAETGPRVAAEGLVIDEWKERREKYLARQQSQAV 816
>AW774166 similar to GP|17979373|gb| putative membrane associated protein
{Arabidopsis thaliana}, partial (21%)
Length = 656
Score = 130 bits (328), Expect = 4e-31
Identities = 70/127 (55%), Positives = 90/127 (70%), Gaps = 2/127 (1%)
Frame = +2
Query: 1 MAVAEPKLHSEPKVWNFFKLPF--RNSNTSSTNTTSSSVNNNLHHHHHHNPNSNTPLEGS 58
M V K S+ KVWNF ++PF ++N SS++TT+SS + + H+ HH S ++ S
Sbjct: 59 MEVESEKPGSDGKVWNFCRMPFWQTSNNPSSSSTTTSSSSTSYMHNVHHQSQSIHSVDRS 238
Query: 59 TSHTSNSVSSVARSLLPTRRRLKLDPSNKLYFPYEPGKQVRSAIRIKNTSKSNVAFKFQT 118
+S +VSSVA+SLLPTRRRL+LDP NKLYFPYEPGKQVRSAI IKNT KS+VAFK
Sbjct: 239 VPQSSATVSSVAKSLLPTRRRLRLDPPNKLYFPYEPGKQVRSAITIKNTCKSHVAFKVWF 418
Query: 119 TAPKSCF 125
+P + F
Sbjct: 419 ISPFTYF 439
>BE202885 similar to GP|8809582|dbj| membrane associated protein {Arabidopsis
thaliana}, partial (23%)
Length = 658
Score = 115 bits (289), Expect = 1e-26
Identities = 57/75 (76%), Positives = 65/75 (86%), Gaps = 2/75 (2%)
Frame = +1
Query: 110 SNVAFKFQTTAPKSCFMRPPGAILAPGESIIATVFKFVEQPENNEK--PEKSGLKFKIMS 167
S + F+FQTTAPKSC+MRPPG ILAPGES+IA VFKFVEQPENNEK +K+ +KFKIMS
Sbjct: 433 SLLCFQFQTTAPKSCYMRPPGGILAPGESVIAAVFKFVEQPENNEKLSNQKNKVKFKIMS 612
Query: 168 LKVKGSIDYVPELFD 182
LKVK +DYVPELFD
Sbjct: 613 LKVKLGVDYVPELFD 657
>TC90632 similar to GP|21554053|gb|AAM63134.1 putative VAMP-associated
protein {Arabidopsis thaliana}, partial (63%)
Length = 681
Score = 62.0 bits (149), Expect = 2e-10
Identities = 47/169 (27%), Positives = 86/169 (50%), Gaps = 6/169 (3%)
Frame = +1
Query: 64 NSVSSVARSLLPTRRRLKLDPSNKLYFPYEPGKQVRSAIRIKNTSKSNVAFKFQTTAPKS 123
+SV + ++ T L ++P +L F +E KQ+ ++++ N + S VAFK +TT P+
Sbjct: 139 DSVDRASTMMMSTGDLLNIEPV-ELKFLFELKKQISCSLQLSNKTDSYVAFKVKTTNPRK 315
Query: 124 CFMRPPGAILAPGES--IIATVFKFVEQPENNEKPEKSGLKFKIMSLKVKGSID---YVP 178
+RP ++ P + +I T+ E P + + + KF + S++V D P
Sbjct: 316 YCVRPNTGVVLPQSTCEVIVTMQAQKEVPPDMQCKD----KFLLQSVRVNDGADAKEITP 483
Query: 179 ELFDEQKDQVAVEQILRVVFLDPERP-SPILEKLKRQLADADAALEERK 226
E+F+++ V E LRVV++ P +P SP+ E + + + E K
Sbjct: 484 EMFNKEAGHVVEECKLRVVYVAPPQPQSPVPESSEEGSSPRXSVTENGK 630
>TC87254 similar to PIR|JC7234|JC7234 27k vesicle-associated membrane
protein-associated protein - curled-leaved tobacco,
partial (87%)
Length = 1040
Score = 60.8 bits (146), Expect = 5e-10
Identities = 43/150 (28%), Positives = 78/150 (51%), Gaps = 3/150 (2%)
Frame = +2
Query: 63 SNSVSSVARSLLPTRRRLKLDPSNKLYFPYEPGKQVRSAIRIKNTSKSNVAFKFQTTAPK 122
S + S + + + + L++ P +L FP+E KQ+ ++++ N S + +AFK +TT PK
Sbjct: 32 STATSLLRSTTMSSGELLQIHPQ-ELQFPFELRKQISCSLQLSNKSDNYIAFKVKTTNPK 208
Query: 123 SCFMRPPGAILAPGES--IIATVFKFVEQPENNEKPEKSGLKFKIMSLKVKGSIDYVPEL 180
+RP ++ P + I T+ E P + + +K L+ + S D PE+
Sbjct: 209 KYCVRPNNGVVLPRSTCDITVTMQGQKEAPPDMQCKDKFLLQSVVASPGATAK-DITPEM 385
Query: 181 FDEQKDQVAVEQILRVVFL-DPERPSPILE 209
F+++ E LRVV++ P+ PSP+ E
Sbjct: 386 FNKESGYEVDECKLRVVYVAPPQPPSPVRE 475
>TC81745 similar to PIR|JC7234|JC7234 27k vesicle-associated membrane
protein-associated protein - curled-leaved tobacco,
partial (94%)
Length = 945
Score = 60.1 bits (144), Expect = 9e-10
Identities = 43/145 (29%), Positives = 77/145 (52%), Gaps = 6/145 (4%)
Frame = +2
Query: 71 RSLLPTRRRLKLDPSNKLYFPYEPGKQVRSAIRIKNTSKSNVAFKFQTTAPKSCFMRPPG 130
+ + P L ++P +L FP+E KQ+ ++++ N + S VAFK +TT P+ +RP
Sbjct: 95 QEMSPDGDLLSIEPL-ELKFPFELKKQISCSLQLSNKTDSYVAFKVKTTNPRKYCVRPNT 271
Query: 131 AILAPGES--IIATVFKFVEQPENNEKPEKSGLKFKIMSLKVKGSI---DYVPELFDEQK 185
I+ P + ++ T+ E P + + + KF + S+K + D E+F+++
Sbjct: 272 GIVLPRSTCDVMVTMQAQKEAPADMQCKD----KFLLQSVKTNDGVSPKDISAEMFNKEA 439
Query: 186 DQVAVEQILRVVFLD-PERPSPILE 209
V E LRVV++ P+ PSP+ E
Sbjct: 440 GHVVEECKLRVVYVSPPQPPSPVPE 514
>TC78518 similar to PIR|JC7234|JC7234 27k vesicle-associated membrane
protein-associated protein - curled-leaved tobacco,
partial (83%)
Length = 827
Score = 57.0 bits (136), Expect = 7e-09
Identities = 43/142 (30%), Positives = 67/142 (46%), Gaps = 6/142 (4%)
Frame = +1
Query: 74 LPTRRRLKLDPSNKLYFPYEPGKQVRSAIRIKNTSKSNVAFKFQTTAPKSCFMRPPGAIL 133
+ T L + P ++L FP+E KQ+ ++++ N + + VAFK +TT PK +RP ++
Sbjct: 82 MSTGELLNIQP-DELQFPFELRKQISCSLQLSNKTDNYVAFKVKTTNPKKYCVRPNSGVV 258
Query: 134 APGESIIATVFKFVEQPENNEKPEKSGLKFKIMSLKVKGS-----IDYVPELFDEQKDQV 188
P + T V E P K K + V S D PE+F ++
Sbjct: 259 LPRSTCDVT----VTMQAQKEAPSDMQCKDKFLLQSVIASPGTTTKDINPEMFSKEAGHN 426
Query: 189 AVEQILRVVFL-DPERPSPILE 209
E LRVV++ P PSP+ E
Sbjct: 427 VEECKLRVVYVAPPGPPSPVRE 492
>BG452644 similar to GP|21554053|gb putative VAMP-associated protein
{Arabidopsis thaliana}, partial (61%)
Length = 629
Score = 51.2 bits (121), Expect = 4e-07
Identities = 46/173 (26%), Positives = 79/173 (45%), Gaps = 29/173 (16%)
Frame = +2
Query: 64 NSVSSVARSLLPTRRRLKLDPSNKLYFPYEPGKQVRSAIRIKNTSKSNVAFKFQTTAPKS 123
+SV + ++ T L ++P +L F +E KQ+ ++++ N + S VAFK +TT P+
Sbjct: 50 DSVDRASTMMMSTGDLLNIEPV-ELKFLFELKKQISCSLQLSNKTDSYVAFKVKTTNPRK 226
Query: 124 CFMRPPGAILAPGES----------IIATVFKF-------------VEQPENNEKPEKSG 160
+RP ++ P + I + +F F V E P
Sbjct: 227 YCVRPNTGVVLPQSTCEVIGIACYIIFSFLF*FFAG*WHCDLWLFIVTMQAQKEVPPDMQ 406
Query: 161 L--KFKIMSLKVKGSID---YVPELFDEQKDQVAVEQILRVVFLDPERP-SPI 207
KF + S++V D PE+F+++ V E LRVV++ P +P SP+
Sbjct: 407 CKDKFLLQSVRVNDGADAKEITPEMFNKEAGHVVEECKLRVVYVAPPQPQSPV 565
>AW775086 similar to PIR|JC7234|JC7 27k vesicle-associated membrane
protein-associated protein - curled-leaved tobacco,
partial (52%)
Length = 747
Score = 44.7 bits (104), Expect = 4e-05
Identities = 38/133 (28%), Positives = 61/133 (45%), Gaps = 6/133 (4%)
Frame = +2
Query: 80 LKLDPSNKLYFPYEPGKQVRSAIRIKNTSKSNVAFKFQTTAPKSCFMRPPGAILAPGESI 139
L++ P +L+FP+E KQ+ + ++ N S + +AFK +TT P+ +RP ++ P +
Sbjct: 44 LQIHPQ-ELHFPFELKKQISISFQMSNKSDNYLAFKVKTTIPEKYCVRPNNGVVLPRSTC 220
Query: 140 IATVFKFVEQPENNEKPEKSGL-KFKIMSLKVKGSI---DYVPELFDEQKDQVAVEQILR 195
V Q + P + KF I S+ K D E+F + LR
Sbjct: 221 DIQV---TMQAQKKAPPNMHCIDKFLIQSIVAKPGATTEDITSEMFFQDSGYKVEMCKLR 391
Query: 196 VVF--LDPERPSP 206
VV+ P PSP
Sbjct: 392 VVYDAPPPNLPSP 430
>AW694603 homologue to GP|8489871|gb|A DEAD box protein P68 {Pisum sativum},
partial (13%)
Length = 420
Score = 42.7 bits (99), Expect = 1e-04
Identities = 29/76 (38%), Positives = 40/76 (52%)
Frame = +2
Query: 12 PKVWNFFKLPFRNSNTSSTNTTSSSVNNNLHHHHHHNPNSNTPLEGSTSHTSNSVSSVAR 71
PK ++ RN+ +S+T +T++ +N HHHHHHN N L S SH SNS S
Sbjct: 47 PKTMSYVPPHLRNAASSTTVSTNTGTLDNHHHHHHHNNN----LAFSNSH-SNSPSLFNA 211
Query: 72 SLLPTRRRLKLDPSNK 87
S RR PS++
Sbjct: 212 S----RRTSAAPPSSR 247
>BG645472 homologue to GP|8489871|gb| DEAD box protein P68 {Pisum sativum},
partial (31%)
Length = 768
Score = 42.7 bits (99), Expect = 1e-04
Identities = 29/76 (38%), Positives = 40/76 (52%)
Frame = +3
Query: 12 PKVWNFFKLPFRNSNTSSTNTTSSSVNNNLHHHHHHNPNSNTPLEGSTSHTSNSVSSVAR 71
PK ++ RN+ +S+T +T++ +N HHHHHHN N L S SH SNS S
Sbjct: 39 PKTMSYVPPHLRNAASSTTVSTNTGTLDNHHHHHHHNNN----LAFSNSH-SNSPSLFNA 203
Query: 72 SLLPTRRRLKLDPSNK 87
S RR PS++
Sbjct: 204 S----RRTSAAPPSSR 239
>TC91535 similar to PIR|T06521|T06521 pitrilysin (EC 3.4.24.55) - garden
pea, partial (26%)
Length = 1146
Score = 40.0 bits (92), Expect = 0.001
Identities = 24/57 (42%), Positives = 33/57 (57%), Gaps = 4/57 (7%)
Frame = +3
Query: 28 SSTNTTSSSVNNNLHHHHHHNPNSNTPLEGSTSHTSN----SVSSVARSLLPTRRRL 80
+ST+TTSS N +L HHHHH + ++P ST SN S SS++ S RR+
Sbjct: 81 ASTSTTSSLTNLSLPHHHHHRHHRHSPSSISTRFRSNRFFLSSSSLSFSSPQRERRV 251
>TC82506
Length = 676
Score = 39.3 bits (90), Expect = 0.002
Identities = 35/135 (25%), Positives = 62/135 (45%), Gaps = 14/135 (10%)
Frame = +2
Query: 78 RRLKLDPSNKLYFPYEPGKQVRSAIRIKNTSKS-NVAFKFQTTAPKSCFMRPPGAILAPG 136
R +KLDPSN + E G++ I + N + VAF+ Q ++P I++P
Sbjct: 206 RLIKLDPSNIVLIRVEEGQKCLGKITLNNVMYTMPVAFRIQPLIKTRYTIKPQSGIISPL 385
Query: 137 ESIIATV-FKFVEQPENNEKPEK---SGLKFKIMSLKVKGS--------IDYVP-ELFDE 183
S++ + + +Q +N P S F + S+ G+ D VP + F
Sbjct: 386 ASLVIEITYHPPQQQGSNNLPHSFPFSDDSFLLHSVLAPGAAIKEPSSMFDSVPSDWFTT 565
Query: 184 QKDQVAVEQILRVVF 198
+K QV ++ ++V+F
Sbjct: 566 KKKQVFIDSAIKVMF 610
>TC80267 similar to GP|6692128|gb|AAF24593.1| T19E23.14 {Arabidopsis
thaliana}, partial (24%)
Length = 1625
Score = 37.7 bits (86), Expect = 0.005
Identities = 28/84 (33%), Positives = 43/84 (50%), Gaps = 9/84 (10%)
Frame = +1
Query: 16 NFFKLPFRNSNTSST----NTTSSSVNNNLHHHHHHNPNSNTPLEGSTSHTSN-SVSSVA 70
NF N TSST +++S++V ++ H+HHHHN + + G H + +SS+
Sbjct: 136 NFMANSSWNWRTSSTCDYSSSSSTTVKHHKHYHHHHNHQHDLLIPGLPDHIAQLCLSSIN 315
Query: 71 RSLL----PTRRRLKLDPSNKLYF 90
SLL + RRL PS +F
Sbjct: 316 PSLLFKVCHSWRRLIYSPSFPPFF 387
>CA916862 GP|12805231|gb Similar to mesenchyme homeobox 2 {Mus musculus},
partial (5%)
Length = 727
Score = 37.7 bits (86), Expect = 0.005
Identities = 21/78 (26%), Positives = 34/78 (42%), Gaps = 7/78 (8%)
Frame = +2
Query: 24 NSNTSSTNTTSSSVNNNLHHHHHHNPNSNTPLEGSTS-------HTSNSVSSVARSLLPT 76
N +S+T+T S++ +HHHHHH+ + T+ H + + + LL
Sbjct: 365 NPTSSTTSTASTAATPTIHHHHHHHHHQQQSRNNRTNRFVTFHIHRHVDLINSSNGLLCL 544
Query: 77 RRRLKLDPSNKLYFPYEP 94
R PS LY+ P
Sbjct: 545 RASNCRTPSRSLYYICNP 598
Score = 26.9 bits (58), Expect = 8.2
Identities = 14/58 (24%), Positives = 28/58 (48%)
Frame = +1
Query: 6 PKLHSEPKVWNFFKLPFRNSNTSSTNTTSSSVNNNLHHHHHHNPNSNTPLEGSTSHTS 63
P + +P + L +R+ S T S + ++LH H +PN+++ S+ T+
Sbjct: 280 PISYPKP*YFQSLFLLYRHPTFSRFQTLQSHLLHHLHRFHRSHPNNSSSSSSSSPSTT 453
>TC80766 similar to GP|15292967|gb|AAK93594.1 unknown protein {Arabidopsis
thaliana}, partial (59%)
Length = 785
Score = 37.4 bits (85), Expect = 0.006
Identities = 40/150 (26%), Positives = 56/150 (36%), Gaps = 28/150 (18%)
Frame = +1
Query: 9 HSEPKVWNFFKLPFRNSNTSSTNT----TSSSVNNNLHHHHHHNPNSNTPLEGST----- 59
H PK+ + F+ + NT +TN TS NN HH H P+ TPL T
Sbjct: 7 HHPPKISSLFQ----SDNTKNTNGLQFYTSPITTNNHHHLPHQPPHPPTPLPHKTHLHPQ 174
Query: 60 ----------------SHTSNSVSSVARSLLPTRRRLKLDPSNKLYFPYEPGKQVRSAIR 103
+H N + + S R L S+ ++P +V S +
Sbjct: 175 P*THHKTPSLILTNKQAHNPNHIQTTTTSKAQCRNHLLRRWSSLRRPHHQPPPRVHSCMA 354
Query: 104 IKNTSKSNVAF--KFQTTAPKS-CFMRPPG 130
NTS S Q PKS C++R G
Sbjct: 355 PLNTSSSFTCTLPTLQVHKPKSECYIRLNG 444
>TC93366 similar to PIR|E96745|E96745 hypothetical protein T9N14.11
[imported] - Arabidopsis thaliana, partial (27%)
Length = 1082
Score = 37.0 bits (84), Expect = 0.008
Identities = 19/58 (32%), Positives = 32/58 (54%), Gaps = 6/58 (10%)
Frame = +3
Query: 17 FFKLPFRNSNTSSTNTTSSSVNN------NLHHHHHHNPNSNTPLEGSTSHTSNSVSS 68
F +P +TSS++++SS + N +H HHHH+ + +PL ST+ +SS
Sbjct: 147 FTSIPIMPFSTSSSSSSSSFLRNLQRRRFRIHCHHHHHHQTKSPLSNSTNPWQLCLSS 320
>BI266722 similar to GP|17473763|gb| putative protein {Arabidopsis thaliana},
partial (28%)
Length = 404
Score = 35.8 bits (81), Expect = 0.018
Identities = 30/116 (25%), Positives = 55/116 (46%), Gaps = 7/116 (6%)
Frame = +2
Query: 54 PLEGSTSHTSNSVSSVARS--LLPTRRRLKLDPSNKLYF---PYEPGKQVRSAIRIKNTS 108
PL S N + +A S ++ T L ++P + E KQ+ ++++ N +
Sbjct: 38 PLSRSHCCRHNQIL*IAPSTMMMSTGDLLNIEPVELKFLFSVSVELKKQISCSLQLSNKT 217
Query: 109 KSNVAFKFQTTAPKSCFMRPPGAILAPGES--IIATVFKFVEQPENNEKPEKSGLK 162
S VAFK +TT P+ +RP ++ P + +I T+ E P + + +K L+
Sbjct: 218 DSYVAFKVKTTNPRKYCVRPNTGVVLPQSTCEVIVTMQAQKEVPPDMQCKDKXLLQ 385
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.313 0.130 0.369
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,410,556
Number of Sequences: 36976
Number of extensions: 136884
Number of successful extensions: 3880
Number of sequences better than 10.0: 266
Number of HSP's better than 10.0 without gapping: 2463
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3284
length of query: 266
length of database: 9,014,727
effective HSP length: 94
effective length of query: 172
effective length of database: 5,538,983
effective search space: 952705076
effective search space used: 952705076
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 57 (26.6 bits)
Medicago: description of AC146792.8


