

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146792.3 + phase: 0
(518 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
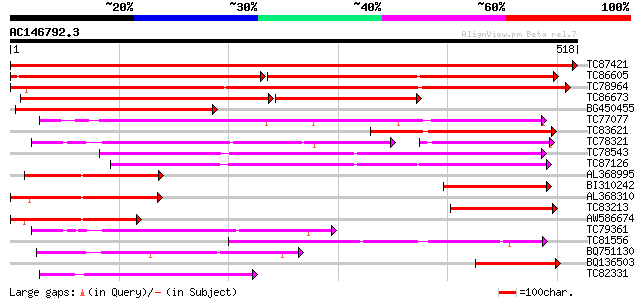
Score E
Sequences producing significant alignments: (bits) Value
TC87421 sugar transporter 1040 0.0
TC86605 similar to PIR|T01853|T01853 probable hexose transport p... 323 e-164
TC78964 weakly similar to PIR|T00450|T00450 probable monosacchar... 513 e-146
TC86673 weakly similar to PIR|S25015|S25015 monosaccharide trans... 297 e-126
BG450455 similar to SP|Q07423|HEX6 Hexose carrier protein HEX6. ... 236 2e-62
TC77077 similar to GP|9293909|dbj|BAB01812.1 sugar transporter p... 208 4e-54
TC83621 similar to PIR|T01506|T01506 probable hexose transport p... 204 9e-53
TC78321 similar to GP|21689841|gb|AAM67564.1 unknown protein {Ar... 177 9e-45
TC78543 similar to PIR|T14545|T14545 probable sugar transporter ... 173 2e-43
TC87126 similar to GP|15724240|gb|AAL06513.1 At1g75220/F22H5_6 {... 171 5e-43
AL368995 similar to GP|2104547|gb| AGAA.1 {Arabidopsis thaliana}... 150 1e-36
BI310242 similar to GP|347853|gb|A glucose transporter {Saccharu... 144 1e-34
AL368310 weakly similar to GP|16945177|em STP5 protein {Arabidop... 143 2e-34
TC83213 weakly similar to GP|16945177|emb|CAC69071. STP5 protein... 139 2e-33
AW586674 weakly similar to SP|Q10710|STA_ Sugar carrier protein ... 130 1e-30
TC79361 similar to GP|17380890|gb|AAL36257.1 putative membrane t... 124 7e-29
TC81556 similar to GP|2688830|gb|AAB88879.1| putative sugar tran... 123 1e-28
BQ751130 weakly similar to GP|15289796|dbj putative monosacchari... 110 1e-24
BQ136503 weakly similar to PIR|T05156|T051 probable glucose tran... 105 3e-23
TC82331 weakly similar to EGAD|120604|128852 hypothetical protei... 99 3e-21
>TC87421 sugar transporter
Length = 1753
Score = 1040 bits (2690), Expect = 0.0
Identities = 517/518 (99%), Positives = 517/518 (99%)
Frame = +2
Query: 1 MAGGGIPIGGGNKEYPGNLTPFVTITCIVAAMGGLIFGYDIGISGGVTSMDPFLKKFFPA 60
MAGGGIPIGGGNKEYPGNLTPFVTITCIVAAMGGLIFGYDIGISGGVTSMDPFLKKFFPA
Sbjct: 29 MAGGGIPIGGGNKEYPGNLTPFVTITCIVAAMGGLIFGYDIGISGGVTSMDPFLKKFFPA 208
Query: 61 VYRKKNKDKSTNQYCQYDSQTLTMFTSSLYLAALLSSLVASTITRRFGRKLSMLFGGLLF 120
VYRKKNKDKSTNQYCQYDSQTLTMFTSSLYLAALLSSLVASTITRRFGRKLSMLFGGLLF
Sbjct: 209 VYRKKNKDKSTNQYCQYDSQTLTMFTSSLYLAALLSSLVASTITRRFGRKLSMLFGGLLF 388
Query: 121 LVGALINGFANHVWMLIVGRILLGFGIGFANQAVPLYLSEMAPYKYRGALNIGFQLSITI 180
LVGALINGFANHVWMLIVGRILLGFGIGFANQ VPLYLSEMAPYKYRGALNIGFQLSITI
Sbjct: 389 LVGALINGFANHVWMLIVGRILLGFGIGFANQPVPLYLSEMAPYKYRGALNIGFQLSITI 568
Query: 181 GILVANVLNYFFAKIKGGWGWRLSLGGAMVPALIITIGSLVLPDTPNSMIERGDRDGAKA 240
GILVANVLNYFFAKIKGGWGWRLSLGGAMVPALIITIGSLVLPDTPNSMIERGDRDGAKA
Sbjct: 569 GILVANVLNYFFAKIKGGWGWRLSLGGAMVPALIITIGSLVLPDTPNSMIERGDRDGAKA 748
Query: 241 QLKRIRGIEDVDEEFNDLVAASEASMQVENPWRNLLQRKYRPQLTMAVLIPFFQQFTGIN 300
QLKRIRGIEDVDEEFNDLVAASEASMQVENPWRNLLQRKYRPQLTMAVLIPFFQQFTGIN
Sbjct: 749 QLKRIRGIEDVDEEFNDLVAASEASMQVENPWRNLLQRKYRPQLTMAVLIPFFQQFTGIN 928
Query: 301 VIMFYAPVLFNSIGFKDDASLMSAVITGVVNVVATCVSIYGVDKWGRRALFLEGGAQMLI 360
VIMFYAPVLFNSIGFKDDASLMSAVITGVVNVVATCVSIYGVDKWGRRALFLEGGAQMLI
Sbjct: 929 VIMFYAPVLFNSIGFKDDASLMSAVITGVVNVVATCVSIYGVDKWGRRALFLEGGAQMLI 1108
Query: 361 CQVAVAAAIGAKFGTSGNPGNLPEWYAIVVVLFICIYVAGFAWSWGPLGWLVPSEIFPLE 420
CQVAVAAAIGAKFGTSGNPGNLPEWYAIVVVLFICIYVAGFAWSWGPLGWLVPSEIFPLE
Sbjct: 1109CQVAVAAAIGAKFGTSGNPGNLPEWYAIVVVLFICIYVAGFAWSWGPLGWLVPSEIFPLE 1288
Query: 421 IRSAAQSVNVSVNMLFTFLVAQVFLIMLCHMKFGLFLFFAFFVLVMSIYVFFLLPETKGI 480
IRSAAQSVNVSVNMLFTFLVAQVFLIMLCHMKFGLFLFFAFFVLVMSIYVFFLLPETKGI
Sbjct: 1289IRSAAQSVNVSVNMLFTFLVAQVFLIMLCHMKFGLFLFFAFFVLVMSIYVFFLLPETKGI 1468
Query: 481 PIEEMDRVWKSHPFWSRFVEHGDHGNGVEMGKGAPKNV 518
PIEEMDRVWKSHPFWSRFVEHGDHGNGVEMGKGAPKNV
Sbjct: 1469PIEEMDRVWKSHPFWSRFVEHGDHGNGVEMGKGAPKNV 1582
>TC86605 similar to PIR|T01853|T01853 probable hexose transport protein
F9D12.17 - Arabidopsis thaliana, partial (91%)
Length = 3111
Score = 323 bits (828), Expect(2) = e-164
Identities = 156/267 (58%), Positives = 203/267 (75%), Gaps = 1/267 (0%)
Frame = +3
Query: 236 DGAKAQLKRIRGIEDVDEEFNDLVAASEASMQVENPWRNLLQRKYRPQLTMAVLIPFFQQ 295
D KA L++IRG ++++ EF +LV AS + +V++P+RNLL+R RPQL +++ + FQQ
Sbjct: 1995 DKGKAVLRKIRGTDNIEPEFLELVEASRVAKEVKHPFRNLLKRNNRPQLVISIALMIFQQ 2174
Query: 296 FTGINVIMFYAPVLFNSIGFKDDASLMSAVITGVVNVVATCVSIYGVDKWGRRALFLEGG 355
FTGIN IMFYAPVLFN++GFK+DA+L SAVITG +NV++T VSIY VDK GRR L LE G
Sbjct: 2175 FTGINAIMFYAPVLFNTLGFKNDAALYSAVITGAINVISTIVSIYSVDKLGRRKLLLEAG 2354
Query: 356 AQMLICQVAVAAAIGAKFGTSGNPGNLPEWYAIVVVLFICIYVAGFAWSWGPLGWLVPSE 415
QML+ Q+ +A +G K + L + YA +VV+ +CI+V+ FAWSWGPL WL+PSE
Sbjct: 2355 VQMLLSQMVIAIVLGIK--VKDHSEELSKGYAALVVVMVCIFVSAFAWSWGPLAWLIPSE 2528
Query: 416 IFPLEIRSAAQSVNVSVNMLFTFLVAQVFLIMLCHMKFGLFLFFAFFVLVMSIYVFFLLP 475
IFPLE RSA QSV V VN LFT ++AQ FL MLC+ KFG+F FF+ ++L MS +VFFL+P
Sbjct: 2529 IFPLETRSAGQSVTVCVNFLFTAVIAQAFLSMLCYFKFGIFFFFSGWILFMSTFVFFLVP 2708
Query: 476 ETKGIPIEEM-DRVWKSHPFWSRFVEH 501
ETK +PIEEM RVWK H FW RFVE+
Sbjct: 2709 ETKNVPIEEMTQRVWKQHWFWKRFVEN 2789
Score = 275 bits (703), Expect(2) = e-164
Identities = 136/234 (58%), Positives = 176/234 (75%), Gaps = 1/234 (0%)
Frame = +2
Query: 1 MAGGGIPIGGGNKEYPGNLTPFVTITCIVAAMGGLIFGYDIGISGGVTSMDPFLKKFFPA 60
M GGG GG ++E+ +TP + I+CI+AA GGL+FGYD+G+SGGV SM PFLKKFFP
Sbjct: 1289 MTGGGFS-GGNDREFEAKITPIIIISCIMAATGGLMFGYDVGVSGGVASMPPFLKKFFPT 1465
Query: 61 VYRKKNK-DKSTNQYCQYDSQTLTMFTSSLYLAALLSSLVASTITRRFGRKLSMLFGGLL 119
V R+ + D S + YC+YD+Q L +FTSSLYLA L + AS TR GR+L+ML G
Sbjct: 1466 VLRQTTESDGSESNYCKYDNQGLQLFTSSLYLAGLTVTFFASYTTRVLGRRLTMLIAGFF 1645
Query: 120 FLVGALINGFANHVWMLIVGRILLGFGIGFANQAVPLYLSEMAPYKYRGALNIGFQLSIT 179
F+ G +N A ++ MLIVGR+LLG GIGFANQAVP++LSE+AP + RGALNI FQL IT
Sbjct: 1646 FIAGVSLNASAQNLLMLIVGRVLLGCGIGFANQAVPVFLSEIAPSRIRGALNILFQLDIT 1825
Query: 180 IGILVANVLNYFFAKIKGGWGWRLSLGGAMVPALIITIGSLVLPDTPNSMIERG 233
+GIL AN++NY KIKG WGWR+SLG +PAL++T+G+ ++ DTPNS+IERG
Sbjct: 1826 LGILYANLVNYATNKIKGHWGWRISLGLGGIPALLLTLGAYLVVDTPNSLIERG 1987
>TC78964 weakly similar to PIR|T00450|T00450 probable monosaccharide
transport protein T14N5.7 - Arabidopsis thaliana,
partial (98%)
Length = 1771
Score = 513 bits (1321), Expect = e-146
Identities = 263/517 (50%), Positives = 358/517 (68%), Gaps = 5/517 (0%)
Frame = +1
Query: 1 MAGGGIPIGGGNKE---YPGNLTPFVTITCIVAAMGGLIFGYDIGISGGVTSMDPFLKKF 57
MAGGG+ GG K Y T + TC+V A+GG +FGYD+G+SGGVTSMD FL+KF
Sbjct: 31 MAGGGLTNGGPGKRAHLYEHKFTAYFAFTCVVGALGGSLFGYDLGVSGGVTSMDDFLEKF 210
Query: 58 FPAVYRKKNKDKSTNQYCQYDSQTLTMFTSSLYLAALLSSLVASTITRRFGRKLSMLFGG 117
FP VYRKK+ YC+YD+Q LT+FTSSLY +AL+ + AS +TR GRK +++ G
Sbjct: 211 FPDVYRKKHAHLKETDYCKYDNQVLTLFTSSLYFSALVMTFFASYLTRNKGRKATIIVGA 390
Query: 118 LLFLVGALINGFANHVWMLIVGRILLGFGIGFANQAVPLYLSEMAPYKYRGALNIGFQLS 177
L FL+GA++N A ++ LI+GR+ LG GIGF NQAVPLYLSEMAP RGA+N FQ +
Sbjct: 391 LSFLIGAILNAAAQNIPTLIIGRVFLGGGIGFGNQAVPLYLSEMAPASSRGAVNQLFQFT 570
Query: 178 ITIGILVANVLNYFFAKIKGGWGWRLSLGGAMVPALIITIGSLVLPDTPNSMIERGDRDG 237
GIL+AN++NYF KI GWR+SLG A +PA+++ +G + +TPNS++E+G D
Sbjct: 571 TCAGILIANLVNYFTDKIH-PHGWRISLGLAGIPAVLMLLGGIFCAETPNSLVEQGRLDE 747
Query: 238 AKAQLKRIRGIEDVDEEFNDLVAASEASMQVENPWRNLLQRKYRPQLTMAVL-IPFFQQF 296
A+ L+++RG ++VD EF DL ASE + V++P++ LL+RKYRPQL + L P FQQ
Sbjct: 748 ARKVLEKVRGTKNVDAEFEDLKDASELAQAVKSPFKVLLKRKYRPQLIIGALGTPAFQQL 927
Query: 297 TGINVIMFYAPVLFNSIGFKDDASLMSAVITGVVNVVATCVSIYGVDKWGRRALFLEGGA 356
TG N I+FYAPV+F S GF +A+L S+ IT +VAT +S++ VDK+G R FLE G
Sbjct: 928 TGNNSILFYAPVIFQSSGFGSNAALFSSFITNGALLVATVISMFLVDKFGTRKFFLEAGF 1107
Query: 357 QMLICQVAVAAAIGAKFGTSGNPGNLPEWYAIVVVLFICIYVAGFAWSWGPLGWLVPSEI 416
+M+ C + A + +F G+ L + + +V+ I +V + SWGPLGWLVPSE+
Sbjct: 1108EMICCMIITAVVLAVEF---GHGKELSKGISAFLVIMIFWFVLAYGRSWGPLGWLVPSEL 1278
Query: 417 FPLEIRSAAQSVNVSVNMLFTFLVAQVFLIMLCHMKFGLFLFFAFFVLVMSIYVFFLLPE 476
FPLEIRSAAQS+ V VNM+FT LVAQ+FL+ LCH+K+G+FL F ++VMS++VFFLLPE
Sbjct: 1279FPLEIRSAAQSIAVCVNMIFTALVAQLFLLSLCHLKYGIFLLFGGLIVVMSVFVFFLLPE 1458
Query: 477 TKGIPIEEMDRVWKSHPFWSRFVEHG-DHGNGVEMGK 512
TK +PIEE+ ++++H FW V G D G GK
Sbjct: 1459TKQVPIEEIYLLFENHWFWKNIVREGTDQEQGKPNGK 1569
>TC86673 weakly similar to PIR|S25015|S25015 monosaccharide transport
protein MST1 - common tobacco, partial (65%)
Length = 1162
Score = 297 bits (760), Expect(2) = e-126
Identities = 146/232 (62%), Positives = 186/232 (79%), Gaps = 1/232 (0%)
Frame = +2
Query: 11 GNKEYPGNLTPFVTITCIVAAMGGLIFGYDIGISGGVTSMDPFLKKFFPAVYRKKNKDK- 69
G +YPG LT V ITCI+AA GGLIFGYD G+SGGVTSMD FLK+FFP+VY +++ K
Sbjct: 59 GPSKYPGKLTFRVIITCIMAATGGLIFGYDHGVSGGVTSMDSFLKRFFPSVYEQESNLKP 238
Query: 70 STNQYCQYDSQTLTMFTSSLYLAALLSSLVASTITRRFGRKLSMLFGGLLFLVGALINGF 129
S NQYC+++SQ LT+FTSSLY++AL++ L AS+ITR GR+ +M+ GG+ F+ GAL+NG
Sbjct: 239 SANQYCKFNSQILTLFTSSLYISALVAGLGASSITRALGRRTTMILGGIFFVSGALLNGL 418
Query: 130 ANHVWMLIVGRILLGFGIGFANQAVPLYLSEMAPYKYRGALNIGFQLSITIGILVANVLN 189
A ++ MLI GR+LLGFGIG ANQAVP+YLSEMAPYKYRGALN+ FQLSITIGI AN+ N
Sbjct: 419 AMNIAMLIAGRLLLGFGIGCANQAVPIYLSEMAPYKYRGALNMCFQLSITIGIFTANMFN 598
Query: 190 YFFAKIKGGWGWRLSLGGAMVPALIITIGSLVLPDTPNSMIERGDRDGAKAQ 241
Y+F+KI G GWRLSLG VPA++ IGS+ LPD+PNS++ RG + A+ +
Sbjct: 599 YYFSKILNGEGWRLSLGLGAVPAVVFIIGSICLPDSPNSLVTRGRHEEARKE 754
Score = 174 bits (441), Expect(2) = e-126
Identities = 86/133 (64%), Positives = 103/133 (76%)
Frame = +3
Query: 244 RIRGIEDVDEEFNDLVAASEASMQVENPWRNLLQRKYRPQLTMAVLIPFFQQFTGINVIM 303
+IRG +DVD EF D+VAASEAS +V++PW+ L +RKYRPQL A+LIPFFQQFTG+NVI
Sbjct: 762 KIRGTDDVDAEFRDIVAASEASAKVKHPWKTLQERKYRPQLVFAILIPFFQQFTGLNVIT 941
Query: 304 FYAPVLFNSIGFKDDASLMSAVITGVVNVVATCVSIYGVDKWGRRALFLEGGAQMLICQV 363
FYAP+LF +IGF ASLMSA I G V+T VSI+ VDK+GRRALFLEGG QMLI +
Sbjct: 942 FYAPILFRTIGFGSQASLMSAAIIGSFKPVSTLVSIFVVDKFGRRALFLEGGVQMLISXI 1121
Query: 364 AVAAAIGAKFGTS 376
+ AI FGTS
Sbjct: 1122IMTVAIAVTFGTS 1160
>BG450455 similar to SP|Q07423|HEX6 Hexose carrier protein HEX6. [Castor
bean] {Ricinus communis}, partial (36%)
Length = 573
Score = 236 bits (601), Expect = 2e-62
Identities = 114/185 (61%), Positives = 144/185 (77%)
Frame = +2
Query: 6 IPIGGGNKEYPGNLTPFVTITCIVAAMGGLIFGYDIGISGGVTSMDPFLKKFFPAVYRKK 65
+PI + Y G +TP V ++C+VAA GG+IFGYDIGISGGVTSM PFL+KFFP VY K
Sbjct: 14 VPIPSNGRGYNGKMTPIVILSCMVAATGGIIFGYDIGISGGVTSMVPFLEKFFPDVYTKM 193
Query: 66 NKDKSTNQYCQYDSQTLTMFTSSLYLAALLSSLVASTITRRFGRKLSMLFGGLLFLVGAL 125
+D + YC++DSQ LT FTSSLY+A LL+S AS+ITR FGRK S+L GG FL+GA
Sbjct: 194 KQDNKISNYCKFDSQLLTTFTSSLYIAGLLASFFASSITRAFGRKPSILVGGAAFLIGAA 373
Query: 126 INGFANHVWMLIVGRILLGFGIGFANQAVPLYLSEMAPYKYRGALNIGFQLSITIGILVA 185
+ G A +++MLI+GR+LLG GIGFANQAVPLYLS MA +YRGA+NIGFQL + IG+L
Sbjct: 374 LGGAALNIYMLILGRVLLGVGIGFANQAVPLYLSXMALPRYRGAINIGFQLCVGIGVLSX 553
Query: 186 NVLNY 190
N++N+
Sbjct: 554 NLINF 568
>TC77077 similar to GP|9293909|dbj|BAB01812.1 sugar transporter protein
{Arabidopsis thaliana}, partial (85%)
Length = 1752
Score = 208 bits (530), Expect = 4e-54
Identities = 137/488 (28%), Positives = 242/488 (49%), Gaps = 25/488 (5%)
Frame = +1
Query: 28 IVAAMGGLIFGYDIGISGGVTSMDPFLKKFFPAVYRKKNKDKSTNQYCQYDSQTLTMFTS 87
++A+M ++ GYDIG+ G A+Y K++ S + + +
Sbjct: 124 MLASMTSILLGYDIGVMSGA------------AIYIKRDLKVSDGK--------IEVLLG 243
Query: 88 SLYLAALLSSLVASTITRRFGRKLSMLFGGLLFLVGALINGFANHVWMLIVGRILLGFGI 147
+ + +L+ S +A + GR+ +++F G +F VGAL+ GF+ + L+ GR + G GI
Sbjct: 244 IINIYSLIGSCLAGRTSDWIGRRYTIVFAGAIFFVGALLMGFSPNYNFLMFGRFVAGVGI 423
Query: 148 GFANQAVPLYLSEMAPYKYRGALNIGFQLSITIGILVANVLNYFFAKIKGGWGWRLSLGG 207
G+A P+Y +E++P RG L ++ I GIL+ + NY F+K+ GWR+ LG
Sbjct: 424 GYALMIAPVYTAEVSPASSRGFLTSFPEVFINSGILLGYISNYAFSKLSLKLGWRMMLGV 603
Query: 208 AMVPALIITIGSLVLPDTPNSMIERG--------------DRDGAKAQLKRIRGIEDVDE 253
+P++I+ +G L +P++P ++ RG ++ A+ +L I+ + E
Sbjct: 604 GAIPSVILAVGVLAMPESPRWLVMRGRLGDAIKVLNKTSDSKEEAQLRLAEIKQAAGIPE 783
Query: 254 EFNDLVAASEASMQVENPWRNLL---QRKYRPQLTMAVLIPFFQQFTGINVIMFYAPVLF 310
+ ND V + E W+ L R + A+ I FFQQ +G++ ++ Y+P +F
Sbjct: 784 DCNDDVVEVKVKNTGEGVWKELFLYPTPAVRHIVIAALGIHFFQQASGVDAVVLYSPTIF 963
Query: 311 NSIGFKDDASLMSAVI-TGVVNVVATCVSIYGVDKWGRRALFLE--GGAQMLICQVAVAA 367
G D L+ A I G V + V+ + +D++GRR L L GG + + +AV+
Sbjct: 964 KKAGINGDTHLLIATIAVGFVKTLFILVATFMLDRYGRRPLLLTSVGGMVVSLLTLAVSL 1143
Query: 368 AIGAKFGTSGNPGNLPEWYAIVVVLFICIYVAGFAWSWGPLGWLVPSEIFPLEIRSAAQS 427
I T N W + + + YVA F+ GP+ W+ SEIFPL +R+ +
Sbjct: 1144TIIDHSNTKLN------WAIGLSIATVLSYVATFSIGAGPITWVYSSEIFPLRLRAQGAA 1305
Query: 428 VNVSVNMLFTFLVAQVFLIMLCHMKF-GLFLFFAFFVLVMSIYVFFLLPETKGIPIEEMD 486
V VN + + +++ FL + + G F F + I+ + +LPET+G +EEM+
Sbjct: 1306CGVVVNRVTSGVISMTFLSLSKGITIGGAFFLFGGIATIGWIFFYIMLPETQGKTLEEME 1485
Query: 487 ----RVWK 490
++W+
Sbjct: 1486ASFGKIWR 1509
>TC83621 similar to PIR|T01506|T01506 probable hexose transport protein
T10M13.6 - Arabidopsis thaliana, partial (32%)
Length = 636
Score = 204 bits (518), Expect = 9e-53
Identities = 97/170 (57%), Positives = 126/170 (74%)
Frame = +2
Query: 330 VNVVATCVSIYGVDKWGRRALFLEGGAQMLICQVAVAAAIGAKFGTSGNPGNLPEWYAIV 389
V +++T +SI VD+ GRR L + GG QM+ICQV VA +G KFG + L + Y++
Sbjct: 2 VLLLSTFISIAIVDRLGRRPLLISGGIQMIICQVIVAIILGIKFGDNQE---LSKGYSLS 172
Query: 390 VVLFICIYVAGFAWSWGPLGWLVPSEIFPLEIRSAAQSVNVSVNMLFTFLVAQVFLIMLC 449
VV+ IC++V F WSWGPLGW VPSEIFPLEIRSA QS+ V+VN+LFTF++AQ FL +LC
Sbjct: 173 VVVAICLFVLAFGWSWGPLGWTVPSEIFPLEIRSAGQSITVAVNLLFTFIIAQTFLSLLC 352
Query: 450 HMKFGLFLFFAFFVLVMSIYVFFLLPETKGIPIEEMDRVWKSHPFWSRFV 499
KFG+FLFFA ++ +M+I+V LPETKGIPIEEM +WK H FW R +
Sbjct: 353 SFKFGIFLFFAGWITIMTIFVVLFLPETKGIPIEEMAIMWKKHWFWKRIL 502
>TC78321 similar to GP|21689841|gb|AAM67564.1 unknown protein {Arabidopsis
thaliana}, partial (80%)
Length = 2093
Score = 177 bits (449), Expect = 9e-45
Identities = 115/337 (34%), Positives = 177/337 (52%), Gaps = 5/337 (1%)
Frame = +2
Query: 21 PFVTITCIVAAMGGLIFGYDIGISGGVTSMDPFLKKFFPAVYRKKNKDKSTNQYCQYDSQ 80
P+V A +GGL+FGYD G+ G +++ FPAV +K ++
Sbjct: 215 PYVLRLAFSAGIGGLLFGYDTGVISGALL---YIRDEFPAVEKKTWLQEA---------- 355
Query: 81 TLTMFTSSLYLAALLSSLVASTITRRFGRKLSMLFGGLLFLVGALINGFANHVWMLIVGR 140
S+ A++ + + I RFGRK+S++ LFL+G++I A + LIVGR
Sbjct: 356 ----IVSTAIAGAIIGAAIGGWINDRFGRKVSIIVADTLFLLGSIILAAAPNPATLIVGR 523
Query: 141 ILLGFGIGFANQAVPLYLSEMAPYKYRGALNIGFQLSITIGILVANVLNYFFAKIKGGWG 200
+ +G G+G A+ A PLY+SE +P + RGAL IT G ++ ++N F K G W
Sbjct: 524 VFVGLGVGMASMASPLYISEASPTRVRGALVSLNSFLITGGQFLSYLINLAFTKAPGTWR 703
Query: 201 WRLSLGGAMVPALIITIGSLVLPDTPNSMIERGDRDGAKAQLKRIRGIEDVDEEFNDLVA 260
W LG A PA+I + L LP++P + +G + AK LK+I +ED D E L
Sbjct: 704 W--MLGVAAAPAVIQIVLMLSLPESPRWLYRKGKEEEAKVILKKIYEVEDYDNEIQALKE 877
Query: 261 ASEASMQVENPWRNLLQ----RKYRPQLTMAVLIPFFQQFTGINVIMFYAPVLFNSIGFK 316
+ E ++ E +++Q R L V + FFQQFTGIN +M+Y+P + GF
Sbjct: 878 SVEMELK-ETEKISIMQLVKTTSVRRGLYAGVGLAFFQQFTGINTVMYYSPSIVQLAGFA 1054
Query: 317 DD-ASLMSAVITGVVNVVATCVSIYGVDKWGRRALFL 352
+L+ ++IT +N + +SIY +DK GR+ L L
Sbjct: 1055SKRTALLLSLITSGLNAFGSILSIYFIDKTGRKKLAL 1165
Score = 61.6 bits (148), Expect = 7e-10
Identities = 35/128 (27%), Positives = 63/128 (48%), Gaps = 5/128 (3%)
Frame = +2
Query: 375 TSGNPGNLPEWYAIVVVLFICIYVAGFAWSWGPLGWLVPSEIFPLEIRSAAQSVNVSVNM 434
T G P N + + +L + +Y+ F+ G + W+V SEI+PL R + +
Sbjct: 1460 TKGCPSN----FGWIAILALALYIIFFSPGMGTVPWVVNSEIYPLRYRGICGGIASTTVW 1627
Query: 435 LFTFLVAQVFLIMLCHM-KFGLFLFFAFFVLVMSIYVFFLLPETKGIPIEEMDRVWKSH- 492
+ +V+Q FL + + F+ FA +V +V +PETKG+P+EE++ + +
Sbjct: 1628 VSNLVVSQSFLSLTVAIGPAWTFMIFAIIAIVAIFFVIIFVPETKGVPMEEVESMLEKRF 1807
Query: 493 ---PFWSR 497
FW +
Sbjct: 1808 VQIKFWKK 1831
>TC78543 similar to PIR|T14545|T14545 probable sugar transporter protein -
beet, partial (93%)
Length = 1653
Score = 173 bits (438), Expect = 2e-43
Identities = 110/410 (26%), Positives = 208/410 (49%), Gaps = 2/410 (0%)
Frame = +1
Query: 83 TMFTSSLYLAALLSSLVASTITRRFGRKLSMLFGGLLFLVGALINGFANHVWMLIVGRIL 142
++F S + A++ ++ + I GRK S++ + ++G L+ FAN L +GR+L
Sbjct: 268 SLFGSLSNVGAMVGAIASGQIAEYMGRKGSLMIASIPNIIGWLMISFANDSSFLYMGRLL 447
Query: 143 LGFGIGFANQAVPLYLSEMAPYKYRGALNIGFQLSITIGILVANVLNYFFAKIKGGWGWR 202
GFG+G + VP+Y++E++P RG+L QLS+T+GI++A +L F WR
Sbjct: 448 EGFGVGIISYTVPVYIAEISPQNLRGSLVSVNQLSVTLGIMLAYLLGLFVE-------WR 606
Query: 203 LSLGGAMVPALIITIGSLVLPDTPNSMIERGDRDGAKAQLKRIRGIE-DVDEEFNDL-VA 260
++P ++ G +P++P + + G + + L+ +RG E D+ E N++ A
Sbjct: 607 FLAILGIIPCTLLIPGLFFIPESPRWLAKMGMTEEFENSLQVLRGFETDISVEVNEIKTA 786
Query: 261 ASEASMQVENPWRNLLQRKYRPQLTMAVLIPFFQQFTGINVIMFYAPVLFNSIGFKDDAS 320
+ A+ + + L QR+Y L + + + QQ +GIN ++FY+ +F + G +S
Sbjct: 787 VASANRRTTVRFSELKQRRYWLPLMIGIGLLVLQQLSGINGVLFYSSTIFQNAGI--SSS 960
Query: 321 LMSAVITGVVNVVATCVSIYGVDKWGRRALFLEGGAQMLICQVAVAAAIGAKFGTSGNPG 380
++ G V V+AT ++++ DK GRR L + + M + + V+ + K
Sbjct: 961 DVATFGVGAVQVLATTLTLWLADKSGRRLLLIVSSSAMTLSLLVVSISFYLKDYYISADS 1140
Query: 381 NLPEWYAIVVVLFICIYVAGFAWSWGPLGWLVPSEIFPLEIRSAAQSVNVSVNMLFTFLV 440
+L +++ V + + V F+ G + W++ SEI P+ I+ A S N F++L+
Sbjct: 1141SLYGILSLLSVAGVVVMVIAFSLGMGAMPWIIMSEILPINIKGLAGSFATLANWFFSWLI 1320
Query: 441 AQVFLIMLCHMKFGLFLFFAFFVLVMSIYVFFLLPETKGIPIEEMDRVWK 490
++L G F + +V +PETKG +EE+ + ++
Sbjct: 1321TLTANLLLDWSSGGTFTIYTVVCAFTVGFVAIWVPETKGKTLEEIQQFFR 1470
>TC87126 similar to GP|15724240|gb|AAL06513.1 At1g75220/F22H5_6 {Arabidopsis
thaliana}, partial (79%)
Length = 1524
Score = 171 bits (434), Expect = 5e-43
Identities = 111/405 (27%), Positives = 202/405 (49%), Gaps = 2/405 (0%)
Frame = +3
Query: 93 ALLSSLVASTITRRFGRKLSMLFGGLLFLVGALINGFANHVWMLIVGRILLGFGIGFANQ 152
A++ ++ + I GRK S++ + ++G L FA L +GR+L GFG+G +
Sbjct: 9 AMVGAIASGQIAEYVGRKGSLMIASIPNIIGWLAISFAKDSSFLFMGRLLEGFGVGIISY 188
Query: 153 AVPLYLSEMAPYKYRGALNIGFQLSITIGILVANVLNYFFAKIKGGWGWRLSLGGAMVPA 212
VP+Y++E+AP RG+L QLS+TIGI++A +L F WR+ ++P
Sbjct: 189 VVPVYIAEIAPENMRGSLGSVNQLSVTIGIMLAYLLGLFA-------NWRVLAILGILPC 347
Query: 213 LIITIGSLVLPDTPNSMIERGDRDGAKAQLKRIRGIE-DVDEEFNDL-VAASEASMQVEN 270
++ G +P++P + + G + + L+ +RG + D+ E +++ A + +
Sbjct: 348 TVLIPGLFFIPESPRWLAKMGMMEEFETSLQVLRGFDTDISVEVHEIKKAVASNGKRATI 527
Query: 271 PWRNLLQRKYRPQLTMAVLIPFFQQFTGINVIMFYAPVLFNSIGFKDDASLMSAVITGVV 330
+ +L +++Y L++ + + QQ +GIN ++FY+ +F + G +S + V G +
Sbjct: 528 RFADLQRKRYWFPLSVGIGLLVLQQLSGINGVLFYSTSIFANAGI--SSSNAATVGLGAI 701
Query: 331 NVVATCVSIYGVDKWGRRALFLEGGAQMLICQVAVAAAIGAKFGTSGNPGNLPEWYAIVV 390
V+AT V+ + VDK GRR L + + M + V+ A + G I+
Sbjct: 702 QVIATGVATWLVDKSGRRVLLIISSSLMTASLLVVSIAFYLE-GVVEKDSQYFSILGIIS 878
Query: 391 VLFICIYVAGFAWSWGPLGWLVPSEIFPLEIRSAAQSVNVSVNMLFTFLVAQVFLIMLCH 450
V+ + + V GF+ GP+ WL+ SEI P+ I+ A S N L +++ ++L
Sbjct: 879 VVGLVVMVIGFSLGLGPIPWLIMSEILPVNIKGLAGSTATMANWLVAWIITMTANLLLTW 1058
Query: 451 MKFGLFLFFAFFVLVMSIYVFFLLPETKGIPIEEMDRVWKSHPFW 495
G FL + ++ +PETKG +EE+ + FW
Sbjct: 1059SSGGTFLIYTVVAAFTVVFTSLWVPETKGRTLEEIQFSLR*SMFW 1193
>AL368995 similar to GP|2104547|gb| AGAA.1 {Arabidopsis thaliana}, partial
(57%)
Length = 447
Score = 150 bits (379), Expect = 1e-36
Identities = 71/127 (55%), Positives = 99/127 (77%)
Frame = +1
Query: 14 EYPGNLTPFVTITCIVAAMGGLIFGYDIGISGGVTSMDPFLKKFFPAVYRKKNKDKSTNQ 73
+Y G +T V + CIVAA GG +FGYD+GISGGVTSMD FL KFFP+VY++K N
Sbjct: 70 QYNGRVTVHVILACIVAATGGSLFGYDVGISGGVTSMDDFLLKFFPSVYKQK-MHAHENN 246
Query: 74 YCQYDSQTLTMFTSSLYLAALLSSLVASTITRRFGRKLSMLFGGLLFLVGALINGFANHV 133
YC+Y++Q L FTS LY++ L++SLVASTITR++GRK+S++ GG+ FL+G+++N A ++
Sbjct: 247 YCKYNNQVLAAFTSVLYISGLVASLVASTITRKYGRKISIIVGGISFLIGSILNAAAANL 426
Query: 134 WMLIVGR 140
MLI+GR
Sbjct: 427 GMLIIGR 447
>BI310242 similar to GP|347853|gb|A glucose transporter {Saccharum hybrid
cultivar H65-7052}, partial (36%)
Length = 519
Score = 144 bits (362), Expect = 1e-34
Identities = 63/99 (63%), Positives = 79/99 (79%)
Frame = -1
Query: 397 YVAGFAWSWGPLGWLVPSEIFPLEIRSAAQSVNVSVNMLFTFLVAQVFLIMLCHMKFGLF 456
+V F WSWGPLGW VPSEIFPLEIRSA QS+ V+VN+LFTF++AQ FL +LC K+G+F
Sbjct: 519 FVLAFGWSWGPLGWTVPSEIFPLEIRSAGQSITVAVNLLFTFIIAQAFLSLLCFFKYGIF 340
Query: 457 LFFAFFVLVMSIYVFFLLPETKGIPIEEMDRVWKSHPFW 495
LFFA + +M+++VF LPETKGIPIEEM + + H FW
Sbjct: 339 LFFAGWTALMTLFVFLFLPETKGIPIEEMSILLRKHWFW 223
>AL368310 weakly similar to GP|16945177|em STP5 protein {Arabidopsis
thaliana}, partial (22%)
Length = 477
Score = 143 bits (360), Expect = 2e-34
Identities = 74/144 (51%), Positives = 97/144 (66%), Gaps = 5/144 (3%)
Frame = +2
Query: 1 MAGGGIPIGGGNKEYP-----GNLTPFVTITCIVAAMGGLIFGYDIGISGGVTSMDPFLK 55
MAGG +P+ G LT + ITCIVAA GGL++GYD+G+SGGVT+M PFL+
Sbjct: 47 MAGGVLPVDSTPVAVTAINIGGKLTLSIIITCIVAASGGLLYGYDLGVSGGVTTMVPFLQ 226
Query: 56 KFFPAVYRKKNKDKSTNQYCQYDSQTLTMFTSSLYLAALLSSLVASTITRRFGRKLSMLF 115
KFFP + RK N YC YDSQ LT+FTSSLYLA L+SS+ AS +T +GR+ ++
Sbjct: 227 KFFPDILRKA-ASAEVNMYCVYDSQILTLFTSSLYLAGLVSSIAASXVTAAYGRRNVIII 403
Query: 116 GGLLFLVGALINGFANHVWMLIVG 139
GG LF+ G ING + ++ MLI+G
Sbjct: 404 GGALFIAGGAINGGSENIPMLILG 475
>TC83213 weakly similar to GP|16945177|emb|CAC69071. STP5 protein
{Arabidopsis thaliana}, partial (18%)
Length = 563
Score = 139 bits (351), Expect = 2e-33
Identities = 58/98 (59%), Positives = 79/98 (80%)
Frame = +1
Query: 403 WSWGPLGWLVPSEIFPLEIRSAAQSVNVSVNMLFTFLVAQVFLIMLCHMKFGLFLFFAFF 462
WSWGPL WL+PSEIFP++IR+ QS+ V+V + F+++Q FL MLCHMKFG F+F+AF+
Sbjct: 1 WSWGPLTWLIPSEIFPVKIRTTGQSIAVAVQFIIIFVLSQTFLTMLCHMKFGAFVFYAFW 180
Query: 463 VLVMSIYVFFLLPETKGIPIEEMDRVWKSHPFWSRFVE 500
V+VM+++V F LPETKGIP+E M +W H FWSR+V+
Sbjct: 181 VIVMTMFVIFFLPETKGIPLESMYTIWGRHWFWSRYVK 294
>AW586674 weakly similar to SP|Q10710|STA_ Sugar carrier protein A. [Castor
bean] {Ricinus communis}, partial (21%)
Length = 531
Score = 130 bits (327), Expect = 1e-30
Identities = 63/126 (50%), Positives = 89/126 (70%), Gaps = 6/126 (4%)
Frame = +3
Query: 1 MAGGGIPIGGGN------KEYPGNLTPFVTITCIVAAMGGLIFGYDIGISGGVTSMDPFL 54
M G P GGG ++Y G +T V I CIVAA GG +FGYD+GISGGV SMD FL
Sbjct: 156 MEDGSSPTGGGTVDKGRAEQYKGRVTVHVIIACIVAATGGSLFGYDVGISGGVASMDDFL 335
Query: 55 KKFFPAVYRKKNKDKSTNQYCQYDSQTLTMFTSSLYLAALLSSLVASTITRRFGRKLSML 114
+ FFPAVY+ K + N YC+Y++Q ++ FTS+LY++ L++S++A+ ITRR+GR+ S++
Sbjct: 336 QNFFPAVYKHK-LEAHENNYCKYNNQGISAFTSTLYISGLVASIIAAPITRRYGRRTSII 512
Query: 115 FGGLLF 120
GG+ F
Sbjct: 513 IGGINF 530
>TC79361 similar to GP|17380890|gb|AAL36257.1 putative membrane transporter
protein {Arabidopsis thaliana}, partial (55%)
Length = 949
Score = 124 bits (312), Expect = 7e-29
Identities = 84/281 (29%), Positives = 139/281 (48%), Gaps = 3/281 (1%)
Frame = +1
Query: 21 PFVTITCIVAAMGGLIFGYDIGISGGVTSMDPFLKKFFPAVYRKKNKDKSTNQYCQYDSQ 80
P++ VA +GGL+FGYD G+ G ++K FP V R N + T
Sbjct: 163 PYILGLAAVAGIGGLLFGYDTGVISGALL---YIKDDFPQV-RNSNFLQET--------- 303
Query: 81 TLTMFTSSLYLAALLSSLVASTITRRFGRKLSMLFGGLLFLVGALINGFANHVWMLIVGR 140
S A++ + + +GRK + L ++F++GA++ A ++LI GR
Sbjct: 304 ----IVSMAIAGAIVGAAFGGWLNDAYGRKKATLLADVIFILGAILMAAAPDPYVLIAGR 471
Query: 141 ILLGFGIGFANQAVPLYLSEMAPYKYRGALNIGFQLSITIGILVANVLNYFFAKIKGGWG 200
+L+G G+G A+ P+Y++E+AP + RG+L L IT G V+ ++N F ++ G W
Sbjct: 472 LLVGLGVGIASVTAPVYIAEVAPSEIRGSLVSTNVLMITGGQFVSYLVNLVFTQVPGTWR 651
Query: 201 WRLSLGGAMVPALIITIGSLVLPDTPNSMIERGDRDGAKAQLKRIRGIEDVDEEFNDLVA 260
W L + G VPALI I L LP++P + + ++ A + +I + +++E + L A
Sbjct: 652 WMLGVSG--VPALIQFICMLFLPESPRWLFIKNRKNEAVDVISKIYDLSRLEDEIDFLTA 825
Query: 261 ASEASMQVENP---WRNLLQRKYRPQLTMAVLIPFFQQFTG 298
SE Q + W ++ R + FQQFTG
Sbjct: 826 QSEQERQRRSTIKFWHVFRSKETRLAFLDGGGLLAFQQFTG 948
>TC81556 similar to GP|2688830|gb|AAB88879.1| putative sugar transporter
{Prunus armeniaca}, partial (59%)
Length = 1074
Score = 123 bits (309), Expect = 1e-28
Identities = 79/294 (26%), Positives = 150/294 (50%), Gaps = 3/294 (1%)
Frame = +1
Query: 201 WRLSLGGAMVPALIITIGSLVLPDTPNSMIERGDRDGAKAQLKRIRGIEDVDEEFNDLVA 260
WR G A+VP++++ +G + P++P + ++G A+ +K + G E V DL
Sbjct: 19 WRTMFGIAVVPSILLALGMAISPESPRWLFQQGKIAEAEKAIKTLYGKERVATVMYDLRT 198
Query: 261 ASEASMQVENPWRNLLQRKYRPQLTMAVLIPFFQQFTGINVIMFYAPVLFNSIGFKDDAS 320
AS+ S + E W +L +Y +++ + FQQ GIN +++Y+ +F S G D +
Sbjct: 199 ASQGSSEPEAGWFDLFSSRYWKVVSVGAALFLFQQLAGINAVVYYSTSVFRSAGIASDVA 378
Query: 321 LMSAVITGVVNVVATCVSIYGVDKWGRRALFLEGGAQMLICQVAVAAAIGAKFGTSGNPG 380
++ + G NV+ T ++ +DK GR++L + + M + ++ + K
Sbjct: 379 --ASALVGASNVIGTAIASSLMDKQGRKSLLITSFSGMAASMLLLSLSFTWKV------- 531
Query: 381 NLPEWYAIVVVLFICIYVAGFAWSWGPLGWLVPSEIFPLEIRSAAQSVNVSVNMLFTFLV 440
L + + VL +YV F+ GP+ L+ EIF IR+ A S+++ + + F++
Sbjct: 532 -LAPYSGTLAVLGTVLYVLSFSLGAGPVPALLLPEIFASRIRAKAVSLSLGTHWISNFVI 708
Query: 441 AQVFLIMLCHMKFGL---FLFFAFFVLVMSIYVFFLLPETKGIPIEEMDRVWKS 491
FL ++ KFG+ +L F+ ++ +Y+ + ETKG +EE++R S
Sbjct: 709 GLYFLSVV--NKFGISSVYLGFSAVCVLAVLYIAGNVVETKGRSLEEIERALSS 864
>BQ751130 weakly similar to GP|15289796|dbj putative monosaccharide
transporter 3 {Oryza sativa (japonica cultivar-group)},
partial (7%)
Length = 795
Score = 110 bits (275), Expect = 1e-24
Identities = 77/253 (30%), Positives = 125/253 (48%), Gaps = 9/253 (3%)
Frame = +3
Query: 25 ITCIVAAMGGLIFGYDIGISGGVTSMDPFLKKFF-PAVYRKKNKDKSTNQYCQYDSQTLT 83
+ C A+ GG+ FGYD G GV + F++ P +S
Sbjct: 45 VICAFASFGGIFFGYDSGYINGVLGANQFIEHVEGPGAEAISESHQS------------- 185
Query: 84 MFTSSLYLAALLSSLVASTITRRFGRKLSMLFGGLLFLVGALIN---GFANHVWMLIVGR 140
+ S L +L+A ++ GRK +++ G ++++G LI G + ++ GR
Sbjct: 186 LIVSILSCGTFFGALIAGDVSDWIGRKWTVIIGCGIYMIGVLIQMFTGMGAPLGAIVAGR 365
Query: 141 ILLGFGIGFANQAVPLYLSEMAPYKYRGALNIGFQLSITIGILVANVLNYFFAKIKGGWG 200
++ G G+GF + V LY+SE+ P K RGAL G+Q ITIG+L+A + Y K +G G
Sbjct: 366 LIAGIGVGFESAIVILYMSEICPKKVRGALVAGYQFCITIGLLLAACVVY-ATKDRGDTG 542
Query: 201 -WRLSLGGAMVPALIITIGSLVLPDTPNSMIERGDRDGAKAQLKRIRG----IEDVDEEF 255
+R+ + ALI+ G L+LP++P +++G A A L R+RG E + E
Sbjct: 543 VYRIPIAIQFPWALILGGGLLLLPESPRFFVKKGRIADATAALSRLRGQPKESEYIQVEL 722
Query: 256 NDLVAASEASMQV 268
++VA E Q+
Sbjct: 723 AEIVANEEYERQL 761
>BQ136503 weakly similar to PIR|T05156|T051 probable glucose transport
protein F18E5.100 - Arabidopsis thaliana, partial (15%)
Length = 490
Score = 105 bits (263), Expect = 3e-23
Identities = 43/78 (55%), Positives = 59/78 (75%)
Frame = +3
Query: 426 QSVNVSVNMLFTFLVAQVFLIMLCHMKFGLFLFFAFFVLVMSIYVFFLLPETKGIPIEEM 485
QS+ V+VNM TF +AQ+F MLCH KFGLF+FF FV++M+ +++ PETKG+P+EEM
Sbjct: 9 QSITVAVNMTSTFFIAQLFTEMLCHFKFGLFIFFGCFVILMTFFIYKFFPETKGVPLEEM 188
Query: 486 DRVWKSHPFWSRFVEHGD 503
VW+ HPFW +F+E D
Sbjct: 189 HMVWRKHPFWGKFLEAED 242
>TC82331 weakly similar to EGAD|120604|128852 hypothetical protein M01F1.5
{Caenorhabditis elegans}, partial (10%)
Length = 848
Score = 99.4 bits (246), Expect = 3e-21
Identities = 56/201 (27%), Positives = 109/201 (53%), Gaps = 2/201 (0%)
Frame = +3
Query: 28 IVAAMGGLIFGYDIGISGGVTSMDPFLKKFFPAVYRKKNKDKSTNQYCQYDSQTLTMFTS 87
+V ++GG +FGYD G + F ++F + N + S ++ S
Sbjct: 255 VVTSIGGFLFGYDTGQISSMLLFTDFRQRFATG-------PEDANGVKTWVSIIQSLMVS 413
Query: 88 SLYLAALLSSLVASTITRRFGRKLSMLFGGLLFLVGALINGFANHVWM-LIVGRILLGFG 146
+ + L+ +L + +GR+ S+ FG ++F++G ++ A W+ +++GR + G G
Sbjct: 414 LMSIGTLIGALSGAYTADWWGRRKSLSFGVVVFVIGNIVQITAMDTWVHMMMGRFIAGLG 593
Query: 147 IGFANQAVPLYLSEMAPYKYRGALNIGFQLSITIGILVANVLNYFFAKI-KGGWGWRLSL 205
+G + VP++ SE +P + RGA+ +QL IT GIL++N++N+ I + WR+ +
Sbjct: 594 VGNLSVGVPMFQSECSPKEIRGAVVASYQLMITFGILISNLINFGVRNIQESDASWRIVI 773
Query: 206 GGAMVPALIITIGSLVLPDTP 226
G + A+ + +G LV+P++P
Sbjct: 774 GLGIAFAMPLGLGILVVPESP 836
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.327 0.143 0.444
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,551,234
Number of Sequences: 36976
Number of extensions: 245372
Number of successful extensions: 2374
Number of sequences better than 10.0: 116
Number of HSP's better than 10.0 without gapping: 2245
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2302
length of query: 518
length of database: 9,014,727
effective HSP length: 100
effective length of query: 418
effective length of database: 5,317,127
effective search space: 2222559086
effective search space used: 2222559086
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 61 (28.1 bits)
Medicago: description of AC146792.3


