

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146790.7 + phase: 0
(546 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
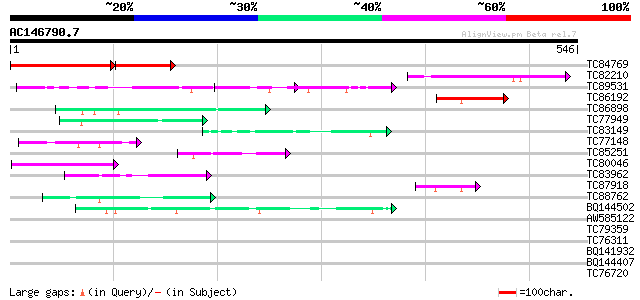
Score E
Sequences producing significant alignments: (bits) Value
TC84769 similar to PIR|E96665|E96665 protein F22C12.16 [imported... 207 6e-84
TC82210 weakly similar to GP|15450527|gb|AAK96556.1 AT5g21160/T1... 97 1e-20
TC89531 similar to GP|19034049|gb|AAL83681.1 putative extensin {... 75 8e-14
TC86192 weakly similar to GP|20161211|dbj|BAB90138. RNA-binding ... 55 5e-08
TC86898 similar to GP|18376007|emb|CAB91741. probable ATP citrat... 47 1e-05
TC77949 weakly similar to GP|11762218|gb|AAG40387.1 AT4g27520 {A... 47 1e-05
TC83149 similar to GP|13561982|gb|AAK30594.1 flagelliform silk p... 45 7e-05
TC77148 ENOD20 44 2e-04
TC85251 homologue to PIR|JQ2128|JQ2128 metallothionein - soybean... 44 2e-04
TC80046 similar to SP|P22699|GUN6_DICDI Endoglucanase precursor ... 44 2e-04
TC83962 similar to PIR|B86289|B86289 hypothetical protein AAF719... 43 3e-04
TC87918 weakly similar to GP|15810397|gb|AAL07086.1 unknown prot... 43 4e-04
TC88762 similar to GP|22136928|gb|AAM91808.1 unknown protein {Ar... 42 5e-04
BQ144502 weakly similar to GP|6523547|emb hydroxyproline-rich gl... 42 5e-04
AW585122 41 0.001
TC79359 weakly similar to PIR|S49915|S49915 extensin-like protei... 40 0.002
TC76311 homologue to GP|3204132|emb|CAA07235.1 extensin {Cicer a... 40 0.002
BQ141932 weakly similar to GP|6523547|emb hydroxyproline-rich gl... 40 0.002
BQ144407 weakly similar to GP|6523547|emb| hydroxyproline-rich g... 40 0.002
TC76720 similar to GP|11121502|emb|CAC14888. putative extensin {... 40 0.002
>TC84769 similar to PIR|E96665|E96665 protein F22C12.16 [imported] -
Arabidopsis thaliana, partial (4%)
Length = 723
Score = 207 bits (527), Expect(2) = 6e-84
Identities = 102/102 (100%), Positives = 102/102 (100%)
Frame = +2
Query: 1 MVTTQNNHSPTSVPSPAPPSAETNNPNFIRKNLPSPWAQVVRGGSGWDTESQSQSPTGIH 60
MVTTQNNHSPTSVPSPAPPSAETNNPNFIRKNLPSPWAQVVRGGSGWDTESQSQSPTGIH
Sbjct: 245 MVTTQNNHSPTSVPSPAPPSAETNNPNFIRKNLPSPWAQVVRGGSGWDTESQSQSPTGIH 424
Query: 61 QSLPSSSSSSSSSLTTVDQPPPSDDSPKAAVVSSSPKAVVVS 102
QSLPSSSSSSSSSLTTVDQPPPSDDSPKAAVVSSSPKAVVVS
Sbjct: 425 QSLPSSSSSSSSSLTTVDQPPPSDDSPKAAVVSSSPKAVVVS 550
Score = 122 bits (306), Expect(2) = 6e-84
Identities = 57/57 (100%), Positives = 57/57 (100%)
Frame = +3
Query: 103 SPLPAPMEKNSNTIASDGGDGGNAGGSKKPAWNKQPLNGVAVEIGPVMGAESWPALS 159
SPLPAPMEKNSNTIASDGGDGGNAGGSKKPAWNKQPLNGVAVEIGPVMGAESWPALS
Sbjct: 552 SPLPAPMEKNSNTIASDGGDGGNAGGSKKPAWNKQPLNGVAVEIGPVMGAESWPALS 722
>TC82210 weakly similar to GP|15450527|gb|AAK96556.1 AT5g21160/T10F18_190
{Arabidopsis thaliana}, partial (24%)
Length = 1436
Score = 97.4 bits (241), Expect = 1e-20
Identities = 62/167 (37%), Positives = 90/167 (53%), Gaps = 10/167 (5%)
Frame = +2
Query: 384 PFRGMPVFPHMPSPATFFPAAESSLSNVIVNQIDYYFSDINLANDEFLKSNMDEEGWVPI 443
P+ + P P+P T SL I+ QI+YYFSD NL ND +L MD++GWVPI
Sbjct: 44 PYPPVNSAPQSPTPET------QSLRASILKQIEYYFSDENLHNDRYLIGLMDDQGWVPI 205
Query: 444 TLIANFPRVKNLTSNIQLILDSMRNSNVVDVQGDKLRRRN----WIDRLTS------AQV 493
+ +A+F RVK ++++I I+D ++NS+ V+VQ DK+R+RN WI + AQV
Sbjct: 206 STVADFKRVKRMSTDIPFIVDVLQNSDNVEVQDDKIRKRNNWSKWIQTSSGNSGSSVAQV 385
Query: 494 QADAGSVSPIESRDNSFTADFQTITLDKTTKDEGESSRQSQLSNGSD 540
Q D S S NS T +T + T ++ S N S+
Sbjct: 386 QQDQHVESTANSCQNSDTVVDKTKESSEATLNDSAHDSTSTEQNQSN 526
>TC89531 similar to GP|19034049|gb|AAL83681.1 putative extensin {Oryza
sativa (japonica cultivar-group)}, partial (8%)
Length = 1184
Score = 74.7 bits (182), Expect = 8e-14
Identities = 83/285 (29%), Positives = 116/285 (40%), Gaps = 12/285 (4%)
Frame = +1
Query: 7 NHSPTSVPSPAPPSAETNNPNFIRKNLPSPWAQVVRGGSGWDTESQSQSPTGIHQSLPSS 66
NHS S+ S+ + N + LP PW QVVRG ++ES + P S
Sbjct: 181 NHSLLSMEMIGNHSSNHSPDNLQSRRLP-PWNQVVRG----ESESIAAVPA-------VS 324
Query: 67 SSSSSSSLTTVDQPPPSDDSPKAAVVSSSPKAVVVSSPLPAPMEKNSNTIASDGGDGGNA 126
S S + T P DDS A V+S + +D G N
Sbjct: 325 LSEESFPIVTA----PVDDSTSAEVISDN----------------------ADNGGERNG 426
Query: 127 GGSKKPAWNKQPLNGVAVEIGPVMGAESWPALSESAKIPGKLPPESS-----SSKIAPPA 181
G K+PAWN+ NG E+ PVM A SWPALS+SA+ G ESS S ++P
Sbjct: 427 GTGKRPAWNRSSGNGGVSEVQPVMDAHSWPALSDSAR--GSTKSESSKGLLDGSSVSPWQ 600
Query: 182 AVDGSPSTSQGPIISHSPQRQGSTSNTKSSSMANNNLPNRPRPMRRISGSNIGPCPAQSS 241
++ +PS+ S + T +A N N M + G A S
Sbjct: 601 GMESTPSSLYAKTTS-----EIM*M*TIWHQLARNQ-SNTTVQMHLLMG-------ATHS 741
Query: 242 LSNPPTP------PPLPPYPVYQLPPAV-SYPNMLPSIPDSSPRD 279
NP PL P ++ LP + S P S+P+++PR+
Sbjct: 742 SLNPKLR*LQRDLTPLLPETIHSLPETIHSLPETTHSLPETTPRE 876
Score = 63.9 bits (154), Expect = 1e-10
Identities = 56/181 (30%), Positives = 79/181 (42%), Gaps = 6/181 (3%)
Frame = +3
Query: 198 SPQRQGSTSNTKSSSMANNNLPNRPRPMRRISGSNIGPCPAQSSLSNPPTPPPLPPYPVY 257
+P RQ S + S++ +N + P I+ + + +S S T P
Sbjct: 672 APTRQKSIKHNSSNASSNGGHTQQSEPQVTIAATG-----SHTSSSRDHTQSPRDH---- 824
Query: 258 QLPPAVSYPNMLPSIPDSSPRDHHRNNNW--DARPFVGGSSRRGHFGSHPRGDGSYHQNS 315
P SPRDH + + + P S R + G H RGDGS+H ++
Sbjct: 825 ---------TQSPRDHAQSPRDHTQRSGFVPSDHPQQRNSFRHRNGGPHQRGDGSHHHHN 977
Query: 316 YSSRREHD---RGNYANTRDAHAPQPRMPPRGILRPP-PPSTAAFLGPQTIGPFPAPVAY 371
Y +RR+ D R NY N RD H P PR+ PR I+RP PP++A TI F P
Sbjct: 978 YGNRRDQDWNSRRNY-NGRDMHVP-PRVSPR-IIRPSLPPNSAPLHSSTTIAAFWWPHGI 1148
Query: 372 P 372
P
Sbjct: 1149P 1151
>TC86192 weakly similar to GP|20161211|dbj|BAB90138. RNA-binding
protein-like {Oryza sativa (japonica cultivar-group)},
partial (34%)
Length = 1617
Score = 55.5 bits (132), Expect = 5e-08
Identities = 28/71 (39%), Positives = 47/71 (65%), Gaps = 2/71 (2%)
Frame = +2
Query: 412 IVNQIDYYFSDINLANDEFLKS--NMDEEGWVPITLIANFPRVKNLTSNIQLILDSMRNS 469
I+ Q++YYFSD NL D+++ S ++EG+VPI IA+F ++K LT + I +++ S
Sbjct: 356 IIKQVEYYFSDENLPTDKYMLSLVKRNKEGFVPIPAIASFRKMKRLTRDHLFIAAALKES 535
Query: 470 NVVDVQGDKLR 480
+++ V GD R
Sbjct: 536 SLLVVSGDGKR 568
>TC86898 similar to GP|18376007|emb|CAB91741. probable ATP citrate lyase
subunit 2 {Neurospora crassa}, complete
Length = 2389
Score = 47.4 bits (111), Expect = 1e-05
Identities = 59/222 (26%), Positives = 87/222 (38%), Gaps = 15/222 (6%)
Frame = +3
Query: 45 SGWDTESQSQSPTGIHQSLPSSSS--SSSSSLTTVDQP--------PPSDDSPKAAVVSS 94
+ W T S + +P+ S P+S S SSSS T+ QP P S P + VSS
Sbjct: 1176 TSWSTSSAACTPSTSTASSPTSRSTPSSSSPTRTLPQPPCTSWILLPSSTRPPTLSAVSS 1355
Query: 95 SP-KAVVVSS---PLPAPMEKNSNTIASDGGDGGNAGGSKKPAWNKQPLNGVAVEIGPVM 150
P A+ + S LP P K S ++ + P + + P
Sbjct: 1356 GPLPALPLPSA*PTLPLPTAKRSTSMRARRWSSPLLSVVS*PRRRLTSPTLMPRPVPPSS 1535
Query: 151 GAESWPALSESAKIPGKLPPESSSSKIAPPAAVDGSPSTSQGPIISHSPQRQGSTSNTKS 210
S P P +PP S+ + PPA+ SP+T ++ SP R +T S
Sbjct: 1536 LLSSTPTAVSGLSSPVVVPPSSTPTPSPPPASPTSSPTTVSTLVLPPSP-RPTTTPVLSS 1712
Query: 211 SSMANNNLPNRPRPMRRISGSNIGP-CPAQSSLSNPPTPPPL 251
+S + P R R + P PA S +S+ P+ L
Sbjct: 1713 TSCSALL*PRRARSSLSVVVLPTSPTSPAHSRVSSGPSATTL 1838
Score = 31.6 bits (70), Expect = 0.82
Identities = 37/119 (31%), Positives = 49/119 (41%), Gaps = 24/119 (20%)
Frame = +3
Query: 9 SPTSVPSPAPPSAETNNPNFIRKNLPSPWAQVVRGGSGWDT-----ESQSQSPTGIH--- 60
SPT +P P PPS+ ++ L SP VV S T S + SPT +
Sbjct: 1497 SPTLMPRPVPPSSLLSSTPTAVSGLSSP---VVVPPSSTPTPSPPPASPTSSPTTVSTLV 1667
Query: 61 -------QSLPSSSSSSSSSL---------TTVDQPPPSDDSPKAAVVSSSPKAVVVSS 103
+ P SS+S S+L +V P S SP + VSS P A +SS
Sbjct: 1668 LPPSPRPTTTPVLSSTSCSALL*PRRARSSLSVVVLPTSPTSPAHSRVSSGPSATTLSS 1844
>TC77949 weakly similar to GP|11762218|gb|AAG40387.1 AT4g27520 {Arabidopsis
thaliana}, partial (34%)
Length = 1297
Score = 47.4 bits (111), Expect = 1e-05
Identities = 43/148 (29%), Positives = 56/148 (37%), Gaps = 6/148 (4%)
Frame = +3
Query: 49 TESQSQSPTGIHQSLPSSS------SSSSSSLTTVDQPPPSDDSPKAAVVSSSPKAVVVS 102
T S SP+ S PS S S S S + V PP S P + +SSP VV +
Sbjct: 495 TPKSSPSPSAGGLSPPSPSPTTTTPSPSGSPPSPVAIPPASSPVPTSGPTASSPSPVVST 674
Query: 103 SPLPAPMEKNSNTIASDGGDGGNAGGSKKPAWNKQPLNGVAVEIGPVMGAESWPALSESA 162
P PM + + + S GG S PA L+ GP A PA
Sbjct: 675 PPAGGPMASSPSPVVSTPPAGGPMASSPSPAGGPPALSPAG---GPSTAAGGPPAPGPGG 845
Query: 163 KIPGKLPPESSSSKIAPPAAVDGSPSTS 190
P + +S AA GSP ++
Sbjct: 846 AATSPGPGGAGASGPGGAAAAPGSPGSN 929
Score = 29.3 bits (64), Expect = 4.1
Identities = 32/127 (25%), Positives = 43/127 (33%), Gaps = 19/127 (14%)
Frame = +3
Query: 86 SPKAAVVSSSPKAVVVSSPLPAPMEKNSNTIASDGG------------DGGNAGGSKKPA 133
SP+ S SP A +S P P+P + S G S P
Sbjct: 486 SPRTPKSSPSPSAGGLSPPSPSPTTTTPSPSGSPPSPVAIPPASSPVPTSGPTASSPSPV 665
Query: 134 WNKQPLNG-VAVEIGPVMGAESWPALSESAKIPGKLPPE------SSSSKIAPPAAVDGS 186
+ P G +A PV+ S+ P PP S++ PPA G
Sbjct: 666 VSTPPAGGPMASSPSPVVSTPPAGGPMASSPSPAGGPPALSPAGGPSTAAGGPPAPGPGG 845
Query: 187 PSTSQGP 193
+TS GP
Sbjct: 846 AATSPGP 866
>TC83149 similar to GP|13561982|gb|AAK30594.1 flagelliform silk protein
{Argiope trifasciata}, partial (3%)
Length = 1177
Score = 45.1 bits (105), Expect = 7e-05
Identities = 56/188 (29%), Positives = 68/188 (35%), Gaps = 6/188 (3%)
Frame = +3
Query: 186 SPSTSQGPIISHSPQRQGSTSNTKSSSMANNNLPNRPRPMRRISGSNIGPCPAQSSLSNP 245
+PSTS S +P R ST T+ SS P+ PR R + S P++ S
Sbjct: 12 TPSTSPS-YASRTPLR--STPPTRPSSPT----PSAPRTSSRSTPST---SPSRRPRSKT 161
Query: 246 PTPPPLPPYPVYQLPPAVSYPNMLPSIPDSSPRDHHRNNNWDARPFVGGSSRRGHFGSHP 305
P PPP PP P + PP S P M S P SP W R
Sbjct: 162 PLPPPPPPLPQHPKPPQKSPPRMRRSKP--SPSSPAATPLWPQR---------------- 287
Query: 306 RGDGSYHQNSYSSRREHDRGNYANTRDAHAPQPRMPPRGIL------RPPPPSTAAFLGP 359
+ S S+ R + AP P PPR L RPP PST P
Sbjct: 288 ----TTPPPSPSTPRPSP*TPATPSTSPTAPLPTPPPRTTLPPALTPRPPSPSTPPTPKP 455
Query: 360 QTIGPFPA 367
+ PA
Sbjct: 456 GPVSVSPA 479
Score = 35.0 bits (79), Expect = 0.074
Identities = 52/191 (27%), Positives = 68/191 (35%), Gaps = 2/191 (1%)
Frame = +3
Query: 64 PSSSSSSSSSLTTVDQPPPSDDSPKAAVVSSSPKAVVVSSPLPAPMEKNSNTIASDGGDG 123
PS+S S +S PP SP S+P+ S+P +P
Sbjct: 15 PSTSPSYASRTPLRSTPPTRPSSP----TPSAPRTSSRSTPSTSP--------------- 137
Query: 124 GNAGGSKKPAWNKQPLNGVAVEIGPVMGAESWPALSESAKIPGKLPPESSSSKIAP--PA 181
S++P +K PL P L + K P K PP SK +P PA
Sbjct: 138 -----SRRPR-SKTPLPPPP------------PPLPQHPKPPQKSPPRMRRSKPSPSSPA 263
Query: 182 AVDGSPSTSQGPIISHSPQRQGSTSNTKSSSMANNNLPNRPRPMRRISGSNIGPCPAQSS 241
A P + P +P+ T T S+S P P R + P P S
Sbjct: 264 ATPLWPQRTTPPPSPSTPRPSP*TPATPSTSPT---APLPTPPPRTTLPPALTPRPP--S 428
Query: 242 LSNPPTPPPLP 252
S PPTP P P
Sbjct: 429 PSTPPTPKPGP 461
Score = 33.1 bits (74), Expect = 0.28
Identities = 45/196 (22%), Positives = 81/196 (40%), Gaps = 1/196 (0%)
Frame = +3
Query: 4 TQNNHSPTSVPSPAPPSAETNNPNFIRKNLPSPWAQVVRGGSGWDTESQSQSPTGIHQSL 63
T + +P++ PS P S P P P Q + +SP + +S
Sbjct: 105 TSSRSTPSTSPSRRPRSKTPLPPP------PPPLPQ--------HPKPPQKSPPRMRRSK 242
Query: 64 PSSSSSSSSSLTTV-DQPPPSDDSPKAAVVSSSPKAVVVSSPLPAPMEKNSNTIASDGGD 122
PS SS +++ L PPPS +P+ + + + + ++PLP P + + A
Sbjct: 243 PSPSSPAATPLWPQRTTPPPSPSTPRPSP*TPATPSTSPTAPLPTPPPRTTLPPAL---- 410
Query: 123 GGNAGGSKKPAWNKQPLNGVAVEIGPVMGAESWPALSESAKIPGKLPPESSSSKIAPPAA 182
+ +P P + GPV + + P S + ++P + +SS+K+ P AA
Sbjct: 411 ------TPRP---PSPSTPPTPKPGPVSVSPASP--SATQRVPWRPTARASSTKV-PAAA 554
Query: 183 VDGSPSTSQGPIISHS 198
+T + +S S
Sbjct: 555 KP*RRATRRPSAVSRS 602
>TC77148 ENOD20
Length = 1108
Score = 43.9 bits (102), Expect = 2e-04
Identities = 43/131 (32%), Positives = 58/131 (43%), Gaps = 12/131 (9%)
Frame = +3
Query: 9 SPTSVPSPAPPSAETNNPNFIRKNLPSPWAQVVRGGSGWDTESQSQSPTGIHQSLP---- 64
SP PSP P + T P+ R++LPSP S + S S SP+ +S P
Sbjct: 423 SPPPPPSPPTPRSSTPIPHPPRRSLPSP-------PSPSPSPSPSPSPSPSPRSTPIPHP 581
Query: 65 --SSSSSSSSSLTTVDQPPPSDD-----SPKAAVVSSSPKAVVV-SSPLPAPMEKNSNTI 116
S +S S S + P PS+ SP +V S +P + SP PAP +
Sbjct: 582 RKRSPASPSPSPSLSKSPSPSESPSLAPSPSDSVASLAPSSSPSDESPSPAP------SP 743
Query: 117 ASDGGDGGNAG 127
+S G GG AG
Sbjct: 744 SSSGSKGGGAG 776
Score = 29.6 bits (65), Expect = 3.1
Identities = 17/36 (47%), Positives = 18/36 (49%)
Frame = +3
Query: 242 LSNPPTPPPLPPYPVYQLPPAVSYPNMLPSIPDSSP 277
LS+PP PPP PP P P LPS P SP
Sbjct: 417 LSSPP-PPPSPPTPRSSTPIPHPPRRSLPSPPSPSP 521
Score = 29.3 bits (64), Expect = 4.1
Identities = 28/120 (23%), Positives = 48/120 (39%), Gaps = 13/120 (10%)
Frame = +3
Query: 142 VAVEIGPVMGAESWPALSESAKIPGKLPPESSSSKIAPPAAVDGSPSTSQGPIISHSPQ- 200
V V + PV+ + P + + +P S +PP+ SPS S P S SP+
Sbjct: 393 VVVMVAPVLSSPPPPPSPPTPRSSTPIPHPPRRSLPSPPSP---SPSPSPSPSPSPSPRS 563
Query: 201 -------RQGSTSNTKSSSMANNNLPNR-----PRPMRRISGSNIGPCPAQSSLSNPPTP 248
++ S + S S++ + P+ P P ++ P+ S S P+P
Sbjct: 564 TPIPHPRKRSPASPSPSPSLSKSPSPSESPSLAPSPSDSVASLAPSSSPSDESPSPAPSP 743
Score = 28.1 bits (61), Expect = 9.0
Identities = 21/64 (32%), Positives = 27/64 (41%), Gaps = 5/64 (7%)
Frame = +3
Query: 219 PNRPRPMRRISGSNIGPCPAQSSLSNPPTPPPLP-----PYPVYQLPPAVSYPNMLPSIP 273
P P P S + I P P + SL +PP+P P P P P + P P+ P
Sbjct: 429 PPPPSPPTPRSSTPI-PHPPRRSLPSPPSPSPSPSPSPSPSPSPRSTPIPHPRKRSPASP 605
Query: 274 DSSP 277
SP
Sbjct: 606 SPSP 617
>TC85251 homologue to PIR|JQ2128|JQ2128 metallothionein - soybean, partial
(93%)
Length = 1554
Score = 43.9 bits (102), Expect = 2e-04
Identities = 34/113 (30%), Positives = 47/113 (41%), Gaps = 4/113 (3%)
Frame = -2
Query: 162 AKIPGKLPPESSSS----KIAPPAAVDGSPSTSQGPIISHSPQRQGSTSNTKSSSMANNN 217
A P KLPP + ++ PP V P TSQ P+ + +P+ TS + A++
Sbjct: 845 ATAPTKLPPTTPTAITPVTTQPPTVVASPPITSQPPV-TVAPKSAPVTSPAPKIAPASSP 669
Query: 218 LPNRPRPMRRISGSNIGPCPAQSSLSNPPTPPPLPPYPVYQLPPAVSYPNMLP 270
P+P P S +S P PPPLPP P P + P P
Sbjct: 668 KVPPPQP------------PKSSPVSTPTLPPPLPPPPKISPTPVQTPPAPAP 546
Score = 37.4 bits (85), Expect = 0.015
Identities = 41/152 (26%), Positives = 55/152 (35%), Gaps = 1/152 (0%)
Frame = -2
Query: 132 PAWNKQPLNGVAVEIGPVMGAESWPALSESAKIPGKLPPESSSSKIAPPAAVDGSPSTSQ 191
P QP VA + PV A + S K+P PP+SS P + P
Sbjct: 761 PPITSQPPVTVAPKSAPVTSPAPKIAPASSPKVPPPQPPKSS-----PVSTPTLPPPLPP 597
Query: 192 GPIISHSPQRQGSTSNTKSSSMANNNLPNRPRPMRRISGSNIGPCPAQSSLSNPPTPPPL 251
P IS +P + P P P++ P PA + PTP P
Sbjct: 596 PPKISPTPVQ----------------TPPAPAPVKAT------PVPAPAPAKQAPTPAPA 483
Query: 252 PPYPVYQLPPAVSYPNMLPSIPDSSP-RDHHR 282
P+ PA+ P +P+ S P R HR
Sbjct: 482 TSPPIPAPTPAIEAP--VPAPESSKPKRRRHR 393
Score = 37.0 bits (84), Expect = 0.019
Identities = 56/250 (22%), Positives = 95/250 (37%), Gaps = 10/250 (4%)
Frame = -2
Query: 46 GWDTESQSQSPTGIHQSLPSSSSSSSSSLTTVDQPPPSDDSP-------KAAVVSSSPKA 98
G++ ++ + +PT + + P++ + ++ TV PP P A V S +PK
Sbjct: 866 GFNAQAPATAPTKLPPTTPTAITPVTTQPPTVVASPPITSQPPVTVAPKSAPVTSPAPKI 687
Query: 99 VVVSSP-LPAPMEKNSNTIASDGGDGGNAGGSKKPAWNKQP-LNGVAVEIGPVMGAESWP 156
SSP +P P S+ +++ + P P ++ V+ P P
Sbjct: 686 APASSPKVPPPQPPKSSPVSTP---------TLPPPLPPPPKISPTPVQTPPA------P 552
Query: 157 ALSESAKIPGKLPPESSSSKIAPPAAVDGSPSTSQGPIISHSPQRQGSTSNTKSSSMANN 216
A ++ +P P +K AP A SP PI + +P + +SS
Sbjct: 551 APVKATPVPAPAP-----AKQAPTPAPATSP-----PIPAPTPAIEAPVPAPESSKPKR- 405
Query: 217 NLPNRPRPMRRISGSNIGPCPAQSSLSNPPTPPPLPPYPVYQLPPAVSYPNMLPSIP-DS 275
R RP R + P PA + + P PP PA + PS +
Sbjct: 404 ---RRHRPKHR---RHQAPAPAPTVIHKSPPAPPTDTTADSDTAPAPA-----PSFNLNG 258
Query: 276 SPRDHHRNNN 285
+P +H + N
Sbjct: 257 APSNHFQGGN 228
Score = 28.5 bits (62), Expect = 6.9
Identities = 17/75 (22%), Positives = 28/75 (36%)
Frame = -2
Query: 9 SPTSVPSPAPPSAETNNPNFIRKNLPSPWAQVVRGGSGWDTESQSQSPTGIHQSLPSSSS 68
+P P + P + P R P+P +PT IH+S P+ +
Sbjct: 443 APVPAPESSKPKRRRHRPKHRRHQAPAP------------------APTVIHKSPPAPPT 318
Query: 69 SSSSSLTTVDQPPPS 83
+++ T P PS
Sbjct: 317 DTTADSDTAPAPAPS 273
>TC80046 similar to SP|P22699|GUN6_DICDI Endoglucanase precursor (EC
3.2.1.4) (Endo-1 4-beta-glucanase) (Spore germination
protein 270-6), partial (11%)
Length = 643
Score = 43.5 bits (101), Expect = 2e-04
Identities = 23/103 (22%), Positives = 46/103 (44%)
Frame = +3
Query: 2 VTTQNNHSPTSVPSPAPPSAETNNPNFIRKNLPSPWAQVVRGGSGWDTESQSQSPTGIHQ 61
+T + + T PSP S ET+ + +P + TE+ +++PT
Sbjct: 231 ITVTDTSTNTVSPSPTETSTETSTETPTETSTETPTETSTETSTETPTETSTETPTETST 410
Query: 62 SLPSSSSSSSSSLTTVDQPPPSDDSPKAAVVSSSPKAVVVSSP 104
P+ +S+ +S+ T+ + PPPS ++ S+ + + P
Sbjct: 411 ETPTETSTETSTETSTETPPPSTETSTETTPPSTETSTETTPP 539
>TC83962 similar to PIR|B86289|B86289 hypothetical protein AAF71991.1
[imported] - Arabidopsis thaliana, partial (11%)
Length = 679
Score = 43.1 bits (100), Expect = 3e-04
Identities = 43/144 (29%), Positives = 64/144 (43%), Gaps = 2/144 (1%)
Frame = +2
Query: 53 SQSPTGI-HQSLPSSSSSSSSSLTTVDQPPPSDDSPKAAVVSSSPKAVVVSSPLPAPMEK 111
++SP I + S PS SSS+S+S ++ PPP+ + ++ S P +SSP P
Sbjct: 50 NKSPA*ISNSSTPSPSSSTSTSTSSSSSPPPTPST--SSTTPSPPPPTSLSSPTP----- 208
Query: 112 NSNTIASDGGDGGNAGGSKKPAWNKQPLNGV-AVEIGPVMGAESWPALSESAKIPGKLPP 170
SN ++S A P N + AV P+ S L + +P LP
Sbjct: 209 TSNHLSS--------------A*PTTPTNTLTAVPSTPLNSPCSNLHLPQQTSLPSPLPS 346
Query: 171 ESSSSKIAPPAAVDGSPSTSQGPI 194
S S + PPA S S+S P+
Sbjct: 347 SSPSYQTTPPALASVSLSSSATPL 418
Score = 35.4 bits (80), Expect = 0.057
Identities = 30/111 (27%), Positives = 46/111 (41%), Gaps = 10/111 (9%)
Frame = +2
Query: 159 SESAKIPGKLPPESSSSKIAPPAAVDGSPSTSQGPIISHSPQRQGST-----SNTKSSSM 213
+ S+ P P SS++ PP SP+ + + S P +T S +S
Sbjct: 116 TSSSSSPPPTPSTSSTTPSPPPPTSLSSPTPTSNHLSSA*PTTPTNTLTAVPSTPLNSPC 295
Query: 214 ANNNLPNR-----PRPMRRISGSNIGPCPAQSSLSNPPTPPPLPPYPVYQL 259
+N +LP + P P S P A SLS+ TP PLP + + +
Sbjct: 296 SNLHLPQQTSLPSPLPSSSPSYQTTPPALASVSLSSSATPLPLPAHSLVNI 448
Score = 32.7 bits (73), Expect = 0.37
Identities = 32/109 (29%), Positives = 45/109 (40%)
Frame = +2
Query: 169 PPESSSSKIAPPAAVDGSPSTSQGPIISHSPQRQGSTSNTKSSSMANNNLPNRPRPMRRI 228
P S + I+ + S STS S SP STS+T S +L + +
Sbjct: 44 PANKSPA*ISNSSTPSPSSSTSTSTSSSSSPPPTPSTSSTTPSPPPPTSLSSPTPTSNHL 223
Query: 229 SGSNIGPCPAQSSLSNPPTPPPLPPYPVYQLPPAVSYPNMLPSIPDSSP 277
S + P ++L+ P+ P P LP S P+ LPS SSP
Sbjct: 224 SSA--*PTTPTNTLTAVPSTPLNSPCSNLHLPQQTSLPSPLPS---SSP 355
>TC87918 weakly similar to GP|15810397|gb|AAL07086.1 unknown protein
{Arabidopsis thaliana}, partial (67%)
Length = 1746
Score = 42.7 bits (99), Expect = 4e-04
Identities = 24/70 (34%), Positives = 42/70 (59%), Gaps = 7/70 (10%)
Frame = +2
Query: 391 FPHMPSPATFFPA-AESSL----SNVIVNQIDYYFSDINLANDEFLKS--NMDEEGWVPI 443
F +P ++FF + A+ SL + ++ Q+++YFSD NL D+FL N E+G V +
Sbjct: 17 FSPIPPLSSFFSSMAKHSLDEETTKKVIRQVEFYFSDSNLPRDDFLSKTVNESEDGMVSL 196
Query: 444 TLIANFPRVK 453
L+ +F R++
Sbjct: 197 ALLCSFNRMR 226
>TC88762 similar to GP|22136928|gb|AAM91808.1 unknown protein {Arabidopsis
thaliana}, partial (3%)
Length = 805
Score = 42.4 bits (98), Expect = 5e-04
Identities = 43/176 (24%), Positives = 61/176 (34%), Gaps = 9/176 (5%)
Frame = +3
Query: 32 NLPSPWAQVVRGGSGWDTESQSQSPTGIHQSLPSSSSSSSSSLTTVDQPPPSDD------ 85
+LP AQ G + Q +PT PSSS S+SS ++T PP S
Sbjct: 12 SLPPGVAQTPSGNGIAPSPYQFNNPTS-----PSSSPSASSPVSTTTSPPASTPASSPVS 176
Query: 86 ---SPKAAVVSSSPKAVVVSSPLPAPMEKNSNTIASDGGDGGNAGGSKKPAWNKQPLNGV 142
SP A+ +SSP S P P P +T + G+
Sbjct: 177 TTTSPPASTPASSPVPTTTSPPAPTPASSPVSTNSPTASPAGSL---------------- 308
Query: 143 AVEIGPVMGAESWPALSESAKIPGKLPPESSSSKIAPPAAVDGSPSTSQGPIISHS 198
P S + + + P PP + SS P G S S + ++S
Sbjct: 309 -----PAAATPSPSSTTAGSPSPSSTPPGTRSSNETTPGRSIGVASASPSGVWAYS 461
Score = 29.3 bits (64), Expect = 4.1
Identities = 32/126 (25%), Positives = 49/126 (38%), Gaps = 1/126 (0%)
Frame = +3
Query: 172 SSSSKIAPPAAVDGSPSTSQGPIISHSPQRQGSTSNTKSSSMANNNLPNRPRPMRRISGS 231
SSS + P + SP P + + +T++ +S+ A++ +P P
Sbjct: 96 SSSPSASSPVSTTTSP-----PASTPASSPVSTTTSPPASTPASSPVPTTTSPP------ 242
Query: 232 NIGPCPAQSSLS-NPPTPPPLPPYPVYQLPPAVSYPNMLPSIPDSSPRDHHRNNNWDARP 290
P PA S +S N PT P P P S PS P S+P +N
Sbjct: 243 --APTPASSPVSTNSPTASPAGSLPAAATPSPSSTTAGSPS-PSSTPPGTRSSNETTPGR 413
Query: 291 FVGGSS 296
+G +S
Sbjct: 414 SIGVAS 431
>BQ144502 weakly similar to GP|6523547|emb hydroxyproline-rich glycoprotein
DZ-HRGP {Volvox carteri f. nagariensis}, partial (39%)
Length = 1358
Score = 42.4 bits (98), Expect = 5e-04
Identities = 86/349 (24%), Positives = 118/349 (33%), Gaps = 40/349 (11%)
Frame = -2
Query: 64 PSSSSSSSSSLTTVDQPPPSDDSPKAAV----------------VSSSPKAVV------- 100
PS + S + +T + P S SP+AA + P+A +
Sbjct: 1324 PSRTRSPAVGITALSSPFVSSRSPRAARDTLRRRGVGARARRARETHLPRARIRDRPYSR 1145
Query: 101 VSSPLPAPMEKNS-NTIASDGGDGGNAGGSKKPAWNKQPLNGV-AVEIGPVMGAESWPAL 158
P PAP + T S A ++PA G+ AV P A S PA
Sbjct: 1144 APQPAPAPPAHAALRTRRSAPASRRRAPAPRRPA--PSIYGGLCAVRAAPSERAFSAPAP 971
Query: 159 S------ESAKIPGKLPPESSSSKIAPPAAVDGSPSTSQGPIISHSPQRQGSTSNTKSSS 212
S + +LPP S A +D +P+ P P ST ++ +
Sbjct: 970 SGVCPVVSRSSTGARLPPRYSLVPRAHAREIDRAPTFEHFP----GPPDPASTPLDRARA 803
Query: 213 MANNNLPNRPRPMRRISGSNIGPCPAQ----SSLSNPPTPPPLPPYPVYQLPPAVSYPNM 268
P+ RP PA+ +SL+ P PPP P P LPP ++
Sbjct: 802 CCARRPPSPHRPAPPPQAH-----PARERRTASLARTPPPPPSSPPPAPLLPPRNGNASL 638
Query: 269 LPSIPDSSPRDHHRNNNWDARPFVGGSSRRGHFGSHPRGDGSYHQNSYSSRREHDRGNYA 328
LP R VG +RR +S SS H + A
Sbjct: 637 LP------------------RKLVGSPARR---------------SSLSSESLHTPAHLA 557
Query: 329 NTRDAHAPQPRMPPRGILRP-----PPPSTAAFLGPQTIGPFPAPVAYP 372
R + P+P P RP PPP+ A P GP PAP P
Sbjct: 556 PPRQSPRPRPPPEPLHNPRPRPPPRPPPAAPARAPPS--GP-PAPPTPP 419
Score = 40.0 bits (92), Expect = 0.002
Identities = 55/231 (23%), Positives = 76/231 (32%), Gaps = 23/231 (9%)
Frame = -2
Query: 71 SSSLTTVDQPPPSDDSPKAAVVSSSPKAVVVSSPLP-APMEKNSNTIASDGGDGGNAGGS 129
++SL PPPS P + + A ++ L +P ++S + S A
Sbjct: 727 TASLARTPPPPPSSPPPAPLLPPRNGNASLLPRKLVGSPARRSSLSSESLHTPAHLAPPR 548
Query: 130 KKPAWNKQPLNGVAVEIGPVMGAESWPALSESAKIPGKLPPESSSSKIAPPAA----VDG 185
+ P P P+ P P + PP + PPA V
Sbjct: 547 QSPRPRPPP--------EPLHNPRPRPPPRPPPAAPARAPPSGPPAPPTPPATRAPPVRA 392
Query: 186 SP-STSQGPIISHSPQRQGSTSNTKSSSMANNNLPNRPRPMRRISGSNIGPCPAQSSL-- 242
SP QG +PQ + + + A + P P P+ R P P S L
Sbjct: 391 SPWCPGQG---QETPQGRHTGVQSPVPCPALPHPPGPPPPLDRAPPPPPAPLPLASRLRP 221
Query: 243 ---------------SNPPTPPPLPPYPVYQLPPAVSYPNMLPSIPDSSPR 278
S PP PPP PP P + PP P P S PR
Sbjct: 220 RASPPPPPPPPGPAASAPPRPPPPPPPPPQRPPPPPPPHTHQPEPPTSPPR 68
Score = 38.9 bits (89), Expect = 0.005
Identities = 59/234 (25%), Positives = 76/234 (32%), Gaps = 3/234 (1%)
Frame = -2
Query: 168 LPPESSSSKIAPPAAVDGSPS--TSQGPIISHSPQRQGSTSNTKSSSMANNNLPN-RPRP 224
LPP + ++ + P V GSP+ +S H+P + L N RPRP
Sbjct: 667 LPPRNGNASLLPRKLV-GSPARRSSLSSESLHTPAHLAPPRQSPRPRPPPEPLHNPRPRP 491
Query: 225 MRRISGSNIGPCPAQSSLSNPPTPPPLPPYPVYQLPPAVSYPNMLPSIPDSSPRDHHRNN 284
R P PA + + PP+ PP PP P P V P +P+ H
Sbjct: 490 PPR-------PPPAAPARA-PPSGPPAPPTPPATRAPPVRASPWCPGQGQETPQGRHTGV 335
Query: 285 NWDARPFVGGSSRRGHFGSHPRGDGSYHQNSYSSRREHDRGNYANTRDAHAPQPRMPPRG 344
P G R + R R + P PPR
Sbjct: 334 Q-SPVPCPALPHPPGPPPPLDRAPPPPPAPLPLASRLRPRASPPPPPPPPGPAASAPPRP 158
Query: 345 ILRPPPPSTAAFLGPQTIGPFPAPVAYPDFYYFPTVPLDPFRGMPVFPHMPSPA 398
PPPP PQ P P P + P P P R P P P P+
Sbjct: 157 PPPPPPP-------PQRPPPPPPPHTHQ-----PEPPTSPPRRQPPAPRAPPPS 32
Score = 38.1 bits (87), Expect = 0.009
Identities = 67/247 (27%), Positives = 88/247 (35%), Gaps = 50/247 (20%)
Frame = -1
Query: 81 PPS---DDSPKAAVVSSSPKAVVVSSPL-------PAPMEKNSNTIASDGGDGGNAGGSK 130
PPS +P A S P+A ++PL P P E +A GG
Sbjct: 773 PPSATPPGAPCARAPHSIPRAHPAATPLLPPPRPPPPPPEWERVPLAK------KIGGQP 612
Query: 131 KPAWNKQPLNGVAV---------EIGPVMGAESWPALSESAKIPGKLPPESSSSK--IAP 179
+PA PL VA + P P SE+A P PP + + + P
Sbjct: 611 RPAL--VPLVRVATYPRPPRPPPPVPPPPSPPRAPPQSEAAPPPPP-PPRGARPRAPLRP 441
Query: 180 PAAVD--GSP-STSQGPIISHSPQRQGSTSNTKSSSMANNNLPNRPRPMRRISGSNIGPC 236
P+A D G P + QG + P R G++ LP P P R + GP
Sbjct: 440 PSAPDPAGHPRAPRQGVAVVPGP-RSGNSPRAAHRGPEPRPLPRPPPPTRPPAAPGPGPP 264
Query: 237 PAQ--SSLSNPPT---------PPP------------LPPYPVYQLPPAVSYPNM---LP 270
PA + LS PP PPP PP P PPA + P+ P
Sbjct: 263 PAPGPTPLSKPPAAEGIPPATAPPPRARRLRPPSAPAPPPTPPAAAPPAPTPPHAPA*AP 84
Query: 271 SIPDSSP 277
+P P
Sbjct: 83 HLPPPPP 63
Score = 36.6 bits (83), Expect = 0.025
Identities = 49/188 (26%), Positives = 63/188 (33%), Gaps = 2/188 (1%)
Frame = -1
Query: 81 PPSDDSPKAAVVSSSPKAVVVSSPLPAPMEKNSNTIASDGGDGGNAGGSKKPAWNKQPLN 140
PP P + + P++ P P P AG + P
Sbjct: 557 PPPPVPPPPSPPRAPPQSEAAPPPPPPPRGARPRAPLRPPSAPDPAGHPRAPR------Q 396
Query: 141 GVAVEIGPVMGAESWPALSESAKIPGKLPPESSSSKIAPPAAVDGSPSTSQGPI-ISHSP 199
GVAV GP G P + P LP ++ PPAA P + GP +S P
Sbjct: 395 GVAVVPGPRSGNS--PRAAHRGPEPRPLPRPPPPTR--PPAAPGPGPPPAPGPTPLSKPP 228
Query: 200 QRQGSTSNTKSSSMANNNLPNRPRPMRRISGSNIGPCPAQSSLSNPPTP-PPLPPYPVYQ 258
+G T P R R +R S P P ++ PP P PP P
Sbjct: 227 AAEGIPPATAP--------PPRARRLRPPSAPAPPPTPPAAA---PPAPTPPHAPA*APH 81
Query: 259 LPPAVSYP 266
LPP P
Sbjct: 80 LPPPPPAP 57
Score = 35.4 bits (80), Expect = 0.057
Identities = 66/259 (25%), Positives = 86/259 (32%), Gaps = 9/259 (3%)
Frame = -3
Query: 153 ESWPALSESAKIPGK----LPPESSSSKIAPPAAVDG-SPSTSQGPIISHSPQRQGSTSN 207
E+W A A+ P + +PP +S +PPA V SPST +G +P + +
Sbjct: 630 ENWWAAPPGAR-PSRPSRYIPPPTSPPPASPPAPVPPQSPSTIRGRAPPPAPPPRRPPAR 454
Query: 208 TKSSSMANNNLPNRPRPMRRI----SGSNIGPCPAQSSLSNPPTPPPLPPYPVYQLPPAV 263
+ P RPRP R SG G L TP P P PP+
Sbjct: 453 PPPA-------PQRPRPRRPPARPPSGRRRGARAKVRKLPKGGTPGSRAPSPA---PPSP 304
Query: 264 SYPNMLPSIPDSSPRDHHRNNNWDARPFVGGSSRRGHFGSHPRGDGSYHQNSYSSRREHD 323
++P +PR RP+ G R R H
Sbjct: 303 THP---------APR----------RPWTGPPPRP---------------------RPHS 244
Query: 324 RGNYANTRDAHAPQPRMPPRGILRPPPPSTAAFLGPQTIGPFPAPVAYPDFYYFPTVPLD 383
A R H P+ R PP G PPP P +GP P P
Sbjct: 243 P*QAACGR-GHPPRHRPPPPG---PPP--------PPPLGPRPPP--------------H 142
Query: 384 PFRGMPVFPHMPSPATFFP 402
P R P PH P+ + P
Sbjct: 141 PPRSGPPRPHPPTRTSLSP 85
Score = 35.4 bits (80), Expect = 0.057
Identities = 59/266 (22%), Positives = 79/266 (29%)
Frame = -2
Query: 8 HSPTSVPSPAPPSAETNNPNFIRKNLPSPWAQVVRGGSGWDTESQSQSPTGIHQSLPSSS 67
H P P A P+ E + R P P + + S P + S S
Sbjct: 775 HRPAPPPQ-AHPARERRTASLARTPPPPPSSPPPAPLLPPRNGNASLLPRKLVGSPARRS 599
Query: 68 SSSSSSLTTVDQPPPSDDSPKAAVVSSSPKAVVVSSPLPAPMEKNSNTIASDGGDGGNAG 127
S SS SL T P SP+ P + +P P P + + G
Sbjct: 598 SLSSESLHTPAHLAPPRQSPRPR-----PPPEPLHNPRPRPPPRPPPAAPARAPPSGPPA 434
Query: 128 GSKKPAWNKQPLNGVAVEIGPVMGAESWPALSESAKIPGKLPPESSSSKIAPPAAVDGSP 187
PA P+ A P G E+ + P P PP P
Sbjct: 433 PPTPPATRAPPVR--ASPWCPGQGQETPQGRHTGVQSPVPCPALPHPPGPPPPLDRAPPP 260
Query: 188 STSQGPIISHSPQRQGSTSNTKSSSMANNNLPNRPRPMRRISGSNIGPCPAQSSLSNPPT 247
+ P+ S R A + P P P P P + PPT
Sbjct: 259 PPAPLPLASRLRPRASPPPPPPPPGPAASAPPRPPPPPPPPPQRPPPPPPPHTHQPEPPT 80
Query: 248 PPPLPPYPVYQLPPAVSYPNMLPSIP 273
PP P + PP + PS+P
Sbjct: 79 SPPRRQPPAPRAPPPSGI*LIAPSVP 2
Score = 30.4 bits (67), Expect = 1.8
Identities = 64/300 (21%), Positives = 98/300 (32%), Gaps = 26/300 (8%)
Frame = -3
Query: 8 HSPTSVPS-PAP-PSAETNNPNFIRKNLPSPWAQVVRGGSGWDTESQSQSPTGIHQSLPS 65
H P S PS PAP P+ + + + L + R S T Q P + +
Sbjct: 975 HRPASAPSSPAPLPARDCRRGTLLFRALTRARSTARRRSS---TSPAPQIPPLRRSIVRA 805
Query: 66 SSSSSSSSLTTVDQPPPSDDSPKAAVVSSSPKAVVVSSPLPAPMEKNSNTIASDGGDGGN 125
++ + T + PP +A S P P P + + + N
Sbjct: 804 RAARAVRPPRTAQRHPPRRTLRASAAQHPSRAPRRHPPPPPPPPPSSPPGMGTRPSCQEN 625
Query: 126 AGGSKKPAWNKQPLNGVAVEIGPVMGAESWPALSES-AKIPGKLPPESSSSKIAP--PAA 182
+ A +P + P + P +S + I G+ PP + + P P
Sbjct: 624 WWAAPPGARPSRPSRYIPPPTSPPPASPPAPVPPQSPSTIRGRAPPPAPPPRRPPARPPP 445
Query: 183 VDGSPSTSQGPIISHSPQRQGS-----------TSNTKSSSMANNNLPNRPRPMRRISGS 231
P + P S +R+G+ T +++ S A + P P P R +G
Sbjct: 444 APQRPRPRRPPARPPSGRRRGARAKVRKLPKGGTPGSRAPSPAPPS-PTHPAPRRPWTGP 268
Query: 232 NIGPCP----------AQSSLSNPPTPPPLPPYPVYQLPPAVSYPNMLPSIPDSSPRDHH 281
P P PP P P PP P+ PP P P S P H
Sbjct: 267 PPRPRPHSP*QAACGRGHPPRHRPPPPGPPPPPPLGPRPP--------PHPPRSGPPRPH 112
>AW585122
Length = 507
Score = 40.8 bits (94), Expect = 0.001
Identities = 29/96 (30%), Positives = 49/96 (50%), Gaps = 3/96 (3%)
Frame = +3
Query: 24 NNPN-FIRKNLPSPWAQVVRGGSGWDTESQSQSPTGIHQSLPSSSSSSSSSLTTVDQPP- 81
+ PN ++R +PWA ++R G+ ++ +SP S+P +S S S + D PP
Sbjct: 138 SRPN*YMRPMSSAPWADLMRIGA---RSTKPKSPA----SMPPASLSPSMRVANADPPPP 296
Query: 82 -PSDDSPKAAVVSSSPKAVVVSSPLPAPMEKNSNTI 116
PS D+P + ++ P PAP +K S+T+
Sbjct: 297 TPSSDAPPSTTRTTPPDLA------PAPRKKTSSTL 386
>TC79359 weakly similar to PIR|S49915|S49915 extensin-like protein - maize,
partial (3%)
Length = 1170
Score = 40.4 bits (93), Expect = 0.002
Identities = 52/182 (28%), Positives = 74/182 (40%), Gaps = 1/182 (0%)
Frame = +3
Query: 61 QSLPSSSSSSSSSLTTVDQPPPSDDSPKAAVVSSSP-KAVVVSSPLPAPMEKNSNTIASD 119
Q+ P+SS SS V PPS S +SP K+ +S P P S + ++
Sbjct: 219 QNAPASSPKSS-----VTAKPPSSVSVSPTNSPASPAKSPTLSPPSQTPAVSPSGSASTP 383
Query: 120 GGDGGNAGGSKKPAWNKQPLNGVAVEIGPVMGAESWPALSESAKIPGKLPPESSSSKIAP 179
+ +K PA QP + V+ PA+S S + PP SS + P
Sbjct: 384 --PPATSPPAKSPA--VQPPSSVS------------PAISPSNNV-SSTPPVSSPASSPP 512
Query: 180 PAAVDGSPSTSQGPIISHSPQRQGSTSNTKSSSMANNNLPNRPRPMRRISGSNIGPCPAQ 239
AAV S + P +S P + S++ SSS + P P S + G PA
Sbjct: 513 TAAVSPVSSPVEAPSVSSPP--EASSAGIPSSSATPADAPAATLP----SSKSPGTSPAS 674
Query: 240 SS 241
SS
Sbjct: 675 SS 680
Score = 37.7 bits (86), Expect = 0.011
Identities = 48/186 (25%), Positives = 71/186 (37%), Gaps = 1/186 (0%)
Frame = +3
Query: 9 SPTSVPSPAPPSAETNNPNFIRKNLPSPWAQVVRGGSGWDTESQSQSPTGIHQSLPSSSS 68
SP S + PPS+ + +P N P+ S ++SPT S + S
Sbjct: 237 SPKSSVTAKPPSSVSVSPT----NSPA---------------SPAKSPTLSPPSQTPAVS 359
Query: 69 SSSSSLTTVDQPPPSDDSPKAAVVSSSPKAVVVSSPLPAPMEKNSNTIASDGGDGGNAGG 128
S S+ T PPP+ P + P +V SP +P S+T
Sbjct: 360 PSGSAST----PPPATSPPAKSPAVQPPSSV---SPAISPSNNVSSTPPVSSPASSPPTA 518
Query: 129 SKKPAWNKQPLNGVAVEIGPVMGAESWPALSES-AKIPGKLPPESSSSKIAPPAAVDGSP 187
+ P P+ +V P + P+ S + A P P S S +P ++ SP
Sbjct: 519 AVSPV--SSPVEAPSVSSPPEASSAGIPSSSATPADAPAATLPSSKSPGTSPASS---SP 683
Query: 188 STSQGP 193
TSQGP
Sbjct: 684 ETSQGP 701
>TC76311 homologue to GP|3204132|emb|CAA07235.1 extensin {Cicer arietinum},
partial (77%)
Length = 973
Score = 40.4 bits (93), Expect = 0.002
Identities = 40/179 (22%), Positives = 66/179 (36%), Gaps = 7/179 (3%)
Frame = +2
Query: 245 PPTPPPLPPY-------PVYQLPPAVSYPNMLPSIPDSSPRDHHRNNNWDARPFVGGSSR 297
PP+P P PPY P PP Y + P P P ++++ P S
Sbjct: 2 PPSPSPPPPYYYHSPPPPSPSPPPPYYYKSPPPPSPSPPPPYYYKSP-----PPPSPSPP 166
Query: 298 RGHFGSHPRGDGSYHQNSYSSRREHDRGNYANTRDAHAPQPRMPPRGILRPPPPSTAAFL 357
++ Y S H +Y + A P PP + PPPP+++
Sbjct: 167 PPYY---------YQSPPPPSPTPHTPYHYKSPPPPTASPP--PPYHYVSPPPPTSSPPP 313
Query: 358 GPQTIGPFPAPVAYPDFYYFPTVPLDPFRGMPVFPHMPSPATFFPAAESSLSNVIVNQI 416
T P P+P P + Y P PV+ + P + A+++ + ++N +
Sbjct: 314 YHYTSPPPPSPAPAPTYIYKSPPPPMKSPPPPVYIYASPPPPIYK*AQNTTTAFLINHV 490
>BQ141932 weakly similar to GP|6523547|emb hydroxyproline-rich glycoprotein
DZ-HRGP {Volvox carteri f. nagariensis}, partial (30%)
Length = 1338
Score = 40.4 bits (93), Expect = 0.002
Identities = 26/63 (41%), Positives = 34/63 (53%), Gaps = 5/63 (7%)
Frame = -2
Query: 220 NRPRPMRRISGSNIGPCPAQ--SSLSNPPTPPPLPPYPVYQLPPAVSYPNMLPS---IPD 274
+RP P RRI GS+ P P + PP+PPP PP P++ P S P PS +P
Sbjct: 392 HRPSPPRRI-GSSTPPPPFVPIAGFPPPPSPPPPPPLPLHPPPQLSSLPPHAPSSFLLPL 216
Query: 275 SSP 277
+SP
Sbjct: 215 TSP 207
Score = 35.8 bits (81), Expect = 0.043
Identities = 31/103 (30%), Positives = 39/103 (37%), Gaps = 4/103 (3%)
Frame = -3
Query: 157 ALSESAKIPGKLPPESSSSKIAPPAAVDGS----PSTSQGPIISHSPQRQGSTSNTKSSS 212
AL SA P LPP S + P S P P+ P+R S
Sbjct: 454 ALPPSAPPPSDLPPPPRSPTVIDPRPPAASAVPLPLRPLYPLRDSPPRRPPPPPPHSPSI 275
Query: 213 MANNNLPNRPRPMRRISGSNIGPCPAQSSLSNPPTPPPLPPYP 255
+ ++ + P P+ S P PA PPTPPP PP P
Sbjct: 274 LPPSSRLSPPTPLPLSSS----P*PALPLPLPPPTPPPPPPAP 158
Score = 33.5 bits (75), Expect = 0.22
Identities = 44/181 (24%), Positives = 59/181 (32%), Gaps = 2/181 (1%)
Frame = -1
Query: 219 PNRPRPMRRISGSNIGPCPAQSSLSNPPTPPPLPPYPVYQLPPAVSYPNM-LPSIPDSS- 276
P RPRP R + P + S PP P P Q P ++ P P+ P S
Sbjct: 657 PARPRPGRVLLVMQY-PTSFLGTGSAPPAIHHHPNPPASQSPATLNRPTSHTPACPPPSR 481
Query: 277 PRDHHRNNNWDARPFVGGSSRRGHFGSHPRGDGSYHQNSYSSRREHDRGNYANTRDAHAP 336
P H R + P HP SS ++ A
Sbjct: 480 PPPHPRAPSLSPPP-----------PPHPPTSPPRPGRPRSSTLAPPPHRQFHSPSALCT 334
Query: 337 QPRMPPRGILRPPPPSTAAFLGPQTIGPFPAPVAYPDFYYFPTVPLDPFRGMPVFPHMPS 396
+PP + PPPP+ P + P P P +P P+ P P+ P P
Sbjct: 333 HCGIPPPAVPPPPPPTPPPSSPPALVSPPPRPFLFPPPPNQPSPSPSPPPPPPLPPPPPQ 154
Query: 397 P 397
P
Sbjct: 153 P 151
Score = 32.7 bits (73), Expect = 0.37
Identities = 30/103 (29%), Positives = 42/103 (40%), Gaps = 11/103 (10%)
Frame = -3
Query: 184 DGSPSTSQGP--IISHSPQRQGSTSNTKSSSMANNNLPNRPRPMRRIS-----GSNIGPC 236
+GSP T P IS S + S + + ++ NR RP R I
Sbjct: 709 EGSPFTPPPPRRTISGSSRPPASGAGSTGHAIPYFVFGNRQRPTRHPPPP*PPSIPIPRH 530
Query: 237 PAQSSLSNPPTPPPLPPYP----VYQLPPAVSYPNMLPSIPDS 275
P + +P PPP+P P + LPP+ P+ LP P S
Sbjct: 529 PEPTHQPHPCLPPPIPSAPPPTRAFALPPSAPPPSDLPPPPRS 401
Score = 30.0 bits (66), Expect = 2.4
Identities = 21/73 (28%), Positives = 27/73 (36%), Gaps = 17/73 (23%)
Frame = -3
Query: 222 PRPMRRISGSNIGPCPAQSSLSN-----------------PPTPPPLPPYPVYQLPPAVS 264
P P R ISGS+ P S + PP PP P P + P
Sbjct: 688 PPPRRTISGSSRPPASGAGSTGHAIPYFVFGNRQRPTRHPPPP*PPSIPIPRHPEPTHQP 509
Query: 265 YPNMLPSIPDSSP 277
+P + P IP + P
Sbjct: 508 HPCLPPPIPSAPP 470
Score = 29.6 bits (65), Expect = 3.1
Identities = 30/106 (28%), Positives = 42/106 (39%), Gaps = 3/106 (2%)
Frame = -3
Query: 168 LPPESSSSKIAPPAAVDGSPSTSQGPIISHSPQRQGSTSNTKSSSMANNNLPNRPR-PMR 226
LPP S+ PP P ++ P P R + + + + + LP RP P+R
Sbjct: 499 LPPPIPSAP--PPTRAFALPPSAPPPSDLPPPPRSPTVIDPRPPAASAVPLPLRPLYPLR 326
Query: 227 RISGSNIGPCPAQSSLSNPPTPPPLPPYPVYQLPPA--VSYPNMLP 270
S P PPP PP+ LPP+ +S P LP
Sbjct: 325 D---------------SPPRRPPPPPPHSPSILPPSSRLSPPTPLP 233
Score = 28.5 bits (62), Expect = 6.9
Identities = 12/27 (44%), Positives = 18/27 (66%)
Frame = -2
Query: 242 LSNPPTPPPLPPYPVYQLPPAVSYPNM 268
L++PP PPP PP+P P ++PN+
Sbjct: 218 LTSPPPPPP-PPHPPPSPPRPHNHPNL 141
>BQ144407 weakly similar to GP|6523547|emb| hydroxyproline-rich glycoprotein
DZ-HRGP {Volvox carteri f. nagariensis}, partial (23%)
Length = 1217
Score = 40.4 bits (93), Expect = 0.002
Identities = 52/201 (25%), Positives = 74/201 (35%), Gaps = 11/201 (5%)
Frame = -3
Query: 179 PPAAVDGSPSTSQGPIISHSPQRQGSTSNTKSSSMANNNLPNRPRPMRRISGSNIGPCPA 238
PP + P S PI SP S T S+ ++ P P I S P
Sbjct: 684 PPTLIHPLPPPSPYPI---SPPPSLDISITAPSTRPSSLTPTPASPPPPIPRS-----PH 529
Query: 239 QSSLS----NPPTPPPLPPYPVYQLPPAVSYPNMLPS----IPDSSPRDHHRNNNW---D 287
SS S +PP PP L P+ ++ L ++P+ P+ +P PR ++
Sbjct: 528 PSSFSPHSRSPPRPPWLRPFNIHPLISNSAFPSSSPAELRFLPPPRPRAPRPTPSFPRHS 349
Query: 288 ARPFVGGSSRRGHFGSHPRGDGSYHQNSYSSRREHDRGNYANTRDAHAPQPRMPPRGILR 347
A P + R H HP + S + D ++ P P PPR
Sbjct: 348 AAPPLHAPYRLPHLFLHPHSPPAVIPLSLLPVQGPDLPPVSSPPPTPPPPPA-PPRPAAP 172
Query: 348 PPPPSTAAFLGPQTIGPFPAP 368
PP S + P TI P+P
Sbjct: 171 PPSLSISINPAPHTIYILPSP 109
Score = 31.6 bits (70), Expect = 0.82
Identities = 55/213 (25%), Positives = 78/213 (35%), Gaps = 2/213 (0%)
Frame = -3
Query: 51 SQSQSPTGIHQSLPSSSSSSSSSLTTVDQPPPSDDSPKAAVVS--SSPKAVVVSSPLPAP 108
S PT IH P S S PPPS D A + SS S P P P
Sbjct: 696 SSPPPPTLIHPLPPPSPYPIS--------PPPSLDISITAPSTRPSSLTPTPASPPPPIP 541
Query: 109 MEKNSNTIASDGGDGGNAGGSKKPAWNKQPLNGVAVEIGPVMGAESWPALSESAKIPGKL 168
+ ++ + ++ +P W + P N I P++ ++P+ S A++
Sbjct: 540 RSPHPSSFSP------HSRSPPRPPWLR-PFN-----IHPLISNSAFPS-SSPAELRFLP 400
Query: 169 PPESSSSKIAPPAAVDGSPSTSQGPIISHSPQRQGSTSNTKSSSMANNNLPNRPRPMRRI 228
PP + + P P S P + H+P R S A L P +
Sbjct: 399 PPRPRAPRPTP-----SFPRHSAAPPL-HAPYRLPHLFLHPHSPPAVIPLSLLP-----V 253
Query: 229 SGSNIGPCPAQSSLSNPPTPPPLPPYPVYQLPP 261
G ++ P S PPTPPP P P PP
Sbjct: 252 QGPDLPPVS-----SPPPTPPPPPAPPRPAAPP 169
Score = 30.0 bits (66), Expect = 2.4
Identities = 29/100 (29%), Positives = 36/100 (36%), Gaps = 23/100 (23%)
Frame = -1
Query: 336 PQPRMPPRGILRPPPPSTAAFLGPQTIGPFPA-PVAYPDFYYFPTVPLDPF--------- 385
P P PP L P P T P+ P PA P+A+P F + P P
Sbjct: 581 PSPLPPPHHPLLSPVPPT-----PRRSLPIPAHPLAHPGFALSTSTPSSPIPPSHPLHPL 417
Query: 386 ------------RG-MPVFPHMPSPATFFPAAESSLSNVI 412
RG +P P +P P F P S S+ I
Sbjct: 416 NCAFFPHPAPAPRGPLPPSPAIPLPPRFMPLIGSPTSSSI 297
>TC76720 similar to GP|11121502|emb|CAC14888. putative extensin {Nicotiana
sylvestris}, partial (51%)
Length = 1290
Score = 40.0 bits (92), Expect = 0.002
Identities = 35/104 (33%), Positives = 43/104 (40%), Gaps = 7/104 (6%)
Frame = +1
Query: 53 SQSPTGIHQSLPSSSSSSSSSLTTVDQPPPSDDSPKAA----VVSSSPKAVVVSSPLPAP 108
S SPT I P++S + + T P P DSP A SSSP SSP P+P
Sbjct: 598 SPSPTPIPTPSPANSPPAPTPTPT---PSPHSDSPPAPSPDNSPSSSPSPSPSSSPAPSP 768
Query: 109 MEKNSNTIASD---GGDGGNAGGSKKPAWNKQPLNGVAVEIGPV 149
E N S G DG ++GG GV +G V
Sbjct: 769 DEAADNNAISHTGIGEDGKSSGGGMSSGKKAGIAVGVIAAVGVV 900
Score = 32.7 bits (73), Expect = 0.37
Identities = 22/68 (32%), Positives = 30/68 (43%)
Frame = +1
Query: 229 SGSNIGPCPAQSSLSNPPTPPPLPPYPVYQLPPAVSYPNMLPSIPDSSPRDHHRNNNWDA 288
S + P PA +S N P+P P+P PPA + P PS SP +N+ +
Sbjct: 550 SADSPAPTPATNSSLNSPSPTPIPTPSPANSPPAPT-PTPTPSPHSDSPPAPSPDNSPSS 726
Query: 289 RPFVGGSS 296
P SS
Sbjct: 727 SPSPSPSS 750
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.309 0.128 0.384
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 19,680,777
Number of Sequences: 36976
Number of extensions: 440492
Number of successful extensions: 9753
Number of sequences better than 10.0: 555
Number of HSP's better than 10.0 without gapping: 4718
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 7552
length of query: 546
length of database: 9,014,727
effective HSP length: 101
effective length of query: 445
effective length of database: 5,280,151
effective search space: 2349667195
effective search space used: 2349667195
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 61 (28.1 bits)
Medicago: description of AC146790.7


