

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
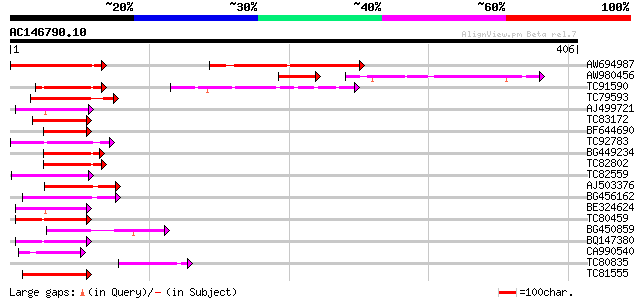
Query= AC146790.10 - phase: 0
(406 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AW694987 weakly similar to GP|10177840|dbj gene_id:MCD7.17~pir||... 74 8e-14
AW980456 65 8e-14
TC91590 weakly similar to GP|10178235|dbj|BAB11667. gene_id:MDJ2... 53 2e-07
TC79593 similar to GP|10177368|dbj|BAB10659. contains similarity... 51 9e-07
AJ499721 weakly similar to PIR|T48291|T482 hypothetical protein ... 49 5e-06
TC83172 weakly similar to GP|20042907|gb|AAM08735.1 Hypothetical... 48 8e-06
BF644690 weakly similar to GP|20042923|gb Putative copia-type po... 47 1e-05
TC92783 similar to PIR|T49020|T49020 hypothetical protein F3C22.... 47 2e-05
BG449234 weakly similar to GP|8979942|gb|A Contains a F-box PF|0... 45 4e-05
TC82802 similar to GP|9802755|gb|AAF99824.1| Unknown protein {Ar... 45 4e-05
TC82559 weakly similar to PIR|T48313|T48313 hypothetical protein... 45 5e-05
AJ503376 weakly similar to PIR|T47778|T477 hypothetical protein ... 45 5e-05
BG456162 weakly similar to PIR|T47890|T478 hypothetical protein ... 45 5e-05
BE324624 weakly similar to GP|20042914|gb| Unknown protein {Oryz... 44 9e-05
TC80459 weakly similar to GP|22831085|dbj|BAC15947. contains EST... 44 9e-05
BG450859 weakly similar to GP|21553400|gb| unknown {Arabidopsis ... 44 1e-04
BQ147380 weakly similar to GP|6017113|gb|A hypothetical protein ... 44 1e-04
CA990540 weakly similar to GP|10092259|gb| hypothetical protein;... 44 1e-04
TC80835 weakly similar to PIR|F84747|F84747 probable SWI/SNF com... 44 1e-04
TC81555 weakly similar to GP|20042914|gb|AAM08742.1 Unknown prot... 44 1e-04
>AW694987 weakly similar to GP|10177840|dbj
gene_id:MCD7.17~pir||T04055~similar to unknown protein
{Arabidopsis thaliana}, partial (5%)
Length = 665
Score = 74.3 bits (181), Expect = 8e-14
Identities = 55/114 (48%), Positives = 69/114 (60%), Gaps = 3/114 (2%)
Frame = +1
Query: 144 FVNFVGRALLLTSISSMELERFSLLINNKRDISLQNTWIS-SILNRRVKILRIHSSFYQL 202
FV+FV RALLLT E SL ++ + D+SL + W +L+R +K LRIHS F +L
Sbjct: 265 FVSFVTRALLLT-----RTETISLSLSGQYDLSLLDAWFGIMLLDRTLKNLRIHSHF-KL 426
Query: 203 PFSALTSHYLFNCTSL-EELELVLHVSSTIKFPSI-SVHFGHLKLLKLYGIFFK 254
PFS S+ LF T L E+LEL S IK PS +HFG+LK LKL I FK
Sbjct: 427 PFSTFASNSLFKITLLLEKLELHPESISRIKVPSKPDIHFGNLKHLKLCKIKFK 588
Score = 64.7 bits (156), Expect = 6e-11
Identities = 35/69 (50%), Positives = 45/69 (64%)
Frame = +1
Query: 1 MEAGSSKVHKSTQMASVIEEDTINSKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRWTS 60
ME+G + + + ED + KL ESLI ++SFLPTKD VRTCVLSK+W +RWT
Sbjct: 34 MESGLTITAQKHRKNKSTCEDDLFYKLTESLITHIISFLPTKDAVRTCVLSKQWKHRWT- 210
Query: 61 SITKLDLDD 69
+TKL L D
Sbjct: 211 FLTKLSLLD 237
>AW980456
Length = 779
Score = 64.7 bits (156), Expect(2) = 8e-14
Identities = 53/149 (35%), Positives = 77/149 (51%), Gaps = 6/149 (4%)
Frame = +3
Query: 241 GHLKLLKLYGIFFKIDTS---SDCLTLNLPLLRKFDIKNCNWSGGKDLIVEAPLLEIVSI 297
G LK+++L GI DT+ S L+L LP+L +NC+W G +IV+APLLE + +
Sbjct: 363 GSLKVVRLRGI----DTNCSLSGYLSLTLPVLETLVTRNCDWLG*V-VIVKAPLLENIHM 527
Query: 298 EQDIEFYNAASHDLHSQSIKFNALHLKQFTYSGYGTAQLIHLFDHGFLSFDSAEIIC--- 354
QD + + H+ S I+F+ HLK+FT+ GY +Q + L D A I
Sbjct: 528 VQD---HTPSFHEPCSCQIEFSDCHLKEFTFDGYAISQPVILSDPSIAKNACANIKLSKW 698
Query: 355 KPSFPETERIPFLVHLLKQFHRVKSIKIE 383
+ PE F LL QF + KSI +
Sbjct: 699 QDGTPEAGLRAFA--LLNQFSQAKSITFD 779
Score = 29.6 bits (65), Expect(2) = 8e-14
Identities = 15/30 (50%), Positives = 19/30 (63%)
Frame = +1
Query: 193 LRIHSSFYQLPFSALTSHYLFNCTSLEELE 222
LR+ SS ++ FS L S LFN L+ELE
Sbjct: 277 LRMRSSRFEWAFSGLASQSLFNSIFLDELE 366
>TC91590 weakly similar to GP|10178235|dbj|BAB11667. gene_id:MDJ22.3~unknown
protein {Arabidopsis thaliana}, partial (9%)
Length = 712
Score = 53.1 bits (126), Expect = 2e-07
Identities = 49/139 (35%), Positives = 77/139 (55%), Gaps = 4/139 (2%)
Frame = +2
Query: 116 RDAEVYERRVEIDFESGMKIESGKK---EQQFVNFVGRALLLTSISSMELERFSL-LINN 171
R +V++ +DF+ + + S KK ++QFVNFV + L+ + SS ++ FSL L ++
Sbjct: 257 RWVDVWKCITNLDFDDSL-LGSRKKRMQKEQFVNFVEKVLIHFTNSS--IQSFSLSLTSH 427
Query: 172 KRDISLQNTWISSILNRRVKILRIHSSFYQLPFSALTSHYLFNCTSLEELELVLHVSSTI 231
+ D S + WIS IL RRV+ ++H + F L S LF C SL ++L L + T+
Sbjct: 428 QYDASKLSEWISFILERRVQ--KLHIQYADKVF--LPSDSLFRCNSL--VDLTLQMRCTL 589
Query: 232 KFPSISVHFGHLKLLKLYG 250
P ISV +L+ L G
Sbjct: 590 SLP-ISVCLQNLQKLSFSG 643
Score = 49.3 bits (116), Expect = 3e-06
Identities = 26/51 (50%), Positives = 35/51 (67%)
Frame = +2
Query: 19 EEDTINSKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRWTSSITKLDLDD 69
E D I S L ES+++++LSF+P D V T VLS+RW++ W IT LD DD
Sbjct: 158 ERDMI-STLHESILSQILSFIPIVDAVSTSVLSRRWVDVW-KCITNLDFDD 304
>TC79593 similar to GP|10177368|dbj|BAB10659. contains similarity to heat
shock transcription factor HSF30~gene_id:K16L22.13,
partial (6%)
Length = 713
Score = 50.8 bits (120), Expect = 9e-07
Identities = 26/63 (41%), Positives = 38/63 (60%)
Frame = +1
Query: 16 SVIEEDTINSKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRWTSSITKLDLDDIDLSYN 75
S+ EE+ S LP+SL++ +LSFLP+KD T VLSKRW W L ++ + +N
Sbjct: 526 SIPEEEDRISALPDSLVHHILSFLPSKDATATTVLSKRWKPLW--------LLELIIHFN 681
Query: 76 YNP 78
+ P
Sbjct: 682 HQP 690
>AJ499721 weakly similar to PIR|T48291|T482 hypothetical protein F9G14.10 -
Arabidopsis thaliana, partial (8%)
Length = 502
Score = 48.5 bits (114), Expect = 5e-06
Identities = 24/66 (36%), Positives = 36/66 (54%), Gaps = 10/66 (15%)
Frame = +1
Query: 5 SSKVHKSTQMASVIEEDTIN----------SKLPESLINRVLSFLPTKDVVRTCVLSKRW 54
S V K ++ ++I + N +P+ LI+ +LSF+ T+D +RTCVLSKRW
Sbjct: 217 SGSVMKRSKKTTLISQHVYNCVATGHQDRLGDMPDCLIHHILSFMETRDAIRTCVLSKRW 396
Query: 55 MNRWTS 60
W S
Sbjct: 397 RYIWKS 414
>TC83172 weakly similar to GP|20042907|gb|AAM08735.1 Hypothetical protein
{Oryza sativa (japonica cultivar-group)}, partial (31%)
Length = 1097
Score = 47.8 bits (112), Expect = 8e-06
Identities = 22/42 (52%), Positives = 29/42 (68%)
Frame = +2
Query: 17 VIEEDTINSKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRW 58
V E++ S LP+ +I +LSFL T D V+TC+LSKRW N W
Sbjct: 191 VKEDEDRLSDLPDCVILHILSFLDTIDAVQTCILSKRWNNLW 316
>BF644690 weakly similar to GP|20042923|gb Putative copia-type polyprotein
{Oryza sativa (japonica cultivar-group)}, partial (2%)
Length = 690
Score = 47.0 bits (110), Expect = 1e-05
Identities = 22/34 (64%), Positives = 26/34 (75%)
Frame = +1
Query: 25 SKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRW 58
S LP S+I +LSFL TKD VRTCVLS+RW + W
Sbjct: 175 SDLPNSVILCILSFLNTKDGVRTCVLSRRWKDIW 276
>TC92783 similar to PIR|T49020|T49020 hypothetical protein F3C22.70 -
Arabidopsis thaliana (fragment), partial (7%)
Length = 679
Score = 46.6 bits (109), Expect = 2e-05
Identities = 31/76 (40%), Positives = 42/76 (54%), Gaps = 1/76 (1%)
Frame = +1
Query: 1 MEAGSSKVHKSTQMASVIEEDTINSKLPESLINRVLSFLPTKDVVRTC-VLSKRWMNRWT 59
ME S K K + I D IN+ LP+SL+ +LSFLPTK V T ++S+RW + W
Sbjct: 34 MEDTSLKTFKRRRKTEQINTDWINT-LPDSLLCHILSFLPTKITVTTIPLVSRRWRHLWE 210
Query: 60 SSITKLDLDDIDLSYN 75
+ L + D L YN
Sbjct: 211 N----LQVFDFYLKYN 246
>BG449234 weakly similar to GP|8979942|gb|A Contains a F-box PF|00646 domain.
{Arabidopsis thaliana}, partial (6%)
Length = 655
Score = 45.4 bits (106), Expect = 4e-05
Identities = 21/44 (47%), Positives = 31/44 (69%)
Frame = +3
Query: 25 SKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRWTSSITKLDLD 68
S L + L+ +LSFL T++ V+TC+LSKRW+N W ++ L LD
Sbjct: 165 SVLADCLLIHILSFLNTREAVQTCILSKRWINLW-KTLPTLTLD 293
>TC82802 similar to GP|9802755|gb|AAF99824.1| Unknown protein {Arabidopsis
thaliana}, partial (50%)
Length = 864
Score = 45.4 bits (106), Expect = 4e-05
Identities = 21/45 (46%), Positives = 32/45 (70%)
Frame = +3
Query: 25 SKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRWTSSITKLDLDD 69
S LP+S+I +L LP +D VRT +LS++W +W S+IT+L D+
Sbjct: 141 SDLPQSIIETILIQLPIRDAVRTSILSRKWRYKW-STITQLVFDE 272
Score = 29.6 bits (65), Expect = 2.2
Identities = 33/122 (27%), Positives = 53/122 (43%), Gaps = 1/122 (0%)
Frame = +1
Query: 182 ISSILNRRVKILRIHSSFYQLPFSALTSHYLFNCTSLEELELVLHVSSTIKFPSISVH-F 240
IS + R IL+I ++ S H F+C L LEL S +P S F
Sbjct: 415 ISGFFSFREMILKIWILNWEKVNSLECRHVFFSCEKLTRLEL----SRCELYPPASFKGF 582
Query: 241 GHLKLLKLYGIFFKIDTSSDCLTLNLPLLRKFDIKNCNWSGGKDLIVEAPLLEIVSIEQD 300
+LK L L+ I + + + L + PLL + ++ L V AP L+ + +E +
Sbjct: 583 SYLKCLNLHQILISPE-AIESLISSCPLLENLSL---SYFDCLALTVRAPNLKYLCLEGE 750
Query: 301 IE 302
+
Sbjct: 751 FK 756
>TC82559 weakly similar to PIR|T48313|T48313 hypothetical protein F9G14.230
- Arabidopsis thaliana, partial (10%)
Length = 693
Score = 45.1 bits (105), Expect = 5e-05
Identities = 24/60 (40%), Positives = 35/60 (58%), Gaps = 1/60 (1%)
Frame = +3
Query: 2 EAGSSKVHKSTQMASVIEEDTINSKLPESLINRVLSFLP-TKDVVRTCVLSKRWMNRWTS 60
++ +S V QM ++E S+LP+ +I +LSFL T+D +RT LSKRW W S
Sbjct: 171 DSANSVVENVEQMIQIVESVDRISQLPDHVIYHILSFLRNTRDAIRTKCLSKRWRTLWFS 350
>AJ503376 weakly similar to PIR|T47778|T477 hypothetical protein F17J16.10 -
Arabidopsis thaliana, partial (7%)
Length = 407
Score = 45.1 bits (105), Expect = 5e-05
Identities = 23/55 (41%), Positives = 38/55 (68%), Gaps = 1/55 (1%)
Frame = +3
Query: 26 KLPESLINRVLSFLPTKDVVRTCVLSKRWMNRWTSSITKLDLD-DIDLSYNYNPL 79
+LP+ +I+ + S L KD+V+T VLSKRW++ W S ++DL+ D+ ++YN L
Sbjct: 156 ELPDCVISYIFSKLCLKDLVKTSVLSKRWLHEWRS---RIDLNFDLHNMFDYNTL 311
>BG456162 weakly similar to PIR|T47890|T478 hypothetical protein T4C21.200 -
Arabidopsis thaliana, partial (6%)
Length = 688
Score = 45.1 bits (105), Expect = 5e-05
Identities = 22/71 (30%), Positives = 43/71 (59%), Gaps = 1/71 (1%)
Frame = +3
Query: 10 KSTQMASVIEEDTINSKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRWTSSITKLDLD- 68
KS + ++ + + SKLP+ +++ + S L KD+V+T LSK+W++ W + DL+
Sbjct: 84 KSVHLPAMKRCENLKSKLPDCIVSYIFSKLSMKDLVKTSTLSKQWLHEWG---FRTDLNF 254
Query: 69 DIDLSYNYNPL 79
D+ ++YN +
Sbjct: 255 DLQNMFHYNTI 287
>BE324624 weakly similar to GP|20042914|gb| Unknown protein {Oryza sativa
(japonica cultivar-group)}, partial (7%)
Length = 671
Score = 44.3 bits (103), Expect = 9e-05
Identities = 24/61 (39%), Positives = 34/61 (55%), Gaps = 7/61 (11%)
Frame = +1
Query: 5 SSKVHKSTQMASVIEEDTIN-------SKLPESLINRVLSFLPTKDVVRTCVLSKRWMNR 57
S K+ +M + + +T N S LPE +I +LSFL +K V+TCVLS RW +
Sbjct: 124 SKKIRIQEKMKKIRQCETENEENEDKLSDLPECVILHILSFLDSKHAVQTCVLSTRWKHL 303
Query: 58 W 58
W
Sbjct: 304 W 306
>TC80459 weakly similar to GP|22831085|dbj|BAC15947. contains EST
AU163066(S15063)~unknown protein {Oryza sativa
(japonica cultivar-group)}, partial (4%)
Length = 1116
Score = 44.3 bits (103), Expect = 9e-05
Identities = 23/54 (42%), Positives = 33/54 (60%)
Frame = +2
Query: 5 SSKVHKSTQMASVIEEDTINSKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRW 58
+++V K +M EED + S LP+ +++ +LS LP KD RT VLSK W W
Sbjct: 35 NNQVKKKKKMEHEEEEDRL-SNLPKIILHNILSRLPKKDGARTSVLSKAWEETW 193
>BG450859 weakly similar to GP|21553400|gb| unknown {Arabidopsis thaliana},
partial (11%)
Length = 549
Score = 43.9 bits (102), Expect = 1e-04
Identities = 25/90 (27%), Positives = 40/90 (43%), Gaps = 2/90 (2%)
Frame = +2
Query: 27 LPESLINRVLSFLPTKDVVRTCVLSKRWMNRWTSSITKLDLDDIDLSYNYNPLCSYCEEI 86
LP+ ++ +LSF+PTK T +LSKRW++ W Y P+ + E
Sbjct: 137 LPDEILIHILSFVPTKQAFTTSILSKRWIHLW----------------RYVPILDFTETN 268
Query: 87 C--TDYCYDYDKYKLTVCPRCHVCGNHKAN 114
D ++++ +V H GNH N
Sbjct: 269 LEDRDSVIRFEEFIFSVIRSRHSAGNHSIN 358
>BQ147380 weakly similar to GP|6017113|gb|A hypothetical protein {Arabidopsis
thaliana}, partial (7%)
Length = 647
Score = 43.9 bits (102), Expect = 1e-04
Identities = 22/54 (40%), Positives = 32/54 (58%)
Frame = +2
Query: 5 SSKVHKSTQMASVIEEDTINSKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRW 58
S K K T+ E D I S LP + ++++S+LP +D VRT VLS+ W +W
Sbjct: 251 SKKERKRTKSTLDAEPDRI-SWLPGHVTDQIMSYLPIRDAVRTSVLSRNWRKKW 409
>CA990540 weakly similar to GP|10092259|gb| hypothetical protein; 40655-42260
{Arabidopsis thaliana}, partial (9%)
Length = 761
Score = 43.5 bits (101), Expect = 1e-04
Identities = 21/48 (43%), Positives = 29/48 (59%)
Frame = +1
Query: 7 KVHKSTQMASVIEEDTINSKLPESLINRVLSFLPTKDVVRTCVLSKRW 54
K HK T E S+LP+ ++ ++SFL +D VRTC+LSKRW
Sbjct: 61 KSHKGTDET---EAGNRISELPDCILLHIMSFLEARDAVRTCILSKRW 195
>TC80835 weakly similar to PIR|F84747|F84747 probable SWI/SNF complex
subunit SW13 [imported] - Arabidopsis thaliana, partial
(34%)
Length = 830
Score = 43.5 bits (101), Expect = 1e-04
Identities = 18/53 (33%), Positives = 32/53 (59%)
Frame = +2
Query: 79 LCSYCEEICTDYCYDYDKYKLTVCPRCHVCGNHKANLRDAEVYERRVEIDFES 131
+CS C+ +C C+ +K +T+C RC + GN++ + + E +RVEI E+
Sbjct: 572 ICSGCKNLCVMACFACEKNNMTLCARCFIRGNYRIGMSNTEF--KRVEISEET 724
>TC81555 weakly similar to GP|20042914|gb|AAM08742.1 Unknown protein {Oryza
sativa (japonica cultivar-group)}, partial (7%)
Length = 688
Score = 43.5 bits (101), Expect = 1e-04
Identities = 22/49 (44%), Positives = 31/49 (62%)
Frame = +2
Query: 10 KSTQMASVIEEDTINSKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRW 58
K +++S E + S LPES+I +LSFL TK V+TCVLS + + W
Sbjct: 131 KKVKLSSECENEDRLSDLPESVILHILSFLNTKHAVQTCVLSPIYKDLW 277
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.324 0.138 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,633,976
Number of Sequences: 36976
Number of extensions: 247320
Number of successful extensions: 1833
Number of sequences better than 10.0: 97
Number of HSP's better than 10.0 without gapping: 1811
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1829
length of query: 406
length of database: 9,014,727
effective HSP length: 98
effective length of query: 308
effective length of database: 5,391,079
effective search space: 1660452332
effective search space used: 1660452332
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.5 bits)
S2: 59 (27.3 bits)
Medicago: description of AC146790.10


