

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
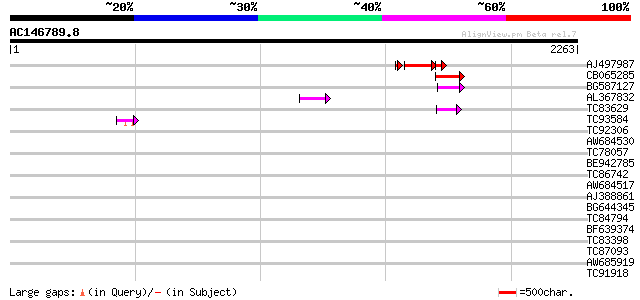
Query= AC146789.8 + phase: 0 /pseudo
(2263 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AJ497987 weakly similar to GP|9927273|dbj Similar to Arabidopsis... 230 1e-79
CB065285 weakly similar to PIR|A84500|A84 probable retroelement ... 192 1e-48
BG587127 weakly similar to PIR|H84506|H84 probable retroelement ... 65 4e-10
AL367832 weakly similar to GP|14091845|gb Putative retroelement ... 62 2e-09
TC83629 weakly similar to GP|6692261|gb|AAF24611.1| putative RNa... 45 2e-04
TC93584 knotted class I homeodomain KNOX 44 7e-04
TC92306 43 0.001
AW684530 similar to GP|15724197|gb At2g39050/T7F6.22 {Arabidopsi... 40 0.008
TC78057 weakly similar to PIR|T03962|T03962 r40g3 protein - rice... 39 0.017
BE942785 38 0.039
TC86742 similar to PIR|T01541|T01541 hypothetical protein A_IG00... 37 0.066
AW684517 similar to GP|9927273|dbj Similar to Arabidopsis thalia... 36 0.19
AJ388861 weakly similar to GP|1754989|gb| proline-rich protein P... 35 0.25
BG644345 weakly similar to GP|19697333|gb putative protein poten... 35 0.25
TC84794 similar to PIR|S54156|S54156 extensin-like protein - cow... 35 0.43
BF639374 similar to GP|8132441|gb| extensin {Pisum sativum}, par... 35 0.43
TC83398 similar to PIR|T05150|T05150 hypothetical protein F18E5.... 35 0.43
TC87093 homologue to GP|10177817|dbj|BAB11183. gene_id:MKD15.14~... 33 0.96
AW685919 homologue to PIR|S25298|S252 extensin (clone Tom J-10) ... 33 1.6
TC91918 homologue to GP|9369375|gb|AAF87124.1| F10A5.29 {Arabido... 27 1.9
>AJ497987 weakly similar to GP|9927273|dbj Similar to Arabidopsis thaliana
chromosome II BAC F26H6; putative retroelement pol
polyprotein, partial (1%)
Length = 636
Score = 230 bits (586), Expect(2) = 1e-79
Identities = 121/127 (95%), Positives = 121/127 (95%)
Frame = -3
Query: 1577 RRVVL*PGLPDASVII*LITLRGWYLKWIQSSTYLRSPL*QEGLHGGRCYYLNMISSTVP 1636
R VVL*PGLP ASVII*LITL GWYLKWIQSSTYLRSPL*QEGLHGGRCYY NMISSTVP
Sbjct: 634 RHVVL*PGLPSASVII*LITLLGWYLKWIQSSTYLRSPL*QEGLHGGRCYYRNMISSTVP 455
Query: 1637 RRQLKAVFLLIT*LTNHLKTIDLSSLTFLMKRLCI*R*KIVTSHYSEKDLIQIQCGV*YL 1696
RRQLKAVFLLIT LTNHLKTIDLSSLTFLMKRLCI*R*KIVTSHYSEK LIQIQCGV*YL
Sbjct: 454 RRQLKAVFLLITWLTNHLKTIDLSSLTFLMKRLCI*R*KIVTSHYSEKVLIQIQCGV*YL 275
Query: 1697 MGLSMYT 1703
MGLSMYT
Sbjct: 274 MGLSMYT 254
Score = 87.8 bits (216), Expect(2) = 1e-79
Identities = 40/44 (90%), Positives = 43/44 (96%)
Frame = -2
Query: 1699 LSMYTNGIGAVLLTPKGAHIPFTARLRFDCTNNIAEYEACIMGI 1742
+++Y NGIGAVLLTPKGAHIPFTARLRFDCTNNIAEYEACIMGI
Sbjct: 266 VNVYGNGIGAVLLTPKGAHIPFTARLRFDCTNNIAEYEACIMGI 135
Score = 45.8 bits (107), Expect = 2e-04
Identities = 23/29 (79%), Positives = 24/29 (82%)
Frame = +3
Query: 1538 KIPWVVCSDNKMKPEEKSTPFII*ARSLL 1566
+I VCSDNKMKPEEKSTPFII*A S L
Sbjct: 24 RIHGFVCSDNKMKPEEKSTPFII*AGSCL 110
>CB065285 weakly similar to PIR|A84500|A84 probable retroelement gag/pol
polyprotein [imported] - Arabidopsis thaliana, partial
(2%)
Length = 592
Score = 192 bits (488), Expect = 1e-48
Identities = 89/115 (77%), Positives = 106/115 (91%)
Frame = -2
Query: 1699 LSMYTNGIGAVLLTPKGAHIPFTARLRFDCTNNIAEYEACIMGIEEAIDLRIKNIEIYGD 1758
++ Y GIGAV+++P+G +IPFTAR+ F+CTNN+AEYEACI GIEEAID+RIK+++IYGD
Sbjct: 510 VNAYGKGIGAVIVSPQGHYIPFTARILFECTNNMAEYEACIFGIEEAIDMRIKHLDIYGD 331
Query: 1759 SALVINQIKGKWETLHAGLIPYRDYARRLLTFFNKVELHHIPRDENQMADALATV 1813
SALVINQIKG+WET HA LIPYRDYARRLLT+F KVELHHIPRDENQMADALAT+
Sbjct: 330 SALVINQIKGEWETHHANLIPYRDYARRLLTYFTKVELHHIPRDENQMADALATL 166
>BG587127 weakly similar to PIR|H84506|H84 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(13%)
Length = 415
Score = 64.7 bits (156), Expect = 4e-10
Identities = 39/107 (36%), Positives = 54/107 (50%)
Frame = +3
Query: 1707 GAVLLTPKGAHIPFTARLRFDCTNNIAEYEACIMGIEEAIDLRIKNIEIYGDSALVINQI 1766
G L +P + + RL F +NN YEA I G+ A L+I+NI Y DS LV +Q
Sbjct: 3 GIRLTSPTNEILKQSFRLEFHASNNETNYEALIAGVRLAHGLKIRNIHAYCDSQLVASQF 182
Query: 1767 KGKWETLHAGLIPYRDYARRLLTFFNKVELHHIPRDENQMADALATV 1813
G++E + Y ++L + L IPR EN ADALA +
Sbjct: 183 SGEYEARDELMDTYLKLVQKLAQKLDYFALTRIPRSENVQADALAAL 323
>AL367832 weakly similar to GP|14091845|gb Putative retroelement {Oryza
sativa}, partial (2%)
Length = 384
Score = 62.4 bits (150), Expect = 2e-09
Identities = 47/127 (37%), Positives = 68/127 (53%)
Frame = -2
Query: 1155 KRGRPFSLMRTN*K*LIWAPKKTRKKSRLGHRSKQVLKSK**SSSKNMLMCLPGPTKICQ 1214
K+G+PFSL R P++T ++SR ++ LK + SSS+N + L KICQ
Sbjct: 383 KKGKPFSLTRKRLSLST*VPRRTSERSRSVLL*RKGLKGRSSSSSENTWIFLHARMKICQ 204
Query: 1215 V*TLILWYITYL*NLNVRQSNKS*EEHVLTWLSKSKKRYRNRSTLVSSSHQITPNG*LT* 1274
V* L LW I +LNV S ++*E + WLS+ + R+++R V S NG
Sbjct: 203 V*ILKLWNIGSQQSLNVLPSGRN*EGLIQIWLSRLRVRFKSRLMRVFS*QLSILNGLPIL 24
Query: 1275 CLFQRKM 1281
L QR+M
Sbjct: 23 YLCQRRM 3
>TC83629 weakly similar to GP|6692261|gb|AAF24611.1| putative RNase H
{Arabidopsis thaliana}, partial (21%)
Length = 1071
Score = 45.4 bits (106), Expect = 2e-04
Identities = 32/100 (32%), Positives = 45/100 (45%)
Frame = +2
Query: 1705 GIGAVLLTPKGAHIPFTARLRFDCTNNIAEYEACIMGIEEAIDLRIKNIEIYGDSALVIN 1764
G GA+LL G+ + + T AEY A ++G++ A K + GDS LVIN
Sbjct: 602 GAGALLLAEDGSLLYGFRQGLGHQTKESAEYRALLLGLKHASMKGFKYVTAKGDSELVIN 781
Query: 1765 QIKGKWETLHAGLIPYRDYARRLLTFFNKVELHHIPRDEN 1804
QI W+ L A L F+ + HI R+ N
Sbjct: 782 QILDPWKIKDEHLKKLCAEALELSDNFHSFRIQHISRERN 901
>TC93584 knotted class I homeodomain KNOX
Length = 1161
Score = 43.9 bits (102), Expect = 7e-04
Identities = 26/103 (25%), Positives = 46/103 (44%), Gaps = 16/103 (15%)
Frame = +1
Query: 425 ILNSHKILHHKILNNHTNNNHTNNSHTNNTLNKI--------SNNNLINNAHNNLDHQEC 476
+ +SH HH I +N+ NNN+ NN++ NNT +NNN N +NN
Sbjct: 106 VTSSHHNAHHPINSNNNNNNNNNNTNANNTTGLFLPIPNSTNNNNNHYTNCNNNTSSIML 285
Query: 477 RLTQYQSPMQNYF--------PAYSRKTLSKLEQLLLSQKNYH 511
+ +P Y+ + S + S ++ +++ +YH
Sbjct: 286 QNNHQNTPGLGYYFMDNINNHGSSSSSSSSSVKSKIMAHPHYH 414
>TC92306
Length = 521
Score = 43.1 bits (100), Expect = 0.001
Identities = 23/73 (31%), Positives = 36/73 (48%)
Frame = +2
Query: 1739 IMGIEEAIDLRIKNIEIYGDSALVINQIKGKWETLHAGLIPYRDYARRLLTFFNKVELHH 1798
I+G+ EA + +++ + GD V Q G W+ + L + A L + F V + H
Sbjct: 2 ILGLNEARNQGYEHVHVRGDFQXVCKQFXGSWKVNNPNLRNLCNXAVELKSNFKSVSVEH 181
Query: 1799 IPRDENQMADALA 1811
+PR N ADA A
Sbjct: 182 VPRGXNXAADAQA 220
>AW684530 similar to GP|15724197|gb At2g39050/T7F6.22 {Arabidopsis thaliana},
partial (33%)
Length = 643
Score = 40.4 bits (93), Expect = 0.008
Identities = 27/76 (35%), Positives = 36/76 (46%), Gaps = 2/76 (2%)
Frame = +3
Query: 383 KLIQPINTLPPSHPPLTLFNKQITILKYRNILKYRNILKYHNILNSHKILHHKILNNH-- 440
K P T PP+ P TL +T NI+ + Y N H HH I+NNH
Sbjct: 15 KWSSPSITTPPT*P--TLITTVVTTTTTNNIIHHLVTTIYLPSTNHH---HHHIINNHLS 179
Query: 441 TNNNHTNNSHTNNTLN 456
T +HT++ H NN +N
Sbjct: 180 TLTHHTHHHHNNNHIN 227
>TC78057 weakly similar to PIR|T03962|T03962 r40g3 protein - rice, partial
(50%)
Length = 878
Score = 39.3 bits (90), Expect = 0.017
Identities = 25/69 (36%), Positives = 34/69 (49%), Gaps = 2/69 (2%)
Frame = +3
Query: 390 TLPPSHPPLTLFNKQITILKYRNILKYRNILKYHNILNSHKILHHKILNNH--TNNNHTN 447
T PP+ P TL +T NI+ + Y N H HH I+NNH T +HT+
Sbjct: 9 TTPPT*P--TLITTVVTTTTTNNIIHHLVTTIYLPSTNHH---HHHIINNHLSTLTHHTH 173
Query: 448 NSHTNNTLN 456
+ H NN +N
Sbjct: 174 HHHNNNHIN 200
>BE942785
Length = 460
Score = 38.1 bits (87), Expect = 0.039
Identities = 27/61 (44%), Positives = 35/61 (57%)
Frame = -3
Query: 987 HLSLSLGQTMSKGHLFKGLL*KIRIPRRMKPLSRL*KMLRRSYKQEGPQAGES**SFQKT 1046
HLSL+ Q + +G LFK L K+R PR M+ L*+M R K+ AG * SF +T
Sbjct: 323 HLSLA*KQGLQRGPLFKD*LSKVRSPRGMELQWLL*RMHREPSKRVKLPAGAG*YSFVRT 144
Query: 1047 S 1047
S
Sbjct: 143 S 141
>TC86742 similar to PIR|T01541|T01541 hypothetical protein A_IG005I10.16 -
Arabidopsis thaliana, partial (19%)
Length = 2073
Score = 37.4 bits (85), Expect = 0.066
Identities = 30/86 (34%), Positives = 47/86 (53%), Gaps = 7/86 (8%)
Frame = -1
Query: 1733 AEYEACIMGIEEAIDLRIKNIEIYGDSALVIN---QIKGKWETLHAGLIPYRDYAR--RL 1787
AE+ AC++ IE+A++L + NI + DS V+N +I G IP++ R
Sbjct: 396 AEFCACMIAIEKAMELGLNNICLETDSLKVVNAFHKIVG---------IPWQMRVRWHNC 244
Query: 1788 LTFFNKVE--LHHIPRDENQMADALA 1811
+ F + + HIPR+ N +ADALA
Sbjct: 243 IRFCHSIACVCVHIPREGNLVADALA 166
>AW684517 similar to GP|9927273|dbj Similar to Arabidopsis thaliana chromosome
II BAC F26H6; putative retroelement pol polyprotein,
partial (1%)
Length = 488
Score = 35.8 bits (81), Expect = 0.19
Identities = 40/115 (34%), Positives = 54/115 (46%), Gaps = 4/115 (3%)
Frame = +2
Query: 1001 LFKGLL*KIRIPRRMKPLSRL*KMLRRSYKQEGPQAGES**SFQKTSTEKDWVSSHLLVC 1060
LFKG L K++ PRR++ L*+ RR ++++ G S* + + S K S V
Sbjct: 140 LFKGCLWKVQSPRRLELQWLL*RTPRRLFRRDRLPTGAS*SNSVRISERKVSDSPQHRVS 319
Query: 1061 PQRRRALSTVRALSMRSLK----MVLKACHEVL*RQEYPATIGLL*MFLLLLTCP 1111
P R L V LS+ LK + L C L* QE IG+ MF CP
Sbjct: 320 P---RELFIVPGLSIHLLKRWLDLFLGPC---L*YQEALRKIGMPLMFPQSCMCP 466
>AJ388861 weakly similar to GP|1754989|gb| proline-rich protein PRP2
precursor {Lupinus luteus}, partial (24%)
Length = 491
Score = 35.4 bits (80), Expect = 0.25
Identities = 18/59 (30%), Positives = 29/59 (48%)
Frame = -1
Query: 418 NILKYHNILNSHKILHHKILNNHTNNNHTNNSHTNNTLNKISNNNLINNAHNNLDHQEC 476
N L + + +H + HH NH NH SH +T N + ++ +N H N++H C
Sbjct: 452 NFLI*XHFMKNHHMNHHHKSTNHFMRNHHK*SHHQST-NHLMKSHHMNIHHQNINHLMC 279
>BG644345 weakly similar to GP|19697333|gb putative protein potential
transcriptional repressor Not4hp - Mus musculus, partial
(7%)
Length = 635
Score = 35.4 bits (80), Expect = 0.25
Identities = 15/33 (45%), Positives = 22/33 (66%)
Frame = +2
Query: 438 NNHTNNNHTNNSHTNNTLNKISNNNLINNAHNN 470
N +N+N+ NN+ +NN N SNNN N ++NN
Sbjct: 335 NFSSNSNNNNNNFSNNNNNNFSNNNNCNCSNNN 433
Score = 33.9 bits (76), Expect = 0.74
Identities = 18/54 (33%), Positives = 28/54 (51%)
Frame = +2
Query: 420 LKYHNILNSHKILHHKILNNHTNNNHTNNSHTNNTLNKISNNNLINNAHNNLDH 473
L + N N+ + NN+ +N+ NN+ +NN SNNN NN +N+ H
Sbjct: 305 LPFRNSNNNFNFSSNSNNNNNNFSNNNNNNFSNNNNCNCSNNN--NNCCSNISH 460
>TC84794 similar to PIR|S54156|S54156 extensin-like protein - cowpea
(fragment), partial (49%)
Length = 478
Score = 34.7 bits (78), Expect = 0.43
Identities = 20/63 (31%), Positives = 28/63 (43%)
Frame = +2
Query: 42 HLLLSELRPRHLLLGHYVLTLQRSPLRNVPRLGFHLSLLGRYFVLSLVKHRCPLTSTRLK 101
H L+S P H LL ++LT PL ++ + HL+ L HR TS L
Sbjct: 254 HQLMSTSHPLHPLLHRHLLTFTNHPLHHLMNIKLHLTSTSHLRHRHLHHHRLTSTSLHLH 433
Query: 102 YPY 104
P+
Sbjct: 434 LPH 442
>BF639374 similar to GP|8132441|gb| extensin {Pisum sativum}, partial (45%)
Length = 381
Score = 34.7 bits (78), Expect = 0.43
Identities = 17/51 (33%), Positives = 27/51 (52%), Gaps = 3/51 (5%)
Frame = +1
Query: 426 LNSHKILHHKILNNHTNNNHTNNSHTNNTLNKI---SNNNLINNAHNNLDH 473
L+ H+ HH +L ++H SHTN L + S N+L+++ NL H
Sbjct: 85 LHHHRCTHHLLLTITVPHHHLQRSHTNTLLLHLQFTSTNHLLHHISTNLHH 237
>TC83398 similar to PIR|T05150|T05150 hypothetical protein F18E5.40 -
Arabidopsis thaliana, partial (6%)
Length = 766
Score = 34.7 bits (78), Expect = 0.43
Identities = 28/87 (32%), Positives = 42/87 (48%), Gaps = 2/87 (2%)
Frame = +2
Query: 1727 DCTNNI--AEYEACIMGIEEAIDLRIKNIEIYGDSALVINQIKGKWETLHAGLIPYRDYA 1784
D +N+I AE A +MG++ A L I ++ Y DS +N I G HA Y
Sbjct: 122 DHSNDILFAELHAILMGLQLAQTLNIVDVVCYSDSLHYVNLINGPSVVYHA----YATLI 289
Query: 1785 RRLLTFFNKVELHHIPRDENQMADALA 1811
+ + +LH + R+ N+ AD LA
Sbjct: 290 QDIKDLIRLSKLHTL-REGNRCADFLA 367
>TC87093 homologue to GP|10177817|dbj|BAB11183. gene_id:MKD15.14~unknown
protein {Arabidopsis thaliana}, partial (49%)
Length = 1313
Score = 33.5 bits (75), Expect = 0.96
Identities = 13/32 (40%), Positives = 24/32 (74%)
Frame = +1
Query: 431 ILHHKILNNHTNNNHTNNSHTNNTLNKISNNN 462
I+H I+N++ NNN T+NS+ + T + I+++N
Sbjct: 601 IIHCFIINSNNNNNSTSNSNNSTTTSSITSSN 696
>AW685919 homologue to PIR|S25298|S252 extensin (clone Tom J-10) - tomato,
partial (23%)
Length = 663
Score = 32.7 bits (73), Expect = 1.6
Identities = 23/89 (25%), Positives = 43/89 (47%), Gaps = 1/89 (1%)
Frame = +2
Query: 381 LNKLIQPINTLPPSHPPLTLFNKQITILKYRNILKYRNILKYHNILNSHKILHHKILNNH 440
L+ L+ T HP L + TI N+ +R+ L +H + S H +L++H
Sbjct: 35 LHHLLHHHTTTNLPHPLHRLLHHHTTI----NLPHHRHPLHHHLTITSLLPHLHHLLHHH 202
Query: 441 TNNNHTNNSHT-NNTLNKISNNNLINNAH 468
T NH ++ H +T + + + +I++ H
Sbjct: 203 TTTNHLHHPHRFTSTTHPLHRSIIIHHHH 289
>TC91918 homologue to GP|9369375|gb|AAF87124.1| F10A5.29 {Arabidopsis
thaliana}, partial (38%)
Length = 1171
Score = 26.6 bits (57), Expect(2) = 1.9
Identities = 11/31 (35%), Positives = 18/31 (57%)
Frame = +2
Query: 440 HTNNNHTNNSHTNNTLNKISNNNLINNAHNN 470
H NN++ NN+ N +N N+ NN +N+
Sbjct: 377 HIQIQPNNNNNINNSFNICNNINMNNNNNNS 469
Score = 24.3 bits (51), Expect(2) = 1.9
Identities = 12/29 (41%), Positives = 15/29 (51%)
Frame = +3
Query: 414 LKYRNILKYHNILNSHKILHHKILNNHTN 442
L +N L IL H +LHH NN+ N
Sbjct: 285 LTSQNFLHPLKILLHHPLLHHLKCNNNNN 371
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.357 0.157 0.557
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 73,991,643
Number of Sequences: 36976
Number of extensions: 1129224
Number of successful extensions: 15859
Number of sequences better than 10.0: 68
Number of HSP's better than 10.0 without gapping: 4767
Number of HSP's successfully gapped in prelim test: 841
Number of HSP's that attempted gapping in prelim test: 9516
Number of HSP's gapped (non-prelim): 7414
length of query: 2263
length of database: 9,014,727
effective HSP length: 112
effective length of query: 2151
effective length of database: 4,873,415
effective search space: 10482715665
effective search space used: 10482715665
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.7 bits)
S2: 66 (30.0 bits)
Medicago: description of AC146789.8


