

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146789.1 + phase: 0
(431 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
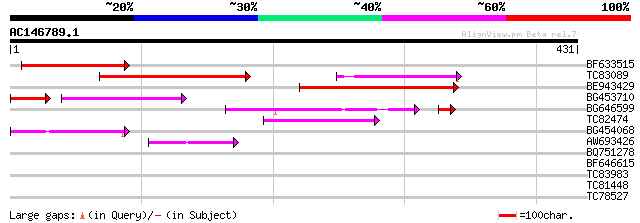
Score E
Sequences producing significant alignments: (bits) Value
BF633515 150 9e-37
TC83089 144 7e-35
BE943429 130 1e-30
BG453710 weakly similar to GP|21537057|gb| unknown {Arabidopsis ... 71 5e-20
BG646599 75 4e-15
TC82474 70 2e-12
BG454068 weakly similar to GP|22202804|dbj hypothetical protein~... 67 1e-11
AW693426 homologue to GP|23307580|dbj similar to serine/threonin... 45 7e-05
BQ751278 similar to PIR|T37665|T37 probable t-complex protein 1 ... 30 2.4
BF646615 homologue to PIR|T48395|T483 GT2-like protein - Arabido... 29 3.1
TC83983 similar to GP|19386796|dbj|BAB86175. putative alpha-gluc... 29 3.1
TC81448 weakly similar to OMNI|NT01MC3624 toxin secretion ATP-bi... 28 5.2
TC78527 similar to GP|14030876|gb|AAK52082.1 nuclease {Nicotiana... 28 8.9
>BF633515
Length = 249
Score = 150 bits (379), Expect = 9e-37
Identities = 70/82 (85%), Positives = 75/82 (91%)
Frame = +2
Query: 10 LDDVACLMHLPIEGRMLAHGKKMPKHEGAALLMTYLGVAQHEAEKICNQEYGGYISYPRL 69
LDDVACL HLPIEGRML+HGKKMPKHEGAALLM +LGV+Q EAEKIC QEYGGYISYPRL
Sbjct: 2 LDDVACLTHLPIEGRMLSHGKKMPKHEGAALLMRHLGVSQQEAEKICGQEYGGYISYPRL 181
Query: 70 RDFYTSYLGRANVLVGTEDPEE 91
RDFYT+YLGRAN+L TEDP E
Sbjct: 182RDFYTTYLGRANLLASTEDPVE 247
>TC83089
Length = 887
Score = 144 bits (363), Expect = 7e-35
Identities = 72/116 (62%), Positives = 87/116 (74%), Gaps = 1/116 (0%)
Frame = +3
Query: 69 LRDFYTSYLGRANVLVGTEDPEEVEELERVRTYCVRCYLLYLVGCLLFGDRSNKRIELIY 128
LR++Y YL A L ++ + +ELE++RT CV+ YLLYLVGCLLFGD+SNKRIEL+Y
Sbjct: 264 LREYYEGYLDSATRLSDPQNLGDPQELEKIRTTCVKWYLLYLVGCLLFGDKSNKRIELVY 443
Query: 129 LTTMADGYAGMRNYSWGAMTLAYLYGELADACRPGHRALGGSVTLLTT-WFLKHFP 183
LTTM DGYAGMRN+SWG MTLAYLY L++ PG AL SVTL+T WFL H P
Sbjct: 444 LTTMEDGYAGMRNHSWGGMTLAYLYHCLSEVSLPGGGALERSVTLVTAGWFLCHLP 611
Score = 68.2 bits (165), Expect = 6e-12
Identities = 38/98 (38%), Positives = 52/98 (52%), Gaps = 3/98 (3%)
Frame = +3
Query: 249 SGWIMCGVRRVYRHLPERVLRQYGYVQTIPRHPTDVRDLPPPSIVQMFIDFRTHTLKADA 308
+GW +C HLPER LR YGYVQT+P PT + L P +V F++F H
Sbjct: 585 AGWFLC-------HLPERGLR*YGYVQTVPIPPTTIEPLEPAEVVTAFLEFALHVQSQQE 743
Query: 309 RGVQAGEDT-RRVADGYVLWYTRVSHPQILPP--IPGD 343
RG ED + + GY+ + RVSH ++ P +P D
Sbjct: 744 RGESVPEDEGHKHSKGYMK*FYRVSHLLMIAPAAVPED 857
>BE943429
Length = 384
Score = 130 bits (327), Expect = 1e-30
Identities = 60/122 (49%), Positives = 78/122 (63%), Gaps = 1/122 (0%)
Frame = +2
Query: 221 DRIQLDDVCWRPYEEHREIQDFEEVFWYSGWIMCGVRRVYRHLPERVLRQYGYVQTIPRH 280
D ++ V WRPYE R++ F++V WYSGWIM G +++ RHLPERVLRQYGYVQT+PR
Sbjct: 2 DSMEHCHVIWRPYEHRRDVTPFQDVCWYSGWIMAGKQKMVRHLPERVLRQYGYVQTVPRP 181
Query: 281 PTDVRDLPPPSIVQMFIDFRTHTLKADARGVQAGEDT-RRVADGYVLWYTRVSHPQILPP 339
PT + L P + F +F H L RG ED + +DGY+ W+ RVSHP I+ P
Sbjct: 182 PTTIVPLAPAEVATAFFEFVVHVLSQQDRGDPVPEDEWWKHSDGYIKWFYRVSHPLIVNP 361
Query: 340 IP 341
P
Sbjct: 362 AP 367
>BG453710 weakly similar to GP|21537057|gb| unknown {Arabidopsis thaliana},
partial (6%)
Length = 651
Score = 70.9 bits (172), Expect(2) = 5e-20
Identities = 40/95 (42%), Positives = 54/95 (56%)
Frame = -3
Query: 40 LLMTYLGVAQHEAEKICNQEYGGYISYPRLRDFYTSYLGRANVLVGTEDPEEVEELERVR 99
L+ LG++ A E G+ISYP L+ Y +L A L EE++E R R
Sbjct: 298 LMTRLLGMSDANAMAEIRTESAGHISYPTLKRVYEDHLIEARRLEDPLTREELQERARRR 119
Query: 100 TYCVRCYLLYLVGCLLFGDRSNKRIELIYLTTMAD 134
+CVR LLYLV C+LF ++N+ I+LIYL MAD
Sbjct: 118 QWCVRSLLLYLVDCVLFTYKTNRHIDLIYLDCMAD 14
Score = 44.7 bits (104), Expect(2) = 5e-20
Identities = 19/31 (61%), Positives = 25/31 (80%)
Frame = -1
Query: 1 MPMGEMTVTLDDVACLMHLPIEGRMLAHGKK 31
+PMGEMT+TLDDV L+H+PI G ML H ++
Sbjct: 414 LPMGEMTITLDDVYNLLHIPIHGCMLDHDEQ 322
>BG646599
Length = 711
Score = 75.1 bits (183), Expect(2) = 4e-15
Identities = 46/151 (30%), Positives = 76/151 (49%), Gaps = 4/151 (2%)
Frame = +3
Query: 165 RALGGSVTLLTTWFLKHFPGFFSVDLNTDYLEKYPV----AARWKLEKGHGEGITYRSLL 220
+ LGG TLL W ++FP + N ++ + A RWK +G+ + R ++
Sbjct: 33 KQLGGYATLLQCWIYEYFPTLGNRVENRVACDEPGMGVARAMRWKYPRGNLKVDQIRRMI 212
Query: 221 DRIQLDDVCWRPYEEHREIQDFEEVFWYSGWIMCGVRRVYRHLPERVLRQYGYVQTIPRH 280
D + +DV W P++ HR++ F+++ SG+I V +LPER LRQ+GY+Q IPR
Sbjct: 213 DDLTPNDVIWCPFDSHRQVIPFDDICLSSGYIR-WCSNVVPYLPERCLRQFGYIQYIPR- 386
Query: 281 PTDVRDLPPPSIVQMFIDFRTHTLKADARGV 311
PPP+ +D + + A +
Sbjct: 387 -------PPPNFNTFNVDVEWNDCHSSANQI 458
Score = 23.9 bits (50), Expect(2) = 4e-15
Identities = 7/13 (53%), Positives = 10/13 (76%)
Frame = +1
Query: 327 WYTRVSHPQILPP 339
WY +VSHP ++ P
Sbjct: 517 WYYKVSHPHLIRP 555
>TC82474
Length = 992
Score = 70.1 bits (170), Expect = 2e-12
Identities = 32/88 (36%), Positives = 50/88 (56%)
Frame = -2
Query: 194 YLEKYPVAARWKLEKGHGEGITYRSLLDRIQLDDVCWRPYEEHREIQDFEEVFWYSGWIM 253
Y E P A R+ L GH Y + LDR+++DDV + Y+ H + F+++ + WI
Sbjct: 883 YTEDRPHATRFSLAMGHKSPNNYMTFLDRVEVDDVSFNQYDFHHQTCPFDDIS*FFRWIE 704
Query: 254 CGVRRVYRHLPERVLRQYGYVQTIPRHP 281
C + Y HL E V++ YG++Q+ RHP
Sbjct: 703 CDRKMNYHHLLELVMK*YGHIQSTLRHP 620
>BG454068 weakly similar to GP|22202804|dbj hypothetical protein~predicted by
GeneMark.hmm etc. {Oryza sativa (japonica
cultivar-group)}, partial (5%)
Length = 673
Score = 67.4 bits (163), Expect = 1e-11
Identities = 35/94 (37%), Positives = 52/94 (55%), Gaps = 3/94 (3%)
Frame = +3
Query: 1 MPMGEMTVTLDDVACLMHLPIEGRMLAHGKKMPKHEGAALLMTYLGVAQHEAEKICNQEY 60
+P+GEMT+ LDD++CL+H+P+ G +L H + + H+G L+ YLG+ E +
Sbjct: 387 LPVGEMTIALDDISCLLHIPVGGNLLFH-ESLSIHQGTEYLVNYLGLEFEEIAAETKRLK 563
Query: 61 GGYISYPRLRDFYTSYLGRANVLV---GTEDPEE 91
+I+Y L YTSYL A G ED E
Sbjct: 564 SAHITYDTLLSIYTSYLTEAKSYANQPGEEDSME 665
>AW693426 homologue to GP|23307580|dbj similar to serine/threonine
phosphatase PP7 {Oryza sativa (japonica
cultivar-group)}, partial (2%)
Length = 209
Score = 44.7 bits (104), Expect = 7e-05
Identities = 25/69 (36%), Positives = 36/69 (51%)
Frame = +1
Query: 106 YLLYLVGCLLFGDRSNKRIELIYLTTMADGYAGMRNYSWGAMTLAYLYGELADACRPGHR 165
+LL LVGC LF D+S + + YL D Y+WG LA+LY L A +
Sbjct: 1 FLLCLVGCTLFSDKSTFVVNVAYLEFFRD-LDSCGGYAWGVAALAHLYDNLRYASFHHMK 177
Query: 166 ALGGSVTLL 174
++ G +TL+
Sbjct: 178 SISGYLTLI 204
>BQ751278 similar to PIR|T37665|T37 probable t-complex protein 1 epsilon
subunit - fission yeast (Schizosaccharomyces pombe),
partial (26%)
Length = 596
Score = 29.6 bits (65), Expect = 2.4
Identities = 21/71 (29%), Positives = 32/71 (44%), Gaps = 1/71 (1%)
Frame = +1
Query: 360 RYEARSSPDTYDMV-SGAVAYADAQLGQEEVMSMTPQQWYEAMTHMREQIAPILTRRRAQ 418
R E R+ P Y SG A Q ++E S P + H ++ + P +R+
Sbjct: 115 RDEGRAGPSLYRRQRSGKEEEAIWQRSRQEPHSRRP----DCREHCQDLLGPPRSRQDPH 282
Query: 419 RPRRRHHHQDQ 429
PRRRHH ++
Sbjct: 283 FPRRRHHSDER 315
>BF646615 homologue to PIR|T48395|T483 GT2-like protein - Arabidopsis
thaliana, partial (16%)
Length = 654
Score = 29.3 bits (64), Expect = 3.1
Identities = 16/41 (39%), Positives = 25/41 (60%), Gaps = 5/41 (12%)
Frame = -2
Query: 13 VACLMHLPIEG----RMLAHGKKMPKHEGA-ALLMTYLGVA 48
V+CL HLP+E + + K+P +EG AL++ +GVA
Sbjct: 365 VSCLGHLPVEASTAMKAVVSASKLPLNEGVEALVVVLVGVA 243
>TC83983 similar to GP|19386796|dbj|BAB86175. putative alpha-glucosidase 1
{Oryza sativa (japonica cultivar-group)}, partial (11%)
Length = 656
Score = 29.3 bits (64), Expect = 3.1
Identities = 13/60 (21%), Positives = 26/60 (42%), Gaps = 5/60 (8%)
Frame = -2
Query: 331 VSHPQILPPIPGDLPRPANEEQI-----IAEQWQRYEARSSPDTYDMVSGAVAYADAQLG 385
+SHP++ D+ RP+++ + I E W + R D + + A+ + G
Sbjct: 562 LSHPRVYKHASSDIYRPSHQSTLIQPSKIVESWHEFRDRDKDDMFPNTNSLCAHLETDRG 383
>TC81448 weakly similar to OMNI|NT01MC3624 toxin secretion ATP-binding
protein putative {Magnetococcus sp. MC-1}, partial (4%)
Length = 655
Score = 28.5 bits (62), Expect = 5.2
Identities = 20/87 (22%), Positives = 39/87 (43%), Gaps = 3/87 (3%)
Frame = +3
Query: 348 ANEEQIIAEQWQRYEARSSPDTYDMVSGAVAYADAQLGQEEVMSMTPQQWYEAMTHMREQ 407
A +E++ + + E + +V + A +L +E + + Q EAM +E+
Sbjct: 129 ATQEKLQHQLQEATEKLQHQEALQLVKHQLQEATEKLQHQEALQLVKHQHQEAM---QEK 299
Query: 408 IAPILTRRRAQRP---RRRHHHQDQDQ 431
+ L ++ + + RHHHQ Q Q
Sbjct: 300 LPQALHQQEVHQE*HHQVRHHHQQQHQ 380
>TC78527 similar to GP|14030876|gb|AAK52082.1 nuclease {Nicotiana tabacum},
partial (83%)
Length = 1346
Score = 27.7 bits (60), Expect = 8.9
Identities = 20/80 (25%), Positives = 33/80 (41%)
Frame = +2
Query: 265 ERVLRQYGYVQTIPRHPTDVRDLPPPSIVQMFIDFRTHTLKADARGVQAGEDTRRVADGY 324
E +L YG + P +V PPP+ + + + HTL D + V G+
Sbjct: 296 EGLLTFYGL--PLSHTPVEVPVQPPPTSLPHGVQYEIHTLPVDEKAVADGDT-------- 445
Query: 325 VLWYTRVSHPQILPPIPGDL 344
V Y + P+ +P +L
Sbjct: 446 VTVYVSTADPRESSRVPSNL 505
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.323 0.139 0.442
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,872,392
Number of Sequences: 36976
Number of extensions: 218585
Number of successful extensions: 1070
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 1056
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1065
length of query: 431
length of database: 9,014,727
effective HSP length: 99
effective length of query: 332
effective length of database: 5,354,103
effective search space: 1777562196
effective search space used: 1777562196
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 60 (27.7 bits)
Medicago: description of AC146789.1


