

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146784.10 + phase: 0
(168 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
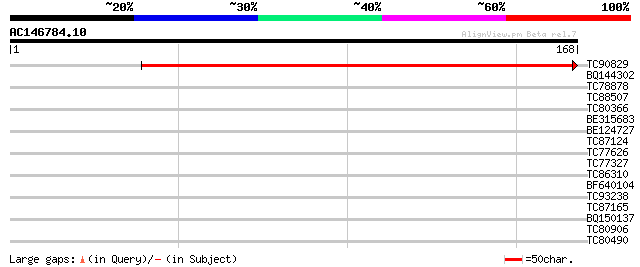
TC90829 weakly similar to PIR|T45789|T45789 hypothetical protein... 273 2e-74
BQ144302 similar to GP|6815109|dbj| hypothetical protein {Oryza ... 29 0.78
TC78878 similar to GP|4519792|dbj|BAA75744.1 Asp1 {Arabidopsis t... 29 1.0
TC88507 similar to GP|13122433|dbj|BAB32914. contains ESTs AU029... 29 1.0
TC80366 similar to GP|13877617|gb|AAK43886.1 protein kinase-like... 29 1.0
BE315683 similar to GP|9651943|gb receptor-like protein kinase 2... 28 2.3
BE124727 weakly similar to GP|9758777|db contains similarity to ... 27 3.0
TC87124 weakly similar to GP|21593647|gb|AAM65614.1 replication ... 27 3.0
TC77626 homologue to GP|5802244|gb|AAD51625.1| seed maturation p... 27 5.1
TC77327 homologue to GP|16224254|gb|AAL15651.1 dehydrin-like pro... 26 6.6
TC86310 similar to GP|16930469|gb|AAL31920.1 At2g26190/T1D16.17 ... 26 6.6
BF640104 26 8.6
TC93238 SP|P03756|VE22_LAMBD EA22 gene protein. {Bacteriophage l... 26 8.6
TC87165 weakly similar to GP|21689637|gb|AAM67440.1 putative sto... 26 8.6
BQ150137 similar to OMNI|NT01MC0313 transposase {Magnetococcus s... 26 8.6
TC80906 similar to GP|8886934|gb|AAF80620.1| F2D10.25 {Arabidops... 26 8.6
TC80490 similar to PIR|E84825|E84825 probable protein kinase [im... 26 8.6
>TC90829 weakly similar to PIR|T45789|T45789 hypothetical protein F26O13.220
- Arabidopsis thaliana, partial (12%)
Length = 676
Score = 273 bits (699), Expect = 2e-74
Identities = 129/129 (100%), Positives = 129/129 (100%)
Frame = +2
Query: 40 LHVTVVTPVPEASFFLRLPSFDKILTPVNGAYFLIFTVIVFAVTWACCCIFKKKPRDEIP 99
LHVTVVTPVPEASFFLRLPSFDKILTPVNGAYFLIFTVIVFAVTWACCCIFKKKPRDEIP
Sbjct: 2 LHVTVVTPVPEASFFLRLPSFDKILTPVNGAYFLIFTVIVFAVTWACCCIFKKKPRDEIP 181
Query: 100 YQELEMALPESASATVVESAEGWDQGWDDDWDDNVAVKSPVVRHAGSISANGLTSRSSNK 159
YQELEMALPESASATVVESAEGWDQGWDDDWDDNVAVKSPVVRHAGSISANGLTSRSSNK
Sbjct: 182 YQELEMALPESASATVVESAEGWDQGWDDDWDDNVAVKSPVVRHAGSISANGLTSRSSNK 361
Query: 160 DGWEDNWDD 168
DGWEDNWDD
Sbjct: 362 DGWEDNWDD 388
>BQ144302 similar to GP|6815109|dbj| hypothetical protein {Oryza sativa
(japonica cultivar-group)}, partial (3%)
Length = 1047
Score = 29.3 bits (64), Expect = 0.78
Identities = 13/43 (30%), Positives = 23/43 (53%)
Frame = +2
Query: 45 VTPVPEASFFLRLPSFDKILTPVNGAYFLIFTVIVFAVTWACC 87
++P+ + FFL +P F TP + +FL+ I F + + C
Sbjct: 593 ISPIHSSIFFLHIPLFPFFPTPPSYPFFLLTPFIPFLMILSFC 721
>TC78878 similar to GP|4519792|dbj|BAA75744.1 Asp1 {Arabidopsis thaliana},
partial (53%)
Length = 1839
Score = 28.9 bits (63), Expect = 1.0
Identities = 14/39 (35%), Positives = 18/39 (45%)
Frame = +2
Query: 130 WDDNVAVKSPVVRHAGSISANGLTSRSSNKDGWEDNWDD 168
W D VK AG + +SN GW+DNWD+
Sbjct: 449 WRDPPVVKENASTRAGK--GKPPLAAASNGGGWDDNWDN 559
>TC88507 similar to GP|13122433|dbj|BAB32914. contains ESTs AU029632(E31167)
AU162277(E31167)~similar to Arabidopsis thaliana
chromosome 3 T6K12., partial (69%)
Length = 1040
Score = 28.9 bits (63), Expect = 1.0
Identities = 19/70 (27%), Positives = 33/70 (47%), Gaps = 15/70 (21%)
Frame = +3
Query: 77 VIVFAVTWACCCIFKKKPRDEIPYQELEMALPESASATVV-------------ESAEGWD 123
V+ F +T+A CIF K PR+++ +++ L ++ T+V +A GW
Sbjct: 558 VLTFVITFAVLCIFLKGPRNDL----MKIWLLAMSTVTLVMVGSAYTGPSMNPANAFGWA 725
Query: 124 --QGWDDDWD 131
W + WD
Sbjct: 726 YLNNWHNTWD 755
>TC80366 similar to GP|13877617|gb|AAK43886.1 protein kinase-like protein
{Arabidopsis thaliana}, partial (16%)
Length = 814
Score = 28.9 bits (63), Expect = 1.0
Identities = 22/69 (31%), Positives = 28/69 (39%), Gaps = 6/69 (8%)
Frame = +3
Query: 77 VIVFAVTWACCCIFKKKPR---DEIPYQELEMALPESASATVVESAEGWDQGWDDDW--- 130
V+V CC F+KK R DE Y + P A VE G Q W ++
Sbjct: 363 VVVLVFFSICCICFRKKKRRRRDEEYYGQQNYQQPPPAQRPKVEPYGGPPQQWQNNAHPP 542
Query: 131 DDNVAVKSP 139
D+V K P
Sbjct: 543 SDHVVSKPP 569
>BE315683 similar to GP|9651943|gb receptor-like protein kinase 2 {Glycine
max}, partial (13%)
Length = 416
Score = 27.7 bits (60), Expect = 2.3
Identities = 12/24 (50%), Positives = 18/24 (75%)
Frame = -3
Query: 116 VESAEGWDQGWDDDWDDNVAVKSP 139
+E+A G + +DDD DD+V V+SP
Sbjct: 357 LEAAVGDSKAFDDDKDDSVIVESP 286
>BE124727 weakly similar to GP|9758777|db contains similarity to unknown
protein~dbj|BAA90625.1~gene_id:MNJ7.8 {Arabidopsis
thaliana}, partial (10%)
Length = 439
Score = 27.3 bits (59), Expect = 3.0
Identities = 12/50 (24%), Positives = 23/50 (46%)
Frame = +3
Query: 87 CCIFKKKPRDEIPYQELEMALPESASATVVESAEGWDQGWDDDWDDNVAV 136
C + +P D P +E ++ A +S + +DW++N+AV
Sbjct: 111 CLLIAGRPNDVFPTEETSISGHPGAVRMPQQSEQAGSNDMLEDWEENLAV 260
>TC87124 weakly similar to GP|21593647|gb|AAM65614.1 replication protein
A1-like {Arabidopsis thaliana}, partial (37%)
Length = 1053
Score = 27.3 bits (59), Expect = 3.0
Identities = 16/49 (32%), Positives = 25/49 (50%)
Frame = -3
Query: 52 SFFLRLPSFDKILTPVNGAYFLIFTVIVFAVTWACCCIFKKKPRDEIPY 100
+FF +LP ++I FL+ ++ + T CCC+F RD I Y
Sbjct: 184 AFFHKLPPIEQISPNP*MISFLLTSL--WRATNNCCCVFWASERDSIFY 44
>TC77626 homologue to GP|5802244|gb|AAD51625.1| seed maturation protein PM37
{Glycine max}, partial (98%)
Length = 1675
Score = 26.6 bits (57), Expect = 5.1
Identities = 15/43 (34%), Positives = 19/43 (43%)
Frame = -2
Query: 21 QIKSESTELTLDAGKGDCVLHVTVVTPVPEASFFLRLPSFDKI 63
Q+ E T G+G CVL+ + T E FL L D I
Sbjct: 1020 QMSQRELETTQSLGEGQCVLYKKIFTLSLELGVFLLLKYKDDI 892
>TC77327 homologue to GP|16224254|gb|AAL15651.1 dehydrin-like protein
{Medicago sativa}, partial (71%)
Length = 1330
Score = 26.2 bits (56), Expect = 6.6
Identities = 10/21 (47%), Positives = 14/21 (66%)
Frame = -2
Query: 66 PVNGAYFLIFTVIVFAVTWAC 86
PV G++ LIF++I F W C
Sbjct: 1005 PVPGSFSLIFSMIPFFSPWCC 943
>TC86310 similar to GP|16930469|gb|AAL31920.1 At2g26190/T1D16.17
{Arabidopsis thaliana}, partial (21%)
Length = 951
Score = 26.2 bits (56), Expect = 6.6
Identities = 11/33 (33%), Positives = 15/33 (45%)
Frame = +3
Query: 86 CCCIFKKKPRDEIPYQELEMALPESASATVVES 118
CCC + K E+PY + L +VES
Sbjct: 639 CCCNYSSKSLQELPY*KKPCRLCSGCGRAMVES 737
>BF640104
Length = 344
Score = 25.8 bits (55), Expect = 8.6
Identities = 11/27 (40%), Positives = 18/27 (65%)
Frame = +1
Query: 40 LHVTVVTPVPEASFFLRLPSFDKILTP 66
L + V+TP+P+ S +RL S ++TP
Sbjct: 241 LSLRVITPIPQTSSLVRLASQLILITP 321
>TC93238 SP|P03756|VE22_LAMBD EA22 gene protein. {Bacteriophage lambda},
partial (93%)
Length = 887
Score = 25.8 bits (55), Expect = 8.6
Identities = 24/89 (26%), Positives = 38/89 (41%), Gaps = 5/89 (5%)
Frame = -1
Query: 60 FDKILTPVNGAYFLIFTVIVFAVTWACCCIFKKKPRDEIPY--QELEMALPESASATVVE 117
F LTP+ F+I T++V + CC K + PY + + + L + +V
Sbjct: 266 FSNTLTPIRDNTFVILTLLV---EF*FCCF---KLNTQFPYC*RNILVLLVAAFDVLLVS 105
Query: 118 SA---EGWDQGWDDDWDDNVAVKSPVVRH 143
+ W + + V VKSPVV H
Sbjct: 104 FPFIQQFQHNRWCYQFMEKVCVKSPVVMH 18
>TC87165 weakly similar to GP|21689637|gb|AAM67440.1 putative storage
protein {Arabidopsis thaliana}, partial (71%)
Length = 1237
Score = 25.8 bits (55), Expect = 8.6
Identities = 13/26 (50%), Positives = 16/26 (61%), Gaps = 2/26 (7%)
Frame = -3
Query: 65 TPVNGAYFLIFTVIVF--AVTWACCC 88
TP+ GAYF IF VI F + + C C
Sbjct: 275 TPM*GAYFAIFCVIKFMLILNFLCYC 198
>BQ150137 similar to OMNI|NT01MC0313 transposase {Magnetococcus sp. MC-1},
partial (9%)
Length = 1056
Score = 25.8 bits (55), Expect = 8.6
Identities = 7/17 (41%), Positives = 12/17 (70%)
Frame = -1
Query: 84 WACCCIFKKKPRDEIPY 100
W+CCCI+++ D P+
Sbjct: 726 WSCCCIYRRAASD*HPW 676
>TC80906 similar to GP|8886934|gb|AAF80620.1| F2D10.25 {Arabidopsis
thaliana}, partial (19%)
Length = 1195
Score = 25.8 bits (55), Expect = 8.6
Identities = 11/25 (44%), Positives = 13/25 (52%), Gaps = 7/25 (28%)
Frame = +3
Query: 114 TVVESAEGWDQG-------WDDDWD 131
T+VE GW G WD+DWD
Sbjct: 897 TLVELPFGWQPGIQEGAADWDEDWD 971
>TC80490 similar to PIR|E84825|E84825 probable protein kinase [imported] -
Arabidopsis thaliana, partial (7%)
Length = 798
Score = 25.8 bits (55), Expect = 8.6
Identities = 13/25 (52%), Positives = 16/25 (64%)
Frame = -2
Query: 71 YFLIFTVIVFAVTWACCCIFKKKPR 95
+FL F VIVF+V C C+F PR
Sbjct: 71 FFLSFFVIVFSV---CDCVFCVNPR 6
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.319 0.135 0.427
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,714,612
Number of Sequences: 36976
Number of extensions: 73490
Number of successful extensions: 573
Number of sequences better than 10.0: 34
Number of HSP's better than 10.0 without gapping: 558
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 563
length of query: 168
length of database: 9,014,727
effective HSP length: 89
effective length of query: 79
effective length of database: 5,723,863
effective search space: 452185177
effective search space used: 452185177
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 55 (25.8 bits)
Medicago: description of AC146784.10


