

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146759.10 + phase: 1 /partial
(178 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
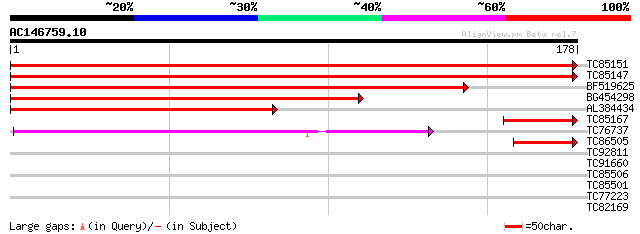
Score E
Sequences producing significant alignments: (bits) Value
TC85151 homologue to GP|9758155|dbj|BAB08712.1 40S ribosomal pro... 354 9e-99
TC85147 homologue to GP|9758155|dbj|BAB08712.1 40S ribosomal pro... 354 1e-98
BF519625 homologue to GP|18496083|em ribosomal protein S3 {Droso... 231 1e-61
BG454298 homologue to GP|14190419|gb At2g31610/T9H9.13 {Arabidop... 216 4e-57
AL384434 homologue to PIR|T39606|T39 40s ribosomal protein s3 - ... 155 8e-39
TC85167 homologue to PIR|S45788|S45788 probable membrane protein... 49 1e-06
TC76737 homologue to SP|P06379|RK2_TOBAC Chloroplast 50S ribosom... 44 3e-05
TC86505 similar to GP|15028127|gb|AAK76687.1 unknown protein {Ar... 43 6e-05
TC92811 homologue to SP|Q04716|RT03_PETHY Mitochondrial ribosoma... 30 0.66
TC91660 similar to GP|10177256|dbj|BAB10724. zinc-binding protei... 27 3.3
TC85506 similar to GP|7208616|gb|AAF40224.1| phenylalanine ammon... 27 5.6
TC85501 homologue to SP|P45732|PALY_STYHU Phenylalanine ammonia-... 27 5.6
TC77223 weakly similar to GP|19911209|dbj|BAB86931. glucosyltran... 26 7.3
TC82169 similar to GP|21703134|gb|AAM74507.1 AT4g27320/M4I22_130... 26 7.3
>TC85151 homologue to GP|9758155|dbj|BAB08712.1 40S ribosomal protein S3
{Arabidopsis thaliana}, partial (91%)
Length = 1066
Score = 354 bits (909), Expect = 9e-99
Identities = 178/178 (100%), Positives = 178/178 (100%)
Frame = +3
Query: 1 EKGRRIRELTSVVQKRFKFPENSVELYAEKVNNRGLCAIAQAESLRYKLLGGLAVRRACY 60
EKGRRIRELTSVVQKRFKFPENSVELYAEKVNNRGLCAIAQAESLRYKLLGGLAVRRACY
Sbjct: 270 EKGRRIRELTSVVQKRFKFPENSVELYAEKVNNRGLCAIAQAESLRYKLLGGLAVRRACY 449
Query: 61 GVLRFVMESGAKGCEVIVSGKLRAQRAKSMKFKDGYMISSGQPVKDYIDSAVRHVLLRQG 120
GVLRFVMESGAKGCEVIVSGKLRAQRAKSMKFKDGYMISSGQPVKDYIDSAVRHVLLRQG
Sbjct: 450 GVLRFVMESGAKGCEVIVSGKLRAQRAKSMKFKDGYMISSGQPVKDYIDSAVRHVLLRQG 629
Query: 121 VLGIKVKIMLDWDPKGKQGPKTPLPDIVTIHTPKEEEEYIRPAAVVANDIEVPVPVPV 178
VLGIKVKIMLDWDPKGKQGPKTPLPDIVTIHTPKEEEEYIRPAAVVANDIEVPVPVPV
Sbjct: 630 VLGIKVKIMLDWDPKGKQGPKTPLPDIVTIHTPKEEEEYIRPAAVVANDIEVPVPVPV 803
>TC85147 homologue to GP|9758155|dbj|BAB08712.1 40S ribosomal protein S3
{Arabidopsis thaliana}, partial (91%)
Length = 1136
Score = 354 bits (908), Expect = 1e-98
Identities = 177/178 (99%), Positives = 178/178 (99%)
Frame = +1
Query: 1 EKGRRIRELTSVVQKRFKFPENSVELYAEKVNNRGLCAIAQAESLRYKLLGGLAVRRACY 60
EKGRRIRELTSVVQKRFKFPENSVELYAEKVNNRGLCAIAQAESLRYKLLGGLAVRRACY
Sbjct: 421 EKGRRIRELTSVVQKRFKFPENSVELYAEKVNNRGLCAIAQAESLRYKLLGGLAVRRACY 600
Query: 61 GVLRFVMESGAKGCEVIVSGKLRAQRAKSMKFKDGYMISSGQPVKDYIDSAVRHVLLRQG 120
GVLRFVMESGAKGCEVIVSGKLRAQRAKSMKFKDGYMISSGQPVKDYIDSAVRHVLLRQG
Sbjct: 601 GVLRFVMESGAKGCEVIVSGKLRAQRAKSMKFKDGYMISSGQPVKDYIDSAVRHVLLRQG 780
Query: 121 VLGIKVKIMLDWDPKGKQGPKTPLPDIVTIHTPKEEEEYIRPAAVVANDIEVPVPVPV 178
VLGIKVKIMLDWDPKGKQGPKTPLPDIVTIHTPKEEEEYIRPAAVVANDIEVPVP+PV
Sbjct: 781 VLGIKVKIMLDWDPKGKQGPKTPLPDIVTIHTPKEEEEYIRPAAVVANDIEVPVPIPV 954
>BF519625 homologue to GP|18496083|em ribosomal protein S3 {Drosophila
virilis}, partial (83%)
Length = 626
Score = 231 bits (588), Expect = 1e-61
Identities = 115/144 (79%), Positives = 124/144 (85%)
Frame = +2
Query: 1 EKGRRIRELTSVVQKRFKFPENSVELYAEKVNNRGLCAIAQAESLRYKLLGGLAVRRACY 60
EKGRRIRELT++VQKRF F +ELYAEKV RGLCAIAQAESLRYKL GGLAVRRACY
Sbjct: 194 EKGRRIRELTAMVQKRFNFEPGRIELYAEKVATRGLCAIAQAESLRYKLTGGLAVRRACY 373
Query: 61 GVLRFVMESGAKGCEVIVSGKLRAQRAKSMKFKDGYMISSGQPVKDYIDSAVRHVLLRQG 120
GVLRF+MESGAKGCEV+VSGKLR QRAKSMKF DG MI SG P DY+++A RHVLLRQG
Sbjct: 374 GVLRFIMESGAKGCEVVVSGKLRGQRAKSMKFVDGLMIHSGDPCNDYVETATRHVLLRQG 553
Query: 121 VLGIKVKIMLDWDPKGKQGPKTPL 144
VLGIKVKIML +DPK K GPK PL
Sbjct: 554 VLGIKVKIMLPYDPKNKIGPKKPL 625
>BG454298 homologue to GP|14190419|gb At2g31610/T9H9.13 {Arabidopsis
thaliana}, partial (68%)
Length = 681
Score = 216 bits (550), Expect = 4e-57
Identities = 110/111 (99%), Positives = 110/111 (99%)
Frame = +1
Query: 1 EKGRRIRELTSVVQKRFKFPENSVELYAEKVNNRGLCAIAQAESLRYKLLGGLAVRRACY 60
EKGRRIRELTSVVQKRFKFPENSVELYAEKVNNRGLCAIAQAESLRYKLLGGLAVRRACY
Sbjct: 346 EKGRRIRELTSVVQKRFKFPENSVELYAEKVNNRGLCAIAQAESLRYKLLGGLAVRRACY 525
Query: 61 GVLRFVMESGAKGCEVIVSGKLRAQRAKSMKFKDGYMISSGQPVKDYIDSA 111
GVLRFVMESGAKGCEVIVSGKLRAQRAKSMKFKDG MISSGQPVKDYIDSA
Sbjct: 526 GVLRFVMESGAKGCEVIVSGKLRAQRAKSMKFKDGXMISSGQPVKDYIDSA 678
>AL384434 homologue to PIR|T39606|T39 40s ribosomal protein s3 - fission
yeast (Schizosaccharomyces pombe), partial (56%)
Length = 475
Score = 155 bits (392), Expect = 8e-39
Identities = 75/84 (89%), Positives = 81/84 (96%)
Frame = +1
Query: 1 EKGRRIRELTSVVQKRFKFPENSVELYAEKVNNRGLCAIAQAESLRYKLLGGLAVRRACY 60
EKGRRIRELT+++QKRFKFPEN+VELYAEKV NRGLCAIAQ ESLRYKLL GLAVRRACY
Sbjct: 223 EKGRRIRELTALIQKRFKFPENTVELYAEKVQNRGLCAIAQCESLRYKLLAGLAVRRACY 402
Query: 61 GVLRFVMESGAKGCEVIVSGKLRA 84
GVLRF+MESGAKGCEV+VSGKLRA
Sbjct: 403 GVLRFIMESGAKGCEVVVSGKLRA 474
>TC85167 homologue to PIR|S45788|S45788 probable membrane protein YBL053w -
yeast (Saccharomyces cerevisiae), partial (9%)
Length = 988
Score = 48.9 bits (115), Expect = 1e-06
Identities = 22/23 (95%), Positives = 23/23 (99%)
Frame = +2
Query: 156 EEEYIRPAAVVANDIEVPVPVPV 178
EEEYIRPAAVVANDIEVPVP+PV
Sbjct: 2 EEEYIRPAAVVANDIEVPVPIPV 70
>TC76737 homologue to SP|P06379|RK2_TOBAC Chloroplast 50S ribosomal protein
L2. [Common tobacco] {Nicotiana tabacum}, partial (95%)
Length = 5466
Score = 44.3 bits (103), Expect = 3e-05
Identities = 35/133 (26%), Positives = 65/133 (48%), Gaps = 1/133 (0%)
Frame = +3
Query: 2 KGRRIRELTSVVQKRFKFPENSVELYAEKVNNRGLCAIAQAESLRYKLLGGLAVRRACYG 61
K RRI EL + VQK+ + + + ++ N AE + +L ++ R+A
Sbjct: 2607 KPRRIEELQTNVQKKLNCVTRKINITSTRIPNAYSDPNILAEFIAGQLKNRISFRKAIKK 2786
Query: 62 VLRFVMESGAKGCEVIVSGKLRAQRAKSMKF-KDGYMISSGQPVKDYIDSAVRHVLLRQG 120
+ ++G KG +V ++G++ + +++ ++G + Q ++ ID V G
Sbjct: 2787 AIELAEQAGTKGVQVQIAGRIDGKEIARVEWIREGRV--PLQTIRAKIDYCSYPVRTIYG 2960
Query: 121 VLGIKVKIMLDWD 133
VLGIKV I L+ D
Sbjct: 2961 VLGIKVWIFLNND 2999
>TC86505 similar to GP|15028127|gb|AAK76687.1 unknown protein {Arabidopsis
thaliana}, partial (11%)
Length = 1100
Score = 43.1 bits (100), Expect = 6e-05
Identities = 19/20 (95%), Positives = 20/20 (100%)
Frame = -1
Query: 159 YIRPAAVVANDIEVPVPVPV 178
YIRPAAVVANDIEVPVP+PV
Sbjct: 257 YIRPAAVVANDIEVPVPIPV 198
>TC92811 homologue to SP|Q04716|RT03_PETHY Mitochondrial ribosomal protein
S3. [Petunia] {Petunia hybrida}, partial (34%)
Length = 668
Score = 29.6 bits (65), Expect = 0.66
Identities = 23/64 (35%), Positives = 33/64 (50%), Gaps = 1/64 (1%)
Frame = +3
Query: 66 VMESGAKGCEVIVSGKLR-AQRAKSMKFKDGYMISSGQPVKDYIDSAVRHVLLRQGVLGI 124
VM+ G +G + SG+L A+ A++ K G +S ID A V R G+LG+
Sbjct: 345 VMKKGVEGIRICCSGRLEGAEIARTECGKYGK--TSCNVFNQKIDYASAEVSTRYGILGV 518
Query: 125 KVKI 128
KV I
Sbjct: 519 KVWI 530
>TC91660 similar to GP|10177256|dbj|BAB10724. zinc-binding protein-like
{Arabidopsis thaliana}, complete
Length = 703
Score = 27.3 bits (59), Expect = 3.3
Identities = 24/77 (31%), Positives = 33/77 (42%), Gaps = 8/77 (10%)
Frame = +2
Query: 52 GLAVRRACYGVLRFVMESGAKGCEVIVSGKLRAQRAKSMKFKDG-YMISSGQPVKD---- 106
G V R L V + C IV K + KS K+K+G +++ G+ V D
Sbjct: 260 GTPVERMMLSGLHTVTDIFCCCCGQIVGWKYESAHEKSQKYKEGKFVLERGRIVDDVDSS 439
Query: 107 ---YIDSAVRHVLLRQG 120
YIDS HV + G
Sbjct: 440 TEFYIDS---HVSMSDG 481
>TC85506 similar to GP|7208616|gb|AAF40224.1| phenylalanine ammonia-lyase 2
{Rubus idaeus}, partial (15%)
Length = 1025
Score = 26.6 bits (57), Expect = 5.6
Identities = 12/37 (32%), Positives = 23/37 (61%)
Frame = +1
Query: 104 VKDYIDSAVRHVLLRQGVLGIKVKIMLDWDPKGKQGP 140
V ++ SAVRH+ L + + + + ++L W + +QGP
Sbjct: 121 VTRWLKSAVRHLPLLRWLELLPMIVVLGWSFRSRQGP 231
>TC85501 homologue to SP|P45732|PALY_STYHU Phenylalanine ammonia-lyase (EC
4.3.1.5). [Townsville stylo] {Stylosanthes humilis},
partial (96%)
Length = 2567
Score = 26.6 bits (57), Expect = 5.6
Identities = 12/37 (32%), Positives = 23/37 (61%)
Frame = +1
Query: 104 VKDYIDSAVRHVLLRQGVLGIKVKIMLDWDPKGKQGP 140
V ++ SAVRH+ L + + + + ++L W + +QGP
Sbjct: 274 VTRWLKSAVRHLPLLRWLELLPMIVVLGWSFRSRQGP 384
>TC77223 weakly similar to GP|19911209|dbj|BAB86931. glucosyltransferase-13
{Vigna angularis}, partial (50%)
Length = 1628
Score = 26.2 bits (56), Expect = 7.3
Identities = 12/20 (60%), Positives = 15/20 (75%)
Frame = +3
Query: 81 KLRAQRAKSMKFKDGYMISS 100
K+ QRAKSM F DG +I+S
Sbjct: 588 KMYIQRAKSMYFVDGILINS 647
>TC82169 similar to GP|21703134|gb|AAM74507.1 AT4g27320/M4I22_130
{Arabidopsis thaliana}, partial (50%)
Length = 600
Score = 26.2 bits (56), Expect = 7.3
Identities = 16/51 (31%), Positives = 25/51 (48%)
Frame = +1
Query: 128 IMLDWDPKGKQGPKTPLPDIVTIHTPKEEEEYIRPAAVVANDIEVPVPVPV 178
+ L PK + P TP + T ++PK E + P A ++P+P PV
Sbjct: 37 LFLSKKPKKCRTPITPHTN-PTHNSPKSESTTLPPLATSPPQHQLPLPAPV 186
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.319 0.138 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,313,323
Number of Sequences: 36976
Number of extensions: 48813
Number of successful extensions: 228
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 226
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 227
length of query: 178
length of database: 9,014,727
effective HSP length: 90
effective length of query: 88
effective length of database: 5,686,887
effective search space: 500446056
effective search space used: 500446056
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 55 (25.8 bits)
Medicago: description of AC146759.10


