

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
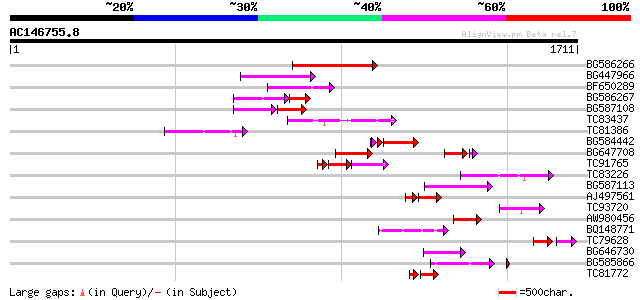
Query= AC146755.8 + phase: 0 /pseudo
(1711 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BG586266 similar to GP|7267666|em RNA-directed DNA polymerase-li... 232 1e-60
BG447966 weakly similar to PIR|G96509|G96 protein F27F5.21 [impo... 172 8e-43
BF650289 weakly similar to GP|9049283|dbj| orf129a {Beta vulgari... 157 3e-38
BG586267 weakly similar to PIR|A84888|A8 hypothetical protein At... 106 5e-34
BG587108 similar to PIR|G96509|G9 protein F27F5.21 [imported] - ... 86 8e-34
TC83437 weakly similar to PIR|D86384|D86384 unknown protein [imp... 117 4e-26
TC81386 similar to GP|10140689|gb|AAG13524.1 putative non-LTR re... 113 7e-25
BG584442 104 9e-24
BG647708 weakly similar to GP|13786450|gb| putative reverse tran... 107 3e-23
TC91765 weakly similar to GP|19881779|gb|AAM01180.1 Putative ret... 94 1e-21
TC83226 weakly similar to PIR|G86419|G86419 probable reverse tra... 95 3e-19
BG587113 weakly similar to PIR|A84888|A8 hypothetical protein At... 91 3e-18
AJ497561 weakly similar to GP|10140689|gb putative non-LTR retro... 72 9e-18
TC93720 similar to GP|21743320|dbj|BAC03314. putative nematode r... 90 9e-18
AW980456 88 3e-17
BQ148771 87 7e-17
TC79628 similar to GP|17104799|gb|AAL34288.1 putative 2-nitropro... 61 1e-16
BG646730 85 2e-16
BG585866 80 3e-15
TC81772 55 1e-11
>BG586266 similar to GP|7267666|em RNA-directed DNA polymerase-like protein
{Arabidopsis thaliana}, partial (18%)
Length = 789
Score = 232 bits (591), Expect = 1e-60
Identities = 111/257 (43%), Positives = 168/257 (65%)
Frame = -3
Query: 854 MKDWRPIALCNVVYKMVAKVLANRLKNVLDKCISTSQSAFVPGRSILDNALVAIELIHYM 913
+ ++R IA CN YK++AK+L+ R++ +L IS SQSAFVPGR+I DN L+ +++HY+
Sbjct: 778 VSEYRTIAPCNTQYKIIAKILSKRMQPLLRSIISPSQSAFVPGRAISDNVLITHKILHYL 599
Query: 914 KAKTKGSQGDVALKLDISKAYDRLDWEYLRDIMIQMRFSEKWIRWIMLCVETVDYTVLVN 973
+ +A+K D++KAYDR+ W +LR+++ ++ F WI WIM CV TV Y+ L+N
Sbjct: 598 RQSGAKKHVSMAVKTDMTKAYDRIAWNFLREVLTRLGFHGIWISWIMECVSTVSYSFLIN 419
Query: 974 GVQVGPLIPGRGIRQGDPLSPYLFILCAEGLSALIRDAERRGVITGTRICTSAPAISHLL 1033
G G ++P RG+RQGDPLSPYLFILC E LS L + A R+G + G ++ + P I+HLL
Sbjct: 418 GGPQGRVLPSRGLRQGDPLSPYLFILCTEVLSGLCQQALRKGTLPGVKVARNCPPINHLL 239
Query: 1034 FADDCFLFFRASEQEASVMKNILTTYEAASGQAINLQKSEMYCSRNTPTECQNRIATTLG 1093
FADD F +++ +++ +I+ Y AASG+ IN KS + S T +R+ L
Sbjct: 238 FADDTMFFGKSNASSCAILLSIMDKYRAASGRCIN*TKSAITFSSKTSQAIIDRVKGELK 59
Query: 1094 VKQVLGTGKYLGLPSMI 1110
+ + GTGKYLG +++
Sbjct: 58 IAKEGGTGKYLGYRNIL 8
>BG447966 weakly similar to PIR|G96509|G96 protein F27F5.21 [imported] -
Arabidopsis thaliana, partial (7%)
Length = 687
Score = 172 bits (437), Expect = 8e-43
Identities = 92/225 (40%), Positives = 130/225 (56%), Gaps = 1/225 (0%)
Frame = +2
Query: 698 WYKEGDLNTKFFHAAATSRRKVNRIEHLENSNGTVCRKEDELQNIARNYFSHLFTKLSST 757
W K+GD NTKFFH+ A+ RRKVN I+ L++ G C+ E+ ++ + YF++LFT + T
Sbjct: 5 WLKDGDKNTKFFHSKASQRRKVNEIKKLKDETGNWCKGEENVERLLITYFNNLFTSSNPT 184
Query: 758 RADVISKVATS-ISDDDNCFLTAAFTLEEFKTAAFSMQADKCPGPDGFNPGFYQNFWNMC 816
+ +V +S + + FT EE A M K PGPDG F+Q +W++
Sbjct: 185 AIEETCEVVKGKLSHEHIVWCEKEFTEEEVLEAINQMHPVKAPGPDGLPALFFQKYWHIV 364
Query: 817 GSEIHKAGCEWLENGVFPPHLNSTNIALIPKGDAQTTMKDWRPIALCNVVYKMVAKVLAN 876
G E+ + + L N + LN T I LIPKG T KD+RPI+LCNVV K++ KV+AN
Sbjct: 365 GKEVQQMVLQVLNNSMETEELNKTFIVLIPKGKNPNTPKDYRPISLCNVVMKIITKVIAN 544
Query: 877 RLKNVLDKCISTSQSAFVPGRSILDNALVAIELIHYMKAKTKGSQ 921
R+K L I QSAFV GR I DNAL+A + + + KG +
Sbjct: 545 RVKQTLPDVIDVEQSAFVQGRLITDNALIAWSVSIG*RXRRKGKR 679
>BF650289 weakly similar to GP|9049283|dbj| orf129a {Beta vulgaris}, partial
(69%)
Length = 616
Score = 157 bits (397), Expect = 3e-38
Identities = 75/203 (36%), Positives = 123/203 (59%)
Frame = +3
Query: 777 LTAAFTLEEFKTAAFSMQADKCPGPDGFNPGFYQNFWNMCGSEIHKAGCEWLENGVFPPH 836
L + FT E K A FSM + K PG DG+N F++ WN+ G + A ++ + G P
Sbjct: 12 LCSEFTAVEVKNALFSMDSSKAPGIDGYNVHFFKCSWNIIGDSVIDAILDFFKTGFMPKI 191
Query: 837 LNSTNIALIPKGDAQTTMKDWRPIALCNVVYKMVAKVLANRLKNVLDKCISTSQSAFVPG 896
+N T + L+PK T++K++RPIA C+V+YK+++K+L +R++ VL+ +S +QSAFV G
Sbjct: 192 INCTYVTLLPKEVNVTSVKNFRPIACCSVIYKIISKILTSRMQGVLNSVVSENQSAFVKG 371
Query: 897 RSILDNALVAIELIHYMKAKTKGSQGDVALKLDISKAYDRLDWEYLRDIMIQMRFSEKWI 956
R I DN +++ EL+ KG +K+D+ KAYD +W +++ +M+++ F K++
Sbjct: 372 RVIFDNIILSHELV--KSYSRKGISPRCMVKIDLXKAYDSXEWPFIKHLMLELGFPYKFV 545
Query: 957 RWIMLCVETVDYTVLVNGVQVGP 979
W+M + T YT NG P
Sbjct: 546 NWVMAXLTTASYTFNXNGDLTXP 614
>BG586267 weakly similar to PIR|A84888|A8 hypothetical protein At2g45230
[imported] - Arabidopsis thaliana, partial (16%)
Length = 794
Score = 106 bits (265), Expect(2) = 5e-34
Identities = 56/170 (32%), Positives = 93/170 (53%), Gaps = 2/170 (1%)
Frame = +3
Query: 675 LKNKLDKLLVQDDLFWKQRAKTFWYKEGDLNTKFFHAAATSRRKVNRIEHLENSNGTVCR 734
L+++L++ +++FW Q+++ W + GD NTKFFHA +RR NRI L + +
Sbjct: 90 LRSELNEEYHNEEIFWMQKSRLNWLRSGDRNTKFFHAVTKNRRAQNRILSLIDDDDKEWF 269
Query: 735 KEDELQNIARNYFSHLFTK--LSSTRADVISKVATSISDDDNCFLTAAFTLEEFKTAAFS 792
E++L +A ++F L++ + T D S + ++++ N L A + EE + A F
Sbjct: 270 VEEDLGRLADSHFKLLYSSEDVGITLEDWNS-IPAIVTEEQNAQLMAQISREEVREAVFD 446
Query: 793 MQADKCPGPDGFNPGFYQNFWNMCGSEIHKAGCEWLENGVFPPHLNSTNI 842
+ KCPGPDG N F+Q FW+ G ++ E+L G +N TNI
Sbjct: 447 INPHKCPGPDGMNVFFFQQFWDTMGDDLTSMAQEFLRTGKLEEGINKTNI 596
Score = 58.2 bits (139), Expect(2) = 5e-34
Identities = 30/63 (47%), Positives = 44/63 (69%)
Frame = +1
Query: 844 LIPKGDAQTTMKDWRPIALCNVVYKMVAKVLANRLKNVLDKCISTSQSAFVPGRSILDNA 903
L+PK + ++RPI+LCNV YK+V+KVL+ RLK+VL I+ +Q+AF + I DN
Sbjct: 601 LVPKKLEAKRLVEFRPISLCNVAYKIVSKVLSKRLKSVLPWIITETQAAFGRRQLISDNI 780
Query: 904 LVA 906
L+A
Sbjct: 781 LIA 789
>BG587108 similar to PIR|G96509|G9 protein F27F5.21 [imported] - Arabidopsis
thaliana, partial (17%)
Length = 677
Score = 85.9 bits (211), Expect(2) = 8e-34
Identities = 45/130 (34%), Positives = 71/130 (54%), Gaps = 1/130 (0%)
Frame = +2
Query: 675 LKNKLDKLLVQDDLFWKQRAKTFWYKEGDLNTKFFHAAATSRRKVNRIEHLENSNGTVCR 734
LK KL + ++ +W+Q+++ W+ GDLN KF+HA R NRI L + +G
Sbjct: 8 LKEKLQEAYKDEEDYWQQKSRNMWHISGDLNKKFYHALTKQRHARNRIVGLYDYDGNWIT 187
Query: 735 KEDELQNIARNYFSHLFTKLSSTRAD-VISKVATSISDDDNCFLTAAFTLEEFKTAAFSM 793
+E ++ +A +YF LF + + T D + ++ +SI+ N L T EE + A F M
Sbjct: 188 EEQGVEKVAVDYFEDLFQRTTPTGFDGFLDEITSSITPQMNQRLLRLATEEEVRLALFIM 367
Query: 794 QADKCPGPDG 803
+K PGPDG
Sbjct: 368 HPEKAPGPDG 397
Score = 78.2 bits (191), Expect(2) = 8e-34
Identities = 41/89 (46%), Positives = 56/89 (62%)
Frame = +3
Query: 808 FYQNFWNMCGSEIHKAGCEWLENGVFPPHLNSTNIALIPKGDAQTTMKDWRPIALCNVVY 867
F+Q+ W++ ++ K +L +G LN+TNI LIPK T M + RPI+LCNV Y
Sbjct: 411 FFQHSWHIIKMDLLKMVNSFLASGKLDTRLNTTNICLIPKKKRPTRMTELRPISLCNVGY 590
Query: 868 KMVAKVLANRLKNVLDKCISTSQSAFVPG 896
K+++KVL RLK L IS +QSAFV G
Sbjct: 591 KIISKVLCQRLKVCLPSLISETQSAFVHG 677
>TC83437 weakly similar to PIR|D86384|D86384 unknown protein [imported] -
Arabidopsis thaliana, partial (6%)
Length = 951
Score = 117 bits (293), Expect = 4e-26
Identities = 105/342 (30%), Positives = 160/342 (46%), Gaps = 12/342 (3%)
Frame = +2
Query: 837 LNSTNIALIPKGDAQTTMKDWRPIALCNVVYKMVAKVLANRLKNVLDKCISTSQSAFVPG 896
+NST IALIPK D + D+RPI+L +YK++ K+LANRL+ V+ IS +QSAFV
Sbjct: 50 INSTFIALIPKVDNPQRLNDFRPISLVGSLYKILGKLLANRLRVVIGSVISDAQSAFVKN 229
Query: 897 RSILDNALVAIELIHYMKAKTKGSQGDVALKLDISKAYDRLDWEYLRDIMIQ-------- 948
R IL+ + ++ +M+ + + K + L W R I +
Sbjct: 230 RQILEMVFL*-QMRLWMRLR------------N*RKIFCCLRWILKRLITLSIGLIWILF 370
Query: 949 ---MRFSEKWIRWIMLCVETVDYTVLVNGVQVGPLIPGRGIRQGDPLSPYLFILCAEGLS 1005
M F W +WI CV T +VLVNG L+ + + Q + Y F
Sbjct: 371 *VGMSFLVLWRKWIKECVSTATTSVLVNGSPTNVLM--KSLVQTQLFTRYSF-------- 520
Query: 1006 ALIRDAERRGVITGTRICTSAPAISHLLFADDCFLFFRASEQEASVMKNILTTYEAASGQ 1065
GV+ +SHL FA+D L + ++ L + A SG
Sbjct: 521 ---------GVVNPV-------VVSHLQFANDTLLLETKNWANIRALRAALVIF*AMSGL 652
Query: 1066 AINLQKSEMYCSRNTPTECQNRIATTLGVKQVLGTGKYLGLPSMIGRSKKATF-KFVKDR 1124
+N KS + C P+ + A+ L K YLG+P + G S++ +F + + +R
Sbjct: 653 KVNFHKSGLVCVNIAPSWL-SEAASVLSWKVGKVPFLYLGMP-IEGNSRRLSFWEPIVNR 826
Query: 1125 IWNRINSWSSRCLSQAGREILIKSVLQSIPSYVMSIFLLPST 1166
I R+ W+SR LS GR +L+KSVL S +S++ LPS+
Sbjct: 827 IKARLTGWNSRFLSFGGRLVLLKSVLTS-----LSVYALPSS 937
>TC81386 similar to GP|10140689|gb|AAG13524.1 putative non-LTR retroelement
reverse transcriptase {Oryza sativa (japonica
cultivar-group)}, partial (1%)
Length = 798
Score = 113 bits (282), Expect = 7e-25
Identities = 75/264 (28%), Positives = 121/264 (45%), Gaps = 12/264 (4%)
Frame = -1
Query: 466 RRKESWAFLKNLARTSSLPWCVIGDFNDILSSEEKKGRSERAPWLIRGFRQAVLDAGLFD 525
+R++ W+ + N+ L WC IGDFN IL S E +G A + F+Q LF
Sbjct: 798 KRRQLWSAISNIQTQHKLSWCCIGDFNTILGSHEHQGSHTPARLPMLDFQQWSDVNNLFH 619
Query: 526 LHMTGYAFTWFKSLGTARAVEEKLDRALVNSDWCNMFPNAGVECLSTTSSDHYPLWLTCK 585
L G AFTW ++LDR++VN + + L+ SDH+P+ +
Sbjct: 618 LPTRGSAFTWTNGRRGRNNTRKRLDRSIVNQLMIDNCESLSACTLTKLRSDHFPILFELQ 439
Query: 586 PVTAVNRNPHRFKFENAWLAEPDFKQQVQQRWKLY----PEEGITQKLSYCAEDLTDWSR 641
T + FKF W A PD ++Q W P + QKL E L W++
Sbjct: 438 --TQNIQFSSSFKFMKMWSAHPDCINIIKQCWANQVVGCPMFVLNQKLKNLKEVLKVWNK 265
Query: 642 NN-NNFRRDISKVQKKIEKLRTHVTAANVSYFNSLKNK-------LDKLLVQDDLFWKQR 693
N N + K+++ ++ + ++ Y + L ++ L+ L +++FW ++
Sbjct: 264 NTFGNVHSQVDNAYKELDDIQVKID--SIGYSDVLMDQEKAAQLNLESALNIEEVFWHEK 91
Query: 694 AKTFWYKEGDLNTKFFHAAATSRR 717
+K W+ EGD NT FFH A +R
Sbjct: 90 SKVNWHCEGDRNTAFFHRVAKIKR 19
>BG584442
Length = 775
Score = 104 bits (260), Expect(3) = 9e-24
Identities = 52/107 (48%), Positives = 71/107 (65%)
Frame = +1
Query: 1128 RINSWSSRCLSQAGREILIKSVLQSIPSYVMSIFLLPSTFIGEIEKMLNAFWWGHNSANS 1187
+IN ++CLS+ E++IK LQSI SYVMSIFLL ++ + EIEK++N F W H N
Sbjct: 382 KINF*RNKCLSKVM*EVMIKYALQSISSYVMSIFLLLNSQVDEIEKIMNTFSWVHVGENR 561
Query: 1188 RGMHWLSWERLSVPKTFGGMGFKSLRAFNLAMIGKQAWKLISNPDSL 1234
+GMHW+S E+L V K +GGMGF FN+ M+GKQ + N +L
Sbjct: 562 KGMHWMS*EKLFVHKNYGGMGFTDFTTFNIPMLGKQV*SFLLNRTTL 702
Score = 24.3 bits (51), Expect(3) = 9e-24
Identities = 9/17 (52%), Positives = 12/17 (69%)
Frame = +3
Query: 1109 MIGRSKKATFKFVKDRI 1125
M+ S KATF ++KD I
Sbjct: 324 MLSTSNKATFSYIKDEI 374
Score = 21.2 bits (43), Expect(3) = 9e-24
Identities = 10/19 (52%), Positives = 11/19 (57%)
Frame = +2
Query: 1088 IATTLGVKQVLGTGKYLGL 1106
I L V+ VLG KY GL
Sbjct: 239 ITYILDVRAVLGRDKYFGL 295
>BG647708 weakly similar to GP|13786450|gb| putative reverse transcriptase
{Oryza sativa}, partial (9%)
Length = 708
Score = 107 bits (268), Expect = 3e-23
Identities = 52/114 (45%), Positives = 76/114 (66%)
Frame = +1
Query: 982 PGRGIRQGDPLSPYLFILCAEGLSALIRDAERRGVITGTRICTSAPAISHLLFADDCFLF 1041
P +G+RQGDPLSPYLFILCA LS L++ + + G ++ S P I+HLLFADD LF
Sbjct: 4 PEKGLRQGDPLSPYLFILCANVLSGLLKREGNKQNLHGIQVARSDPKITHLLFADDSLLF 183
Query: 1042 FRASEQEASVMKNILTTYEAASGQAINLQKSEMYCSRNTPTECQNRIATTLGVK 1095
RA+ EA+ + +L +Y++ASGQ +N +KSE+ S+N P + + I + +K
Sbjct: 184 ARANLTEAATIMQVLHSYQSASGQLVNFEKSEVSYSQNVPNQEKEMICQQIAIK 345
Score = 59.3 bits (142), Expect(2) = 3e-11
Identities = 27/69 (39%), Positives = 43/69 (62%)
Frame = +3
Query: 1312 ALTVGHLFKPNTKEWNENFIRYVFNAETSNQILQTPLLQSVQVDKATWRFEKNGLYSVRS 1371
+L+V L +TK+WN + I + FN ++QI++ PL DK W +EK+G+YSVRS
Sbjct: 393 SLSVDELIDYDTKQWNRDLIFHSFNNYAAHQIIKIPLSMRQPEDKIIWHWEKDGIYSVRS 572
Query: 1372 AYREIINRN 1380
A+ + + N
Sbjct: 573 AHHALCDEN 599
Score = 28.9 bits (63), Expect(2) = 3e-11
Identities = 10/23 (43%), Positives = 13/23 (56%)
Frame = +2
Query: 1389 PGKWNIIWNLKLPPKIKNFLWRV 1411
P W IWN ++NFLWR+
Sbjct: 635 PVIWKAIWNTPASRSVQNFLWRL 703
>TC91765 weakly similar to GP|19881779|gb|AAM01180.1 Putative retroelement
{Oryza sativa (japonica cultivar-group)}, partial (1%)
Length = 625
Score = 94.0 bits (232), Expect = 5e-19
Identities = 52/110 (47%), Positives = 66/110 (59%)
Frame = +3
Query: 1032 LLFADDCFLFFRASEQEASVMKNILTTYEAASGQAINLQKSEMYCSRNTPTECQNRIATT 1091
LLF R E A +MKNIL YE SG+AI+L+KS +YCSRN P + I
Sbjct: 279 LLFEGAPLRSLRVEEHHAQIMKNILILYEEDSGKAISLRKS*IYCSRNVPDILKTSITYI 458
Query: 1092 LGVKQVLGTGKYLGLPSMIGRSKKATFKFVKDRIWNRINSWSSRCLSQAG 1141
LGV+ +LGT KYLGLPSMIGR + TF +K +W + N + R +S G
Sbjct: 459 LGVQFMLGTCKYLGLPSMIGRDRTTTFSSIKGGVWQK-NKFLERHMSIQG 605
Score = 80.9 bits (198), Expect(2) = 1e-21
Identities = 40/70 (57%), Positives = 49/70 (69%)
Frame = +2
Query: 961 LCVETVDYTVLVNGVQVGPLIPGRGIRQGDPLSPYLFILCAEGLSALIRDAERRGVITGT 1020
+CVE+ DY VLVN V P+IP RG++QGD LSPY+FI+C EGLS LI A+ RG GT
Sbjct: 101 MCVESNDYYVLVNNDAVDPIIPSRGLQQGDHLSPYIFIICVEGLSFLIPHAKERGDTHGT 280
Query: 1021 RICTSAPAIS 1030
I AP +S
Sbjct: 281 SI*RGAPPVS 310
Score = 42.4 bits (98), Expect(2) = 1e-21
Identities = 17/32 (53%), Positives = 26/32 (81%)
Frame = +1
Query: 928 LDISKAYDRLDWEYLRDIMIQMRFSEKWIRWI 959
L ISK Y+R+D +YL++IMI+M F+ +WI W+
Sbjct: 1 LYISKVYNRVD*DYLKEIMIKMGFNNRWIYWM 96
Score = 30.4 bits (67), Expect = 6.1
Identities = 13/29 (44%), Positives = 19/29 (64%)
Frame = +1
Query: 1118 FKFVKDRIWNRINSWSSRCLSQAGREILI 1146
F ++ +INSWS CLS+A RE++I
Sbjct: 538 FLLLRVECGKKINSWSDICLSKACREVMI 624
>TC83226 weakly similar to PIR|G86419|G86419 probable reverse transcriptase
100033-105622 [imported] - Arabidopsis thaliana, partial
(2%)
Length = 885
Score = 94.7 bits (234), Expect = 3e-19
Identities = 79/297 (26%), Positives = 128/297 (42%), Gaps = 15/297 (5%)
Frame = +3
Query: 1359 WRFEKNGLYSVRSAYREIINRNDVLLQHRVPGK-----WNIIWNLKLPPKIKNFLWRVCR 1413
W G+YSV+S Y + + + W IW+L P+ K LWR+
Sbjct: 6 WMHNPTGIYSVKSGYNTLRTWQTQQINNTSTSSDETLIWKKIWSLHTIPRHKVLLWRILN 185
Query: 1414 NCLPTRMRLISKGVQCPPTCAICNNHEEDGKHLFFACSKSVGCWQRLGFWPSIQQIWSHN 1473
+ LP R L +G+QC P C C++ E HLF +C S W + + + N
Sbjct: 186 DSLPVRSSLRKRGIQCYPLCPRCHSKTETITHLFMSCPLSKRVWFGSNLCINFDNLPNPN 365
Query: 1474 AFCADIIFSLLQQLDIAHQQIFAVTLWSLWRHRNNKVWNNIVETTEEIGDRAVAFLNSWK 1533
+ ++ + D A +++LW RN V + +I RA ++ +K
Sbjct: 366 FI--N*LYEAIL*KDECITI*IAAIIYNLWHARNLSVLEDQTILEMDIIQRASNCISDYK 539
Query: 1534 AAQETRIRSSPANPHFD---------ISKWSKPSVGRFKCNVDAAFSASLHRVGFGACIR 1584
A T+ S A +D +KW +P++G K N DA + + G G IR
Sbjct: 540 QA-NTQAPPSMARTGYDPRSQHRPAKNTKWKRPNLGLVKVNTDANLQ-NHGKWGLGIIIR 713
Query: 1585 DANGNHVISRTECFTPLLDVEM-GEAIGLLHAMRWAKDLNLVNMDFETDSKVVVENI 1640
D G V++ + T D + EA LL MR+AKD + FE D++ +++ +
Sbjct: 714 DEVG-LVMAASTWETDGNDRALEAEAYALLTGMRFAKDCGFXKVXFEGDNEKLMKMV 881
>BG587113 weakly similar to PIR|A84888|A8 hypothetical protein At2g45230
[imported] - Arabidopsis thaliana, partial (10%)
Length = 767
Score = 91.3 bits (225), Expect = 3e-18
Identities = 61/220 (27%), Positives = 95/220 (42%), Gaps = 13/220 (5%)
Frame = -3
Query: 1251 SASIGHNPSYVWRSLWSVRDYLKHGLKWSIGTGETISVWNQPWLKDPLCL--QPVTEVQI 1308
+A +G SY WRS+ S + +K G K IG GE ++W + C+ P
Sbjct: 759 NAPLGSWASYAWRSIHSAQHLIKQGAKVIIGNGENTNIWEREMAWKLTCVTNHPNKHSSR 580
Query: 1309 MWDALTVG-----HLFKPNTKEWNENFIRYVFNAETSNQILQTPLLQSVQVDKATWRFEK 1363
+ A T+ P +E N N I +F T +IL + D +W + K
Sbjct: 579 AY*APTLYGYEGCRSDDPMRRERNANLINSIFPEGTRRKILSIHPQGPIGEDSYSWEYSK 400
Query: 1364 NGLYSVRSAY---REII---NRNDVLLQHRVPGKWNIIWNLKLPPKIKNFLWRVCRNCLP 1417
+G YSV+S Y II N+ + Q + + +W PK+++FLWR N LP
Sbjct: 399 SGHYSVKSGYYVQTNIIAAANQRGTVDQPSLDDLYQRVWKYNTSPKVRHFLWRCISNSLP 220
Query: 1418 TRMRLISKGVQCPPTCAICNNHEEDGKHLFFACSKSVGCW 1457
T + S+ + +C+ C E H+ F C + W
Sbjct: 219 TAANMRSRHISKDGSCSRCGMESETVNHILFQCPYARLIW 100
>AJ497561 weakly similar to GP|10140689|gb putative non-LTR retroelement
reverse transcriptase {Oryza sativa (japonica
cultivar-group)}, partial (2%)
Length = 621
Score = 72.4 bits (176), Expect(2) = 9e-18
Identities = 32/70 (45%), Positives = 47/70 (66%)
Frame = +2
Query: 1233 SLITKLLKAKYFPHSDYFSASIGHNPSYVWRSLWSVRDYLKHGLKWSIGTGETISVWNQP 1292
SL++K+LK KYFP D+ A++GHNPS+ WRSL S + L G +W IG G I+V +
Sbjct: 410 SLLSKILKFKYFPQWDFSYANLGHNPSFTWRSLLSTQSLLTLGHRWMIGDGSQINVSSMS 589
Query: 1293 WLKDPLCLQP 1302
W+++ L+P
Sbjct: 590 WIRNRPTLRP 619
Score = 37.7 bits (86), Expect(2) = 9e-18
Identities = 16/37 (43%), Positives = 27/37 (72%)
Frame = +3
Query: 1195 WERLSVPKTFGGMGFKSLRAFNLAMIGKQAWKLISNP 1231
+ L+ K+ GG+G ++ + FN++M+GKQ WKL+S P
Sbjct: 303 FNNLARHKSVGGLGLQN-QEFNISMLGKQRWKLLSQP 410
>TC93720 similar to GP|21743320|dbj|BAC03314. putative nematode
resistance-like protein {Oryza sativa (japonica
cultivar-group)}, partial (3%)
Length = 457
Score = 89.7 bits (221), Expect = 9e-18
Identities = 46/141 (32%), Positives = 72/141 (50%), Gaps = 7/141 (4%)
Frame = -2
Query: 1479 IIFSLLQQLDIAHQQIFAVTLWSLWRHRNNKVWNNIVETTEEIGDRAVAFLNSWKAAQET 1538
+IF+LLQQ ++ A +WSLW+ K+W E+ ++ A LN W+AAQ
Sbjct: 441 LIFTLLQQFSSDQCELMATVMWSLWKSCRMKLWQQNNESNSQVVHHATHLLNDWRAAQTI 262
Query: 1539 RIRSS-------PANPHFDISKWSKPSVGRFKCNVDAAFSASLHRVGFGACIRDANGNHV 1591
+ P D W + +KCN+DA+F SL++VG G C+ D + V
Sbjct: 261 *SHNHDQSNMVYPCPMKQDEEAWMTLTPDMYKCNIDASFFTSLNKVGLGMCLADDADDFV 82
Query: 1592 ISRTECFTPLLDVEMGEAIGL 1612
+++T F P D++ GEA+ L
Sbjct: 81 LAKTIWFAPSCDIDAGEAVRL 19
>AW980456
Length = 779
Score = 87.8 bits (216), Expect = 3e-17
Identities = 40/84 (47%), Positives = 58/84 (68%), Gaps = 1/84 (1%)
Frame = -2
Query: 1340 SNQILQTPLLQSVQVDKATWRFEKNGLYSVRSAYREIINRN-DVLLQHRVPGKWNIIWNL 1398
+++I+ TPL+ +D+ W+ EK+G Y V+SAYR + D HR PG W+ IW L
Sbjct: 253 ADKIMSTPLISHA*LDRLIWKDEKHGKYYVKSAYRFCVEELFDSSYLHR-PGNWSGIWKL 77
Query: 1399 KLPPKIKNFLWRVCRNCLPTRMRL 1422
K+PPK++N +WR+CR CLPTR+RL
Sbjct: 76 KVPPKVQNLVWRMCRGCLPTRIRL 5
>BQ148771
Length = 680
Score = 86.7 bits (213), Expect = 7e-17
Identities = 66/215 (30%), Positives = 101/215 (46%), Gaps = 5/215 (2%)
Frame = -3
Query: 1113 SKKATFKFVKDRIWNRINSWSSRCLSQAGREILIKSVLQSIPSYVMSIFLLPSTFIGEIE 1172
+KK+ F V ++ + +W + LS A R L KSV++++P Y M ++P I EI+
Sbjct: 618 TKKSRFS-VYYQVHVMLANWKANHLSLARRVTLAKSVIEAVPLYPMMTTIIPKACIEEIQ 442
Query: 1173 KMLNAFWWGHNSANSRGMHWLSWERLSVPKTFGGMGFKSLRAFNLAMIGKQAWKLISNPD 1232
K+ F WG SR H + WE +S PKT G+G + L N A I K W + S +
Sbjct: 441 KLQRKFVWGDTEV-SRRYHAVGWETMSKPKTIYGLGLRRLDVMNKACIMKLGWSIYSGSN 265
Query: 1233 SLITKLLKAKYFPHSDYFSASIGHNP--SYVWRSLWSVRDYLKHGLKWSIGTGETISVWN 1290
SL T++++ KY S+ P S +W++L + ++ L S G WN
Sbjct: 264 SLCTEVMRGKY-QRSESLEEIFLEKPTDSSLWKALVKLWPEIERNLVDSNGN------WN 106
Query: 1291 QPWLKDPL---CLQPVTEVQIMWDALTVGHLFKPN 1322
LK L LQ + + + L L PN
Sbjct: 105 WEKLKQWLPFDVLQRIMAIATPHENLGSDKLIWPN 1
>TC79628 similar to GP|17104799|gb|AAL34288.1 putative 2-nitropropane
dioxygenase {Arabidopsis thaliana}, partial (97%)
Length = 1560
Score = 60.8 bits (146), Expect(2) = 1e-16
Identities = 27/57 (47%), Positives = 39/57 (68%)
Frame = +1
Query: 1582 CIRDANGNHVISRTECFTPLLDVEMGEAIGLLHAMRWAKDLNLVNMDFETDSKVVVE 1638
CI DA G +V+ RT F PL V++ +A+GL A++W DL + N+DF DSK+VV+
Sbjct: 1165 CI*DATGGYVLGRTTWFAPLCSVDIDKALGLRDALQWVADLGMDNIDFSLDSKLVVD 1335
Score = 45.4 bits (106), Expect(2) = 1e-16
Identities = 24/60 (40%), Positives = 35/60 (58%)
Frame = +3
Query: 1651 AIIHDCRHLLMTDLANSDVKFIRRQANSAAHNLAREALNHASFHYHLNIPHCIHTLINNE 1710
+II+ C L+ L S V+F RRQAN H LA+ AS +++P CI+ L++NE
Sbjct: 1377 SIINYCISYLVLILPISKVEFNRRQANGMTHGLAQTTTLEASSQLFIDLPPCINNLVSNE 1556
>BG646730
Length = 799
Score = 85.1 bits (209), Expect = 2e-16
Identities = 41/127 (32%), Positives = 69/127 (54%)
Frame = +1
Query: 1250 FSASIGHNPSYVWRSLWSVRDYLKHGLKWSIGTGETISVWNQPWLKDPLCLQPVTEVQIM 1309
F+ IG+ PSY WRS+++++D + G +WSI G+ + +W WL + + + +
Sbjct: 373 FALEIGNQPSYAWRSMFNIKDVIDLGSRWSISNGQNVRIWKDDWLPNQTGFKVCSPMVDF 552
Query: 1310 WDALTVGHLFKPNTKEWNENFIRYVFNAETSNQILQTPLLQSVQVDKATWRFEKNGLYSV 1369
+A + L NTK + +R VF+A + QIL PL + D+ +E++ YSV
Sbjct: 553 EEATCISELIDTNTKSSKRDLVRNVFSAFEAKQILNIPLSWRLWPDRRILYWERDRNYSV 732
Query: 1370 RSAYREI 1376
RSA+ I
Sbjct: 733 RSAHHLI 753
>BG585866
Length = 828
Score = 80.5 bits (197), Expect(2) = 3e-15
Identities = 55/195 (28%), Positives = 84/195 (42%), Gaps = 1/195 (0%)
Frame = +3
Query: 1269 RDYLKHGLKWSIGTGETISVWNQPWLKDPLCLQPVTEVQIMWDALTVGHLFKPNTKEWNE 1328
++ LK G W G+G + S W W L V I LTV +F T +
Sbjct: 96 KNVLKSGYTWRAGSGNS-SFWYTNWSSLGLLGTQAPFVDIHDLHLTVKDVF--TTGGQHT 266
Query: 1329 NFIRYVFNAETSNQILQTPLLQSVQV-DKATWRFEKNGLYSVRSAYREIINRNDVLLQHR 1387
+ + + + I T L + + D W NG+Y+ +S Y I+++ + + +
Sbjct: 267 QSLYTILPTDIAEVINNTHLNFNASIGDAYIWPHNSNGVYTAKSGYSWILSQTETVNYNN 446
Query: 1388 VPGKWNIIWNLKLPPKIKNFLWRVCRNCLPTRMRLISKGVQCPPTCAICNNHEEDGKHLF 1447
W+ IW LK+P K K FLW C N +PT L + + C+ C HEE H
Sbjct: 447 --SSWSWIWRLKIPEKYKFFLWLACHNAVPTLSLLNHRNMVNSAICSRCGEHEESFFHCV 620
Query: 1448 FACSKSVGCWQRLGF 1462
C S W ++GF
Sbjct: 621 RDCRFSKIIWHKIGF 665
Score = 21.2 bits (43), Expect(2) = 3e-15
Identities = 6/8 (75%), Positives = 7/8 (87%)
Frame = +2
Query: 1499 LWSLWRHR 1506
LW +WRHR
Sbjct: 755 LWWIWRHR 778
>TC81772
Length = 982
Score = 55.1 bits (131), Expect(2) = 1e-11
Identities = 23/56 (41%), Positives = 34/56 (60%)
Frame = -2
Query: 1239 LKAKYFPHSDYFSASIGHNPSYVWRSLWSVRDYLKHGLKWSIGTGETISVWNQPWL 1294
L++K F + IGHNPSY+W S+ ++R L+ G W IG G +I++W WL
Sbjct: 762 LRSKIFFTLELLGVVIGHNPSYIWCSV*TLRMVLEEGHHWKIGDGSSINIWTDHWL 595
Score = 38.9 bits (89), Expect = 0.017
Identities = 24/70 (34%), Positives = 34/70 (48%), Gaps = 1/70 (1%)
Frame = -1
Query: 1388 VPGKWNIIWNLKLPPKIKNFLWRVCRNCLPTRMRLISKG-VQCPPTCAICNNHEEDGKHL 1446
V W+I +LK K K F+WR NCLP L ++G V CP C + +L
Sbjct: 352 VQNTWSI*RSLKF--KEKFFMWRFGSNCLPN*ANLQTRGVVTCPTVRVCCE*AT*NS*YL 179
Query: 1447 FFACSKSVGC 1456
+ C++S C
Sbjct: 178 YLECTESAKC 149
Score = 33.9 bits (76), Expect(2) = 1e-11
Identities = 12/25 (48%), Positives = 18/25 (72%)
Frame = -1
Query: 1208 GFKSLRAFNLAMIGKQAWKLISNPD 1232
GF + +L M+GKQ WKL++NP+
Sbjct: 853 GFHDIYGLDLVMLGKQVWKLLTNPN 779
Score = 30.8 bits (68), Expect = 4.7
Identities = 36/182 (19%), Positives = 79/182 (42%), Gaps = 13/182 (7%)
Frame = -3
Query: 1220 IGKQAWKLISNPDSLITKLLKAKYFPHSDYFSASIGHNPSYVW-----RSLWSVRDYLKH 1274
+G+ + ++++ P S+++++ + KYF H ++ VW + ++ V L
Sbjct: 818 VGEASLEIVNQPKSIVSRVFEVKYFSHWNFL----------VWLLATIQVIFGVVSRLYG 669
Query: 1275 GLKWSIGTGETISVWNQ---PWLKDPLCLQPVTEVQIMWDALTVGH-LFKPNTKEWNENF 1330
TG+ + V P + +C V + I + +T L + K +
Sbjct: 668 WF*KKDITGK*VMVRQLIFGPIIGLVVCTSHVYKHLIFFHYITNS**LDR*EYKILDWYI 489
Query: 1331 IRYVFNAETSNQILQTPLLQSVQVDKATWRFEKNGLYSVRSAY----REIINRNDVLLQH 1386
+Y F + IL+ LL+ + D+ W F+ G +V++AY ++N ++ +Q
Sbjct: 488 SKYTFGGSDVSAILKITLLRLNKPDERIWIFDGRGTNAVKNAYIVCAEHLVNIKELKIQG 309
Query: 1387 RV 1388
+
Sbjct: 308 EI 303
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.333 0.142 0.482
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 64,401,683
Number of Sequences: 36976
Number of extensions: 1053706
Number of successful extensions: 8150
Number of sequences better than 10.0: 90
Number of HSP's better than 10.0 without gapping: 4471
Number of HSP's successfully gapped in prelim test: 391
Number of HSP's that attempted gapping in prelim test: 3435
Number of HSP's gapped (non-prelim): 5300
length of query: 1711
length of database: 9,014,727
effective HSP length: 110
effective length of query: 1601
effective length of database: 4,947,367
effective search space: 7920734567
effective search space used: 7920734567
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.5 bits)
S2: 65 (29.6 bits)
Medicago: description of AC146755.8


