

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146751.3 + phase: 1 /pseudo
(439 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
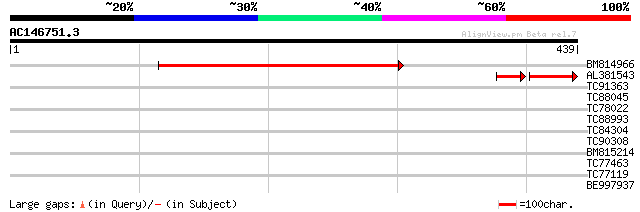
Sequences producing significant alignments: (bits) Value
BM814966 similar to GP|16648987|gb Unknown protein {Arabidopsis ... 397 e-111
AL381543 homologue to GP|16648987|gb| Unknown protein {Arabidops... 74 3e-23
TC91363 similar to PIR|T49093|T49093 hypothetical protein F4F15.... 32 0.63
TC88045 similar to GP|10177239|dbj|BAB10613. PGPD14 protein {Ara... 29 3.1
TC78022 similar to GP|20466746|gb|AAM20690.1 putative protein {A... 29 3.1
TC88993 similar to SP|Q943Z6|SR19_ARATH Signal recognition parti... 29 3.1
TC84304 similar to GP|4105798|gb|AAD02556.1| PGPD14 {Petunia x h... 29 4.1
TC90308 similar to SP|P50288|ASPG_LUPAL L-asparaginase (EC 3.5.1... 28 7.0
BM815214 similar to GP|22136992|gb unknown protein {Arabidopsis ... 28 7.0
TC77463 homologue to GP|10952510|gb|AAG24944.1 putative ammonium... 28 9.1
TC77119 similar to PIR|D96533|D96533 ARP protein [imported] - Ar... 28 9.1
BE997937 similar to GP|10764850|gb F1K23.5 {Arabidopsis thaliana... 28 9.1
>BM814966 similar to GP|16648987|gb Unknown protein {Arabidopsis thaliana},
partial (29%)
Length = 586
Score = 397 bits (1019), Expect = e-111
Identities = 189/190 (99%), Positives = 190/190 (99%)
Frame = +2
Query: 116 I*HD*NCDWD*AFCDINSC*LFNTFKSFGPTFTIHRKVFEFEKSGQEP*MESNL*GTRFT 175
+*HD*NCDWD*AFCDINSC*LFNTFKSFGPTFTIHRKVFEFEKSGQEP*MESNL*GTRFT
Sbjct: 17 L*HD*NCDWD*AFCDINSC*LFNTFKSFGPTFTIHRKVFEFEKSGQEP*MESNL*GTRFT 196
Query: 176 FPRACDHA*RSG*EGKRVCF*RHRCSSTYFR**YNCAWKQYKIYVIHC*FHSLWCCRSFQ 235
FPRACDHA*RSG*EGKRVCF*RHRCSSTYFR**YNCAWKQYKIYVIHC*FHSLWCCRSFQ
Sbjct: 197 FPRACDHA*RSG*EGKRVCF*RHRCSSTYFR**YNCAWKQYKIYVIHC*FHSLWCCRSFQ 376
Query: 236 FCVSNNGVYLGFVYSHYIRVWWSN*TSDAYASNFNLH*S*MC*CFG*SDFRCSFGYR*DC 295
FCVSNNGVYLGFVYSHYIRVWWSN*TSDAYASNFNLH*S*MC*CFG*SDFRCSFGYR*DC
Sbjct: 377 FCVSNNGVYLGFVYSHYIRVWWSN*TSDAYASNFNLH*S*MC*CFG*SDFRCSFGYR*DC 556
Query: 296 IFSRMSYMAF 305
IFSRMSYMAF
Sbjct: 557 IFSRMSYMAF 586
>AL381543 homologue to GP|16648987|gb| Unknown protein {Arabidopsis
thaliana}, partial (7%)
Length = 521
Score = 73.6 bits (179), Expect(2) = 3e-23
Identities = 36/37 (97%), Positives = 37/37 (99%)
Frame = +2
Query: 403 LQGAIMGPLITTVMIALKDLYAEFVLEEPKDRAKKKT 439
L+GAIMGPLITTVMIALKDLYAEFVLEEPKDRAKKKT
Sbjct: 59 LEGAIMGPLITTVMIALKDLYAEFVLEEPKDRAKKKT 169
Score = 52.8 bits (125), Expect(2) = 3e-23
Identities = 22/22 (100%), Positives = 22/22 (100%)
Frame = +1
Query: 378 *CLSYWTQHHWGNDIVPISS*G 399
*CLSYWTQHHWGNDIVPISS*G
Sbjct: 1 *CLSYWTQHHWGNDIVPISS*G 66
>TC91363 similar to PIR|T49093|T49093 hypothetical protein F4F15.250 -
Arabidopsis thaliana, partial (8%)
Length = 724
Score = 31.6 bits (70), Expect = 0.63
Identities = 15/30 (50%), Positives = 16/30 (53%)
Frame = +3
Query: 361 FPYGLWCVRDSGRCSWK*CLSYWTQHHWGN 390
F YG + S CSW *C YW Q WGN
Sbjct: 300 FTYGWPDINRSTPCSWN*CSLYW-QGGWGN 386
>TC88045 similar to GP|10177239|dbj|BAB10613. PGPD14 protein {Arabidopsis
thaliana}, partial (76%)
Length = 1225
Score = 29.3 bits (64), Expect = 3.1
Identities = 13/35 (37%), Positives = 18/35 (51%)
Frame = +2
Query: 12 QEKCN*CFFYKKGGEEVEYYCCNWAYCCYDCRIFD 46
Q CN C + GG+E ++ CN CCY + D
Sbjct: 404 QYHCNGCGICRTGGQE-NFFHCNKCGCCYSTLLRD 505
>TC78022 similar to GP|20466746|gb|AAM20690.1 putative protein {Arabidopsis
thaliana}, partial (31%)
Length = 1701
Score = 29.3 bits (64), Expect = 3.1
Identities = 10/26 (38%), Positives = 13/26 (49%)
Frame = +3
Query: 363 YGLWCVRDSGRCSWK*CLSYWTQHHW 388
Y +WC D G WK S W ++ W
Sbjct: 837 YWIWC*HDQGGRKWKIRAS*WNEYRW 914
>TC88993 similar to SP|Q943Z6|SR19_ARATH Signal recognition particle 19 kDa
protein (SRP19). [Mouse-ear cress] {Arabidopsis
thaliana}, partial (87%)
Length = 640
Score = 29.3 bits (64), Expect = 3.1
Identities = 16/44 (36%), Positives = 22/44 (49%), Gaps = 2/44 (4%)
Frame = +1
Query: 213 WKQYKIYVI--HC*FHSLWCCRSFQFCVSNNGVYLGFVYSHYIR 254
WK+ KI + C* S WC + FC + VY F+ H I+
Sbjct: 433 WKKEKIIMCLKTC*TFSYWCLSKYGFC*ED--VYYSFILVHIIQ 558
>TC84304 similar to GP|4105798|gb|AAD02556.1| PGPD14 {Petunia x hybrida},
partial (75%)
Length = 676
Score = 28.9 bits (63), Expect = 4.1
Identities = 15/44 (34%), Positives = 23/44 (52%), Gaps = 2/44 (4%)
Frame = +2
Query: 5 FFEDQ*FQEK--CN*CFFYKKGGEEVEYYCCNWAYCCYDCRIFD 46
FF+D + + C+ C + GGEE ++ CN CCY + D
Sbjct: 293 FFDDDVSKNQYHCDECGICRTGGEE-NFFHCNKCECCYSTLMKD 421
>TC90308 similar to SP|P50288|ASPG_LUPAL L-asparaginase (EC 3.5.1.1)
(L-asparagine amidohydrolase). [White lupine] {Lupinus
albus}, partial (71%)
Length = 939
Score = 28.1 bits (61), Expect = 7.0
Identities = 12/30 (40%), Positives = 16/30 (53%)
Frame = +1
Query: 16 N*CFFYKKGGEEVEYYCCNWAYCCYDCRIF 45
N F+Y G + E +CC W+Y C C F
Sbjct: 4 NGSFYY--GWKHYEMWCCFWSYYCR*CCFF 87
>BM815214 similar to GP|22136992|gb unknown protein {Arabidopsis thaliana},
partial (24%)
Length = 596
Score = 28.1 bits (61), Expect = 7.0
Identities = 8/15 (53%), Positives = 10/15 (66%)
Frame = -3
Query: 31 YCCNWAYCCYDCRIF 45
+CCN + CY C IF
Sbjct: 147 FCCNMNFLCYSCNIF 103
>TC77463 homologue to GP|10952510|gb|AAG24944.1 putative ammonium transporter
AMT1;1 {Lotus japonicus}, complete
Length = 1952
Score = 27.7 bits (60), Expect = 9.1
Identities = 9/14 (64%), Positives = 9/14 (64%)
Frame = -3
Query: 24 GGEEVEYYCCNWAY 37
GG E YYCC W Y
Sbjct: 1110 GGYEATYYCCPWFY 1069
>TC77119 similar to PIR|D96533|D96533 ARP protein [imported] - Arabidopsis
thaliana, partial (93%)
Length = 2277
Score = 27.7 bits (60), Expect = 9.1
Identities = 10/17 (58%), Positives = 11/17 (63%), Gaps = 3/17 (17%)
Frame = +3
Query: 31 YCC---NWAYCCYDCRI 44
YCC NWA CC C+I
Sbjct: 1404 YCCGRRNWAICCSACKI 1454
>BE997937 similar to GP|10764850|gb F1K23.5 {Arabidopsis thaliana}, partial
(52%)
Length = 538
Score = 27.7 bits (60), Expect = 9.1
Identities = 11/30 (36%), Positives = 15/30 (49%)
Frame = +1
Query: 18 CFFYKKGGEEVEYYCCNWAYCCYDCRIFDW 47
CF KGG+ + + C YCCY R +
Sbjct: 253 CFERSKGGDWIRPFSC---YCCYSSRTLSY 333
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.369 0.166 0.734
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,105,598
Number of Sequences: 36976
Number of extensions: 345929
Number of successful extensions: 5813
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 2419
Number of HSP's successfully gapped in prelim test: 294
Number of HSP's that attempted gapping in prelim test: 3163
Number of HSP's gapped (non-prelim): 3053
length of query: 439
length of database: 9,014,727
effective HSP length: 99
effective length of query: 340
effective length of database: 5,354,103
effective search space: 1820395020
effective search space used: 1820395020
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.8 bits)
S2: 60 (27.7 bits)
Medicago: description of AC146751.3


