

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146751.11 - phase: 0
(399 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
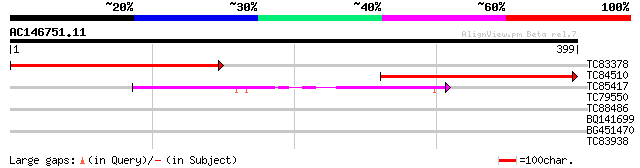
TC83378 weakly similar to PIR|D96631|D96631 RNA polymerase subun... 298 2e-81
TC84510 similar to PIR|D96631|D96631 RNA polymerase subunit (iso... 282 2e-76
TC85417 similar to SP|Q39211|RP3A_ARATH DNA-directed RNA polymer... 113 1e-25
TC79550 similar to GP|21593370|gb|AAM65319.1 DNA-directed RNA po... 39 0.004
TC88486 similar to GP|19386796|dbj|BAB86175. putative alpha-gluc... 31 0.96
BQ141699 similar to GP|334068|gb|A ORF2 {Pseudorabies virus}, pa... 30 1.3
BG451470 similar to GP|2191177|gb|A belongs to the SPOU family o... 29 3.7
TC83938 28 8.2
>TC83378 weakly similar to PIR|D96631|D96631 RNA polymerase subunit (isoform
B) [imported] - Arabidopsis thaliana, partial (27%)
Length = 452
Score = 298 bits (763), Expect = 2e-81
Identities = 150/150 (100%), Positives = 150/150 (100%)
Frame = +1
Query: 1 MSSSGDESMASEIDEQEQNGGRGIPDYIMNLEDVPSTLPTHLELQKTRVFCNLDAPQHTD 60
MSSSGDESMASEIDEQEQNGGRGIPDYIMNLEDVPSTLPTHLELQKTRVFCNLDAPQHTD
Sbjct: 1 MSSSGDESMASEIDEQEQNGGRGIPDYIMNLEDVPSTLPTHLELQKTRVFCNLDAPQHTD 180
Query: 61 TIQYSGAYAALGVDNSVRLDNFYQNFKVEVKRITDEEMEFDMIGIDPAVANAFRRILIAE 120
TIQYSGAYAALGVDNSVRLDNFYQNFKVEVKRITDEEMEFDMIGIDPAVANAFRRILIAE
Sbjct: 181 TIQYSGAYAALGVDNSVRLDNFYQNFKVEVKRITDEEMEFDMIGIDPAVANAFRRILIAE 360
Query: 121 VPTMAIERVYIANNTSLVQDEVLSHRLGLI 150
VPTMAIERVYIANNTSLVQDEVLSHRLGLI
Sbjct: 361 VPTMAIERVYIANNTSLVQDEVLSHRLGLI 450
>TC84510 similar to PIR|D96631|D96631 RNA polymerase subunit (isoform B)
[imported] - Arabidopsis thaliana, partial (32%)
Length = 509
Score = 282 bits (721), Expect = 2e-76
Identities = 138/138 (100%), Positives = 138/138 (100%)
Frame = +3
Query: 262 VKGIGKTHAKWSPVATAWYRMLPEVVLTKDVKDELAEELVSKCPAKVFDIEDIGRGRKKA 321
VKGIGKTHAKWSPVATAWYRMLPEVVLTKDVKDELAEELVSKCPAKVFDIEDIGRGRKKA
Sbjct: 3 VKGIGKTHAKWSPVATAWYRMLPEVVLTKDVKDELAEELVSKCPAKVFDIEDIGRGRKKA 182
Query: 322 VVKNARACTLCRECIREADEGEEGGEGEKWTDRVSLRRVKDHFIFTIESTGAIPPDVLFT 381
VVKNARACTLCRECIREADEGEEGGEGEKWTDRVSLRRVKDHFIFTIESTGAIPPDVLFT
Sbjct: 183 VVKNARACTLCRECIREADEGEEGGEGEKWTDRVSLRRVKDHFIFTIESTGAIPPDVLFT 362
Query: 382 EAVKILEDKCERVITELS 399
EAVKILEDKCERVITELS
Sbjct: 363 EAVKILEDKCERVITELS 416
>TC85417 similar to SP|Q39211|RP3A_ARATH DNA-directed RNA polymerase II 36
kDa polypeptide A (EC 2.7.7.6) (RNA polymerase II
subunit 3)., partial (64%)
Length = 855
Score = 113 bits (283), Expect = 1e-25
Identities = 78/233 (33%), Positives = 122/233 (51%), Gaps = 9/233 (3%)
Frame = +1
Query: 87 KVEVKRITDEEMEFDMIGIDPAVANAFRRILIAEVPTMAIERVYIANNTSLVQDEVLSHR 146
+V+++ + D+ M+F++ D +VANA RR++I+EVPT+AI+ V I N+S++ DE ++HR
Sbjct: 118 RVKIRELKDDYMKFELRDTDASVANALRRVMISEVPTVAIDLVEIEVNSSVLNDEFIAHR 297
Query: 147 LGLIPIDADPKL---FEYPDDA--GGENNEKNTIVFKQHVRCEKGQPRLTVKSDTLKWLP 201
LGLIP+ ++ + F DA G E ++ F V+C Q L V S L
Sbjct: 298 LGLIPLTSERAMAMRFSRDCDACDGDGQCEYCSVEFHLRVKCITDQ-TLDVSSKDL---- 462
Query: 202 NGSELIAEGTKSAADPAPKTFTTFNQKSLPKFSKDPAPYNLDVIVAKLGPGQEIELEAHA 261
+ ++ T + D +P +I+ KL GQE+ L A A
Sbjct: 463 ----ISSDHTVTPVD-------------VPGGDASIESDGRGIIIVKLRRGQELRLRAIA 591
Query: 262 VKGIGKTHAKWSPVATAWYRMLPEVVLTKDVKDELA----EELVSKCPAKVFD 310
KGIGK HAKWSP AT + P++ + +D+ + L +E V P +VFD
Sbjct: 592 RKGIGKDHAKWSPAATVTFMYEPDIHINEDLMESLTLEEKKEWVDSSPTRVFD 750
>TC79550 similar to GP|21593370|gb|AAM65319.1 DNA-directed RNA polymerase II
third largest subunit {Arabidopsis thaliana}, partial
(31%)
Length = 612
Score = 38.9 bits (89), Expect = 0.004
Identities = 25/88 (28%), Positives = 47/88 (53%)
Frame = +2
Query: 307 KVFDIEDIGRGRKKAVVKNARACTLCRECIREADEGEEGGEGEKWTDRVSLRRVKDHFIF 366
+VFDI+ + ++ +V + A + E I++A+ + G + + +D FIF
Sbjct: 11 RVFDIDPV---TQQVMVVDPEAYSYDDEVIKKAEAMGKPG-------LIEITAKQDSFIF 160
Query: 367 TIESTGAIPPDVLFTEAVKILEDKCERV 394
T+ESTGA+ L +++IL+ K + V
Sbjct: 161 TVESTGAVKASQLLLNSIEILKQKLDAV 244
>TC88486 similar to GP|19386796|dbj|BAB86175. putative alpha-glucosidase 1
{Oryza sativa (japonica cultivar-group)}, partial (13%)
Length = 1233
Score = 30.8 bits (68), Expect = 0.96
Identities = 15/46 (32%), Positives = 25/46 (53%), Gaps = 2/46 (4%)
Frame = -1
Query: 307 KVFDIEDIGRGRKKAVVKNARACTLCRECIREA--DEGEEGGEGEK 350
K+ + D+ RG+++ + R C E + E D+ EE G+GEK
Sbjct: 330 KLRKVHDLWRGKRRNDARKMRLCCRSNEVMNEKKDDDDEEEGDGEK 193
>BQ141699 similar to GP|334068|gb|A ORF2 {Pseudorabies virus}, partial (1%)
Length = 1217
Score = 30.4 bits (67), Expect = 1.3
Identities = 12/30 (40%), Positives = 18/30 (60%)
Frame = +3
Query: 340 DEGEEGGEGEKWTDRVSLRRVKDHFIFTIE 369
+EG GGEGE +RRV+ H++ +E
Sbjct: 483 EEGGTGGEGEACASAGFVRRVRPHYVLNVE 572
>BG451470 similar to GP|2191177|gb|A belongs to the SPOU family of rRNA
methylases {Arabidopsis thaliana}, partial (19%)
Length = 656
Score = 28.9 bits (63), Expect = 3.7
Identities = 11/19 (57%), Positives = 14/19 (72%)
Frame = -3
Query: 337 READEGEEGGEGEKWTDRV 355
RE +G EGGEG KW ++V
Sbjct: 228 REKRKGGEGGEGRKWREKV 172
>TC83938
Length = 737
Score = 27.7 bits (60), Expect = 8.2
Identities = 23/102 (22%), Positives = 38/102 (36%), Gaps = 15/102 (14%)
Frame = +3
Query: 284 PEVVLTKDVKDELAEELVSKCPAKVF-----------DIEDIGRGRKKAVVKNARACTLC 332
P + +TK VK + A+VF DIED + +V ++ CT+
Sbjct: 324 PPIAITKPVKHRQETSKTNSNNAQVFLNDDAIMKVSSDIEDQFLKDDEILVSKSKLCTIV 503
Query: 333 RECIREADEGEEGGEGE----KWTDRVSLRRVKDHFIFTIES 370
+C + +G+ E W LR + IE+
Sbjct: 504 SQCPSKPAQGDPSQRDED*SYSWVVTKELRNLVSENYLAIEN 629
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.315 0.134 0.389
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,685,521
Number of Sequences: 36976
Number of extensions: 137664
Number of successful extensions: 645
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 642
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 644
length of query: 399
length of database: 9,014,727
effective HSP length: 98
effective length of query: 301
effective length of database: 5,391,079
effective search space: 1622714779
effective search space used: 1622714779
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.5 bits)
S2: 59 (27.3 bits)
Medicago: description of AC146751.11


