

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146750.9 + phase: 0
(214 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
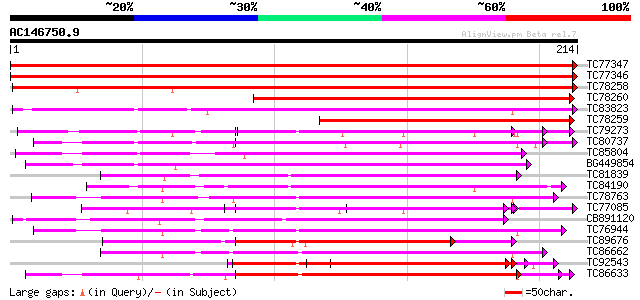
Score E
Sequences producing significant alignments: (bits) Value
TC77347 weakly similar to GP|22324851|gb|AAM95647.1 polygalactur... 444 e-125
TC77346 weakly similar to GP|18542897|gb|AAL75739.1 Putative rec... 405 e-114
TC78258 similar to GP|21593869|gb|AAM65836.1 polygalacturonase i... 366 e-102
TC78260 weakly similar to PIR|T00712|T00712 protein kinase homol... 201 1e-52
TC83823 weakly similar to GP|21593869|gb|AAM65836.1 polygalactur... 168 2e-42
TC78259 weakly similar to PIR|T00712|T00712 protein kinase homol... 156 5e-39
TC79273 weakly similar to PIR|S47965|S47965 polygalacturonase in... 133 4e-32
TC80737 weakly similar to GP|1143381|emb|CAA88846.1 polygalactur... 124 2e-29
TC85804 weakly similar to GP|8778050|gb|AAF79181.1| polygalactur... 122 1e-28
BG449854 weakly similar to GP|5579315|gb| polygalacturonase inhi... 108 2e-24
TC81839 similar to GP|21593619|gb|AAM65586.1 receptor protein ki... 102 8e-23
TC84190 weakly similar to GP|4008006|gb|AAC95351.1| receptor-lik... 102 8e-23
TC78763 somatic embryogenesis receptor kinase 1 [Medicago trunca... 100 7e-22
TC77085 similar to GP|21536600|gb|AAM60932.1 putative disease re... 99 1e-21
CB891120 similar to GP|17979065|gb putative receptor kinase homo... 96 8e-21
TC76944 similar to GP|13899097|gb|AAK48970.1 Unknown protein {Ar... 94 3e-20
TC89676 similar to PIR|D84889|D84889 probable receptor-like prot... 94 4e-20
TC86662 weakly similar to GP|4756646|emb|CAB42250.1 unnamed prot... 94 5e-20
TC92543 weakly similar to GP|6041804|gb|AAF02124.1| putative pro... 93 7e-20
TC86633 weakly similar to SP|Q00874|D100_ARATH DNA-damage-repair... 91 4e-19
>TC77347 weakly similar to GP|22324851|gb|AAM95647.1 polygalacturonase
inhibitory protein {Brassica napus}, partial (21%)
Length = 926
Score = 444 bits (1142), Expect = e-125
Identities = 214/214 (100%), Positives = 214/214 (100%)
Frame = +3
Query: 1 MFLLLHLTLLFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGRLDDWDNNTECC 60
MFLLLHLTLLFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGRLDDWDNNTECC
Sbjct: 120 MFLLLHLTLLFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGRLDDWDNNTECC 299
Query: 61 DWSFVGCGRPYPGRVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFS 120
DWSFVGCGRPYPGRVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFS
Sbjct: 300 DWSFVGCGRPYPGRVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFS 479
Query: 121 KLQRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSF 180
KLQRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSF
Sbjct: 480 KLQRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSF 659
Query: 181 NQLCGAIPAGLKKFNKNVFEHNKCLCGAPLAPCK 214
NQLCGAIPAGLKKFNKNVFEHNKCLCGAPLAPCK
Sbjct: 660 NQLCGAIPAGLKKFNKNVFEHNKCLCGAPLAPCK 761
>TC77346 weakly similar to GP|18542897|gb|AAL75739.1 Putative receptor-like
protein kinase {Oryza sativa}, partial (6%)
Length = 865
Score = 405 bits (1041), Expect = e-114
Identities = 194/214 (90%), Positives = 201/214 (93%)
Frame = +2
Query: 1 MFLLLHLTLLFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGRLDDWDNNTECC 60
MF+LL LTL FLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGP GRLDDWDNNTECC
Sbjct: 56 MFMLLSLTLFFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPNGRLDDWDNNTECC 235
Query: 61 DWSFVGCGRPYPGRVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFS 120
DW+FVGCGRPYPGRVT VTI RGWGLSGTLPAEFG+LPYLSMLSLAEMPKVTGPIPNSFS
Sbjct: 236 DWNFVGCGRPYPGRVTGVTIGRGWGLSGTLPAEFGDLPYLSMLSLAEMPKVTGPIPNSFS 415
Query: 121 KLQRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSF 180
KL+RL+NLDLGSNSLSGPIP FLGKLKRL+EVDLSNNKLSG IPASL NLQSLSQFNVSF
Sbjct: 416 KLKRLENLDLGSNSLSGPIPEFLGKLKRLQEVDLSNNKLSGAIPASLDNLQSLSQFNVSF 595
Query: 181 NQLCGAIPAGLKKFNKNVFEHNKCLCGAPLAPCK 214
NQLCGAIP GL KF K+ FEHNKCLC APLAPCK
Sbjct: 596 NQLCGAIPTGLSKFAKSSFEHNKCLCDAPLAPCK 697
>TC78258 similar to GP|21593869|gb|AAM65836.1 polygalacturonase inhibiting
protein 1 {Arabidopsis thaliana}, partial (18%)
Length = 885
Score = 366 bits (940), Expect = e-102
Identities = 177/215 (82%), Positives = 193/215 (89%), Gaps = 2/215 (0%)
Frame = +2
Query: 2 FLLLHLTLLFLSSIYFSKSNASD-DFFCNADDKAALLKIRDHFGGPKGRLDDWDNNTECC 60
FLLLHLTLLFLSSIYFS SNA D +F CNA+DKA LLKIRDHFGGP GRL DWDN T+CC
Sbjct: 71 FLLLHLTLLFLSSIYFSTSNADDVEFSCNANDKATLLKIRDHFGGPNGRLSDWDNGTDCC 250
Query: 61 -DWSFVGCGRPYPGRVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSF 119
DWSFVGCG+P PGR+TVVTISRGWGLSGTLPAEFG+LPYL+ LSLAEMPKVTGPIPNSF
Sbjct: 251 SDWSFVGCGKPLPGRITVVTISRGWGLSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSF 430
Query: 120 SKLQRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVS 179
SKL+RLQ LDLGSNSLSGPIP+FLG+LK L+E DLSNNKL+G IPAS G++ SLSQFNVS
Sbjct: 431 SKLKRLQKLDLGSNSLSGPIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSIPSLSQFNVS 610
Query: 180 FNQLCGAIPAGLKKFNKNVFEHNKCLCGAPLAPCK 214
FNQLCGAIP GL KF K+ F+HNKCLCGAPLA CK
Sbjct: 611 FNQLCGAIPTGLNKFAKSSFDHNKCLCGAPLAACK 715
>TC78260 weakly similar to PIR|T00712|T00712 protein kinase homolog F22O13.7
- Arabidopsis thaliana, partial (7%)
Length = 563
Score = 201 bits (512), Expect = 1e-52
Identities = 98/121 (80%), Positives = 107/121 (87%)
Frame = +3
Query: 93 EFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKLKRLKEV 152
EFG+LPYL+ LSLAEMPKVTGPIPNSFSKLQRLQ LDLGSNSLSG IP+FLG+LK L+E
Sbjct: 3 EFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQRLQKLDLGSNSLSGSIPTFLGQLKGLQEF 182
Query: 153 DLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKKFNKNVFEHNKCLCGAPLAP 212
DLSNNKL+G IPAS G+L SLSQFNVSFNQLCGAIP GL KF K+ F+HNKCLCGAPL
Sbjct: 183 DLSNNKLTGVIPASFGSLPSLSQFNVSFNQLCGAIPTGLSKFAKSSFDHNKCLCGAPLTA 362
Query: 213 C 213
C
Sbjct: 363 C 365
Score = 31.6 bits (70), Expect = 0.24
Identities = 15/39 (38%), Positives = 24/39 (61%)
Frame = +3
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQR 124
L+G +PA FG+LP LS +++ ++ G IP SK +
Sbjct: 201 LTGVIPASFGSLPSLSQFNVS-FNQLCGAIPTGLSKFAK 314
>TC83823 weakly similar to GP|21593869|gb|AAM65836.1 polygalacturonase
inhibiting protein 1 {Arabidopsis thaliana}, partial
(38%)
Length = 798
Score = 168 bits (425), Expect = 2e-42
Identities = 99/220 (45%), Positives = 124/220 (56%), Gaps = 7/220 (3%)
Frame = +1
Query: 2 FLLLHLTLLFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGRLDDWDNNTECCD 61
FLLL LLFL S+ FS S C+ DD+ LLKI+DHF W T+CC
Sbjct: 52 FLLL---LLFLPSLTFSLSPPPRRHHCHPDDEKVLLKIKDHFHNTS-LFSTWTPKTDCCK 219
Query: 62 WSFVGCGRPYPG----RVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPN 117
W V C + P RV + I L GT+P +LPYL L +P +TGPIP
Sbjct: 220 WRIVLC-KKIPKTTIYRVNFLEIDGADDLVGTIPPLIADLPYLETLIFRNLPNLTGPIPQ 396
Query: 118 SFSKLQRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFN 177
+ +KL L + L NSL+GPIP + KL L + L+NN L+G +PA LG L L +
Sbjct: 397 AIAKLPHLNFVLLNWNSLTGPIPDYFSKLTNLATLGLNNNHLTGRVPAYLGRLPKLVGLD 576
Query: 178 VSFNQLCGAIP---AGLKKFNKNVFEHNKCLCGAPLAPCK 214
VS NQLCG IP L+ F+ + FE+NKCLCGAPLAPCK
Sbjct: 577 VSHNQLCGPIPKDGGKLQGFDPSWFENNKCLCGAPLAPCK 696
>TC78259 weakly similar to PIR|T00712|T00712 protein kinase homolog F22O13.7
- Arabidopsis thaliana, partial (6%)
Length = 522
Score = 156 bits (395), Expect = 5e-39
Identities = 76/96 (79%), Positives = 84/96 (87%)
Frame = +2
Query: 118 SFSKLQRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFN 177
+FSKLQRLQ LDLGSNSLSG IP+FLG+LK L+E DLSNNKL+G IPAS G+L SLSQFN
Sbjct: 32 AFSKLQRLQKLDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFN 211
Query: 178 VSFNQLCGAIPAGLKKFNKNVFEHNKCLCGAPLAPC 213
VSFNQLCGAIP GL KF K+ F+HNKCLCGAPLA C
Sbjct: 212 VSFNQLCGAIPTGLSKFAKSSFDHNKCLCGAPLAAC 319
Score = 31.6 bits (70), Expect = 0.24
Identities = 15/39 (38%), Positives = 24/39 (61%)
Frame = +2
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQR 124
L+G +PA FG+LP LS +++ ++ G IP SK +
Sbjct: 155 LTGVIPASFGSLPSLSQFNVS-FNQLCGAIPTGLSKFAK 268
>TC79273 weakly similar to PIR|S47965|S47965 polygalacturonase inhibitor
protein - tomato, partial (37%)
Length = 1399
Score = 133 bits (335), Expect = 4e-32
Identities = 78/189 (41%), Positives = 105/189 (55%), Gaps = 1/189 (0%)
Frame = +2
Query: 4 LLHLTLLFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGRLDDWDNNTECC-DW 62
L+ +LF+SS+ S S CN +DK LL + F WD T+CC +W
Sbjct: 29 LIFTFILFISSLNLSFSTT----ICNTNDKNVLLGXKSQFNNASV-FTTWDPITDCCKNW 193
Query: 63 SFVGCGRPYPGRVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKL 122
S + C GRVT++ +S + G +P NLP+L + A P V+G IP + +KL
Sbjct: 194 SGIECNSN--GRVTMLAVSDTNDVIGEIPTSVVNLPFLQFFTFAVFPGVSGTIPPAIAKL 367
Query: 123 QRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQ 182
L +LD +SL+GPIP FLG+LK L +DLS N+ +G IPASLG L L N+ NQ
Sbjct: 368 TNLVHLDFSLDSLTGPIPDFLGQLKNLDVIDLSGNRFTGQIPASLGRLTKLRSANLGSNQ 547
Query: 183 LCGAIPAGL 191
L G IPA L
Sbjct: 548 LSGPIPASL 574
Score = 97.4 bits (241), Expect = 4e-21
Identities = 66/178 (37%), Positives = 85/178 (47%), Gaps = 50/178 (28%)
Frame = +2
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQR-LQNLDLGSNSLSGPIPSFLG 144
LSG +PA LP L+ LSL + ++TG IP SF + N+DL SN+LSGPIPS G
Sbjct: 620 LSGPIPASLAQLPKLNELSLFQN-QLTGSIPESFGSFKNPALNIDLSSNNLSGPIPSSFG 796
Query: 145 KLK-----------------------------------------------RLKEVDLSNN 157
K K LK +D+S+N
Sbjct: 797 KAKITALVLSKNKFSGDASFLFGKDKTVLTTMDLSNNIFKFDFSNVDLSPGLKNLDISHN 976
Query: 158 KLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAG--LKKFNKNVFEHNKCLCGAPLAPC 213
+ G++P LG L L + +VSFNQLCG IP G LK+F+ F +N CLCG PL C
Sbjct: 977 MIFGSLPEKLGQLP-LQRLDVSFNQLCGQIPTGRRLKQFSPTKFSNNTCLCGLPLPAC 1147
Score = 72.0 bits (175), Expect = 2e-13
Identities = 44/120 (36%), Positives = 69/120 (56%), Gaps = 3/120 (2%)
Frame = +2
Query: 87 SGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKL 146
+G +PA G L L +L +++GPIP S ++ L+ L + N+LSGPIP+ L +L
Sbjct: 479 TGQIPASLGRLTKLRSANLGSN-QLSGPIPASLGMIKSLEQLYIYINNLSGPIPASLAQL 655
Query: 147 KRLKEVDLSNNKLSGTIPASLGNLQSLS-QFNVSFNQLCGAIPA--GLKKFNKNVFEHNK 203
+L E+ L N+L+G+IP S G+ ++ + ++S N L G IP+ G K V NK
Sbjct: 656 PKLNELSLFQNQLTGSIPESFGSFKNPALNIDLSSNNLSGPIPSSFGKAKITALVLSKNK 835
>TC80737 weakly similar to GP|1143381|emb|CAA88846.1 polygalacturonase
inhibitor {Actinidia deliciosa}, partial (77%)
Length = 1176
Score = 124 bits (312), Expect = 2e-29
Identities = 76/201 (37%), Positives = 106/201 (51%), Gaps = 7/201 (3%)
Frame = +2
Query: 10 LFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGRLDDWDNNTECCDWSFVGCGR 69
L LS I FS+ CN DK ALL+I+ P L WD T+CC W V C
Sbjct: 50 LLLSPIAFSEK-------CNPQDKRALLRIKKELNNPY-LLASWDPQTDCCGWYCVKCDL 205
Query: 70 PYPGRVTVVTISRG---WGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQ 126
R+T + + LSGT+P G+LPYL L ++P++ GPI + +KL +L+
Sbjct: 206 -ITHRITALIMQSSVPDTNLSGTIPPSVGDLPYLENLEFHKLPRLKGPIQPTIAKLTKLK 382
Query: 127 NLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGA 186
L + ++SGPIP FL +LK L+ + LS N LSG IP+SL L +L ++ N+L G
Sbjct: 383 YLFIEYTNVSGPIPPFLAQLKNLQLLHLSTNNLSGPIPSSLSQLPNLESLHLDRNKLTGP 562
Query: 187 IPAGLKKFNKN----VFEHNK 203
IP F K + HN+
Sbjct: 563 IPESFGSFKKPGPDIILSHNQ 625
Score = 83.6 bits (205), Expect = 5e-17
Identities = 58/182 (31%), Positives = 84/182 (45%), Gaps = 53/182 (29%)
Frame = +2
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRL-QNLDLGSNSLSGPIPSFLG 144
LSG +P+ LP L L L + K+TGPIP SF ++ ++ L N LSGPIP+ LG
Sbjct: 479 LSGPIPSSLSQLPNLESLHL-DRNKLTGPIPESFGSFKKPGPDIILSHNQLSGPIPASLG 655
Query: 145 KL-----------------------------------------------KRLKEVDLSNN 157
++ + L +D+++N
Sbjct: 656 QIDPERIDLSRNKLEGDASVLFGSQKRTQILDVSRNLLSFDFSKVDFPKQSLIWLDINHN 835
Query: 158 KLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAG-----LKKFNKNVFEHNKCLCGAPLAP 212
K+ G IP +L +++L QFNVS+N+L G IP G +F+ + HNK LCG PL
Sbjct: 836 KIYGKIPVTLTKVENLQQFNVSYNKLSGEIPQGGTGRLQDRFDVYAYLHNKGLCGPPLPK 1015
Query: 213 CK 214
CK
Sbjct: 1016CK 1021
>TC85804 weakly similar to GP|8778050|gb|AAF79181.1| polygalacturonase
inhibiting protein {Prunus mahaleb}, partial (51%)
Length = 1094
Score = 122 bits (306), Expect = 1e-28
Identities = 77/212 (36%), Positives = 107/212 (50%), Gaps = 19/212 (8%)
Frame = +1
Query: 3 LLLHLTLLFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGRLDDWDNNTECCDW 62
LL +LL L + S+ CN DK ALL+I+ P L W+ CCDW
Sbjct: 28 LLCFFSLLLLFPVAISQK-------CNPQDKKALLQIKKELNNPTS-LSSWNPRKNCCDW 183
Query: 63 SFVGCGRPYPGRVTVVTISRGWGLS-------------------GTLPAEFGNLPYLSML 103
F+ C VT SR L+ G + G+L Y+ L
Sbjct: 184 VFIHCD---------VTTSRVIWLAIQFSSPDQFTTPFPNPEFIGHISPSVGDLSYVERL 336
Query: 104 SLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTI 163
++P VTG IP++ SKL+ L+ L + S+SGPIPSFLG+ K L+ +DL +NKL+G+I
Sbjct: 337 EFNQLPNVTGQIPSTISKLKNLKYLTISGTSVSGPIPSFLGQFKNLELLDLYSNKLTGSI 516
Query: 164 PASLGNLQSLSQFNVSFNQLCGAIPAGLKKFN 195
P+SL L +L Q + N+L G IPA L + N
Sbjct: 517 PSSLSQLTNLKQLFLHENKLSGHIPASLGQLN 612
>BG449854 weakly similar to GP|5579315|gb| polygalacturonase inhibitor
protein {Glycine max}, partial (46%)
Length = 635
Score = 108 bits (269), Expect = 2e-24
Identities = 68/193 (35%), Positives = 100/193 (51%), Gaps = 2/193 (1%)
Frame = +3
Query: 7 LTLLFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGRLDDWDNNTECCD--WSF 64
L LL L++ F S + CN DK L +I+ FG P +L WD+ T+CC+ W
Sbjct: 27 LLLLILTTHLFIPSLSEQ---CNPHDKKTLFQIKKQFGNPT-KLSSWDSTTDCCNGTWYG 194
Query: 65 VGCGRPYPGRVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQR 124
V C + T+ +P NLP+L L+L P + G IP S S L +
Sbjct: 195 VFCDKQTYRVSTLDLQDLDLPHPVPIPPSIFNLPFLLYLTLMNTPNLVGTIPPSISNLTK 374
Query: 125 LQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLC 184
L ++ + ++SG IP+ L ++K L +D+S+NKL+G +PASL L +L+ NQL
Sbjct: 375 LNSIYIIQTNISGEIPNTLSEIKTLVTIDISDNKLTGPLPASLSTLPNLAGIFFYRNQLT 554
Query: 185 GAIPAGLKKFNKN 197
GAIP F K+
Sbjct: 555 GAIPKSYGSFPKS 593
>TC81839 similar to GP|21593619|gb|AAM65586.1 receptor protein kinase-like
protein {Arabidopsis thaliana}, partial (34%)
Length = 960
Score = 102 bits (255), Expect = 8e-23
Identities = 61/160 (38%), Positives = 84/160 (52%), Gaps = 1/160 (0%)
Frame = +2
Query: 35 ALLKIRDHFGGPKGRLDDWDNNT-ECCDWSFVGCGRPYPGRVTVVTISRGWGLSGTLPAE 93
AL+ I++ P G ++WD + + C W+ V C P + V LSGTL +
Sbjct: 239 ALVSIKESLMDPHGIFENWDGDAVDPCSWNMVTCS---PENLVVSLGIPSQNLSGTLSSS 409
Query: 94 FGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKLKRLKEVD 153
GNL L + L + +TGPIP+ KL LQ LDL N G IP LG L+ L+ +
Sbjct: 410 IGNLTNLQTVVL-QNNNITGPIPSELGKLSMLQTLDLSDNLFHGKIPPSLGHLRNLQYLR 586
Query: 154 LSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKK 193
L+NN SG P SL N+ L+ ++SFN L G +P L K
Sbjct: 587 LNNNSFSGECPESLANMAQLAFLDLSFNNLTGNVPRILAK 706
Score = 32.3 bits (72), Expect = 0.14
Identities = 15/42 (35%), Positives = 25/42 (58%)
Frame = +2
Query: 154 LSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKKFN 195
+ + LSGT+ +S+GNL +L + N + G IP+ L K +
Sbjct: 371 IPSQNLSGTLSSSIGNLTNLQTVVLQNNNITGPIPSELGKLS 496
>TC84190 weakly similar to GP|4008006|gb|AAC95351.1| receptor-like protein
kinase {Arabidopsis thaliana}, partial (21%)
Length = 833
Score = 102 bits (255), Expect = 8e-23
Identities = 79/204 (38%), Positives = 97/204 (46%), Gaps = 23/204 (11%)
Frame = +3
Query: 30 ADDKAALLKIRDHFGGPKGRLDDWDNN-TECCDWSFVGCGRPYPGRVTVVTISRGWGLSG 88
A D+A+LL +R GG R W++ T C W+ V C RVT + + GLSG
Sbjct: 162 ASDRASLLTLRATVGG---RTLLWNSTETNPCLWTGVICNNK---RVTALRLP-AMGLSG 320
Query: 89 TLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKLKR 148
LP+ GNL L LSL +TGPIP F+KL L+NL L SN SG +P FL L+
Sbjct: 321 NLPSGIGNLTELQTLSL-RYNALTGPIPMDFAKLVSLRNLYLHSNFFSGEVPEFLYGLQN 497
Query: 149 LKEVDLSNNKLSGTIPASLGNLQSLS----------------------QFNVSFNQLCGA 186
L ++L N SG I NL L QFNVSFN L G
Sbjct: 498 LVRLNLGKNNFSGEISQHFNNLTRLDTLFLEQNMFTGSVPDLNIPPLHQFNVSFNNLTGQ 677
Query: 187 IPAGLKKFNKNVFEHNKCLCGAPL 210
IP + N + F N LCG PL
Sbjct: 678 IPKRFSRLNISAFSGNS-LCGNPL 746
>TC78763 somatic embryogenesis receptor kinase 1 [Medicago truncatula]
Length = 2737
Score = 99.8 bits (247), Expect = 7e-22
Identities = 71/203 (34%), Positives = 94/203 (45%), Gaps = 4/203 (1%)
Frame = +3
Query: 9 LLFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGRLDDWDNN-TECCDWSFVGC 67
LL L ++ +N D AL +R + P L WD C W V C
Sbjct: 471 LLLLHPLWLVSANMEGD---------ALHNLRTNLQDPNNVLQSWDPTLVNPCTWFHVTC 623
Query: 68 GRPYPGRVTVVTISRG-WGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQ 126
+V+ + G LSGTL + G L L L L +TGPIP+ L L
Sbjct: 624 NNDN----SVIRVDLGNAALSGTLVPQLGQLKNLQYLELYSN-NITGPIPSDLGNLTNLV 788
Query: 127 NLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGA 186
+LDL N +GPIP LGKL +L+ + L+NN L G IP SL N+ +L ++S NQL G
Sbjct: 789 SLDLYLNRFNGPIPDSLGKLSKLRFLRLNNNSLMGPIPMSLTNISALQVLDLSNNQLSGV 968
Query: 187 IP--AGLKKFNKNVFEHNKCLCG 207
+P F F +N LCG
Sbjct: 969 VPDNGSFSLFTPISFANNLNLCG 1037
>TC77085 similar to GP|21536600|gb|AAM60932.1 putative disease resistance
protein {Arabidopsis thaliana}, partial (88%)
Length = 1662
Score = 99.0 bits (245), Expect = 1e-21
Identities = 58/169 (34%), Positives = 85/169 (49%), Gaps = 4/169 (2%)
Frame = +2
Query: 28 CNADDKAALLKIRDHF-GGPKGRLDDWDNNTECCDWSFVGC---GRPYPGRVTVVTISRG 83
C+ DD++ LL + P L W T CC W VGC R +T T +
Sbjct: 128 CDPDDESGLLAFKSGIKSDPTSMLKSWIPGTNCCTWVGVGCLDNKRVTSLSLTGDTENPK 307
Query: 84 WGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFL 143
LSGT+ L +L + L + K++GP P+ KL L+ + + +N+LSGPIP +
Sbjct: 308 SFLSGTISPSLSKLKFLDGIYLINLLKISGPFPDFLFKLPNLKYIYIENNTLSGPIPQNI 487
Query: 144 GKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLK 192
G + +L+ L NK +G IP+S+ L L+Q + N L G IP LK
Sbjct: 488 GSMNQLEAFSLQENKFTGPIPSSISALTKLTQLKLGNNFLTGTIPVSLK 634
Score = 76.3 bits (186), Expect = 8e-15
Identities = 48/108 (44%), Positives = 62/108 (56%), Gaps = 1/108 (0%)
Frame = +2
Query: 82 RGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQ-RLQNLDLGSNSLSGPIP 140
+G LSG +P F +L L +L L+ K +G IP S S L L+ L+LG NSLSG IP
Sbjct: 665 QGNQLSGNIPDIFTSLKNLIILQLSHN-KFSGNIPLSISSLYPTLRYLELGHNSLSGKIP 841
Query: 141 SFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIP 188
FLGK K L +DLS N+ GT+P S NL + ++S N L P
Sbjct: 842 DFLGKFKALDTLDLSKNQFKGTVPKSFANLTKIFNLDLSDNFLVDPFP 985
Score = 74.3 bits (181), Expect = 3e-14
Identities = 44/106 (41%), Positives = 59/106 (55%)
Frame = +2
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
LSG +P G++ L SL E K TGPIP+S S L +L L LG+N L+G IP L
Sbjct: 461 LSGPIPQNIGSMNQLEAFSLQEN-KFTGPIPSSISALTKLTQLKLGNNFLTGTIPVSLKN 637
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGL 191
L L + L N+LSG IP +L++L +S N+ G IP +
Sbjct: 638 LTNLTYLSLQGNQLSGNIPDIFTSLKNLIILQLSHNKFSGNIPLSI 775
Score = 59.3 bits (142), Expect = 1e-09
Identities = 39/110 (35%), Positives = 51/110 (45%), Gaps = 23/110 (20%)
Frame = +2
Query: 128 LDLGSNSLSGPIPSFLGKLK-----------------------RLKEVDLSNNKLSGTIP 164
+DL N +SG L K + RLK +DLS+N + G +
Sbjct: 1163 IDLSGNEISGSAVGLLNKTEYLIEFRGSENLLKFDLESLKFGNRLKYLDLSHNLVFGKVT 1342
Query: 165 ASLGNLQSLSQFNVSFNQLCGAIPAGLKKFNKNVFEHNKCLCGAPLAPCK 214
S+ +Q L NVS+N+LCG IP F +VF CLCG PL PCK
Sbjct: 1343 KSVVGIQKL---NVSYNRLCGEIPKN--NFPASVFVGXDCLCGPPLMPCK 1477
>CB891120 similar to GP|17979065|gb putative receptor kinase homolog
{Arabidopsis thaliana}, partial (32%)
Length = 845
Score = 96.3 bits (238), Expect = 8e-21
Identities = 65/188 (34%), Positives = 95/188 (49%), Gaps = 1/188 (0%)
Frame = +2
Query: 2 FLLLHLTLLFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGRLDDWDN-NTECC 60
FLLL L LFLS FS ++ + + AL+ I++ P L +WD + + C
Sbjct: 44 FLLL-LFFLFLSHQPFSSASEPRN-----PEVVALMSIKEALNDPHNVLSNWDEFSVDPC 205
Query: 61 DWSFVGCGRPYPGRVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFS 120
W+ + C + + LSGTL + NL L + L + ++G IP
Sbjct: 206 SWAMITCSSD---SFVIGLGAPSQSLSGTLSSSIANLTNLKQV-LLQNNNISGKIPPELG 373
Query: 121 KLQRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSF 180
L +LQ LDL +N SG IPS L +L L+ + L+NN LSG P SL N+ L+ ++SF
Sbjct: 374 NLPKLQTLDLSNNRFSGFIPSSLNQLNSLQYMRLNNNSLSGPFPVSLSNITQLAFLDLSF 553
Query: 181 NQLCGAIP 188
N L G +P
Sbjct: 554 NNLTGPLP 577
>TC76944 similar to GP|13899097|gb|AAK48970.1 Unknown protein {Arabidopsis
thaliana}, partial (92%)
Length = 1248
Score = 94.4 bits (233), Expect = 3e-20
Identities = 73/204 (35%), Positives = 98/204 (47%), Gaps = 3/204 (1%)
Frame = +1
Query: 10 LFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGRLDDWDNN-TECCDWSFVGCG 68
L L+ I F+ +N+ D AL ++ P L WD C W V C
Sbjct: 172 LTLTLIQFTHANSEGD---------ALYTLKRSLTDPDNVLQSWDPTLVSPCTWFHVTCN 324
Query: 69 RPYPGRVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNL 128
+ RVT V + LSG L E G L +L L L + + G IP L+ L +L
Sbjct: 325 QD--NRVTRVDLGNS-NLSGHLVPELGKLEHLQYLELYKN-NIQGTIPKELGNLKSLVSL 492
Query: 129 DLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIP 188
DL +N++SG IP LGKLK L + L++N+L+G IP L + SL +VS N LCG IP
Sbjct: 493 DLYNNNISGTIPPSLGKLKNLVFLRLNDNRLTGPIPRELIAVTSLKVVDVSNNDLCGTIP 672
Query: 189 AG--LKKFNKNVFEHNKCLCGAPL 210
+ N FE+N L G L
Sbjct: 673 TSGPFEHIPLNNFENNPRLEGPEL 744
>TC89676 similar to PIR|D84889|D84889 probable receptor-like protein kinase
[imported] - Arabidopsis thaliana, partial (28%)
Length = 774
Score = 94.0 bits (232), Expect = 4e-20
Identities = 64/179 (35%), Positives = 92/179 (50%), Gaps = 23/179 (12%)
Frame = +1
Query: 36 LLKIRDHFGGPKGRLDDWDNNTECCDWSFVGCGRPYPGRVTVVTISRGWGLSGTLPAEFG 95
L+ I+D L W+N+++ C +F G G VT +++ +G GLSG +P+ G
Sbjct: 82 LMLIKDSLDPENHVLLSWNNHSDPCSGTFDGVACNEQGLVTNISL-QGKGLSGEIPSVIG 258
Query: 96 NLPYLSMLSL------AEMPK-----------------VTGPIPNSFSKLQRLQNLDLGS 132
L L+ L L +PK ++G IP+ + LQ L L
Sbjct: 259 KLKSLTGLYLHFNALNGILPKEIAGLTQLSDLYLNVNNLSGFIPHEIGNMSNLQVLQLCH 438
Query: 133 NSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGL 191
N L+G IP+ LGKLKRL + L N LSG IPASLG L++L + ++SFN L G IP +
Sbjct: 439 NELNGSIPTELGKLKRLSVLALQYNHLSGAIPASLGELETLERLDLSFNTLLGPIPVNI 615
Score = 67.0 bits (162), Expect = 5e-12
Identities = 38/83 (45%), Positives = 51/83 (60%)
Frame = +1
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
LSG +P E GN+ L +L L ++ G IP KL+RL L L N LSG IP+ LG+
Sbjct: 373 LSGFIPHEIGNMSNLQVLQLCHN-ELNGSIPTELGKLKRLSVLALQYNHLSGAIPASLGE 549
Query: 146 LKRLKEVDLSNNKLSGTIPASLG 168
L+ L+ +DLS N L G IP ++G
Sbjct: 550 LETLERLDLSFNTLLGPIPVNIG 618
>TC86662 weakly similar to GP|4756646|emb|CAB42250.1 unnamed protein product
{unidentified}, partial (76%)
Length = 1161
Score = 93.6 bits (231), Expect = 5e-20
Identities = 65/172 (37%), Positives = 85/172 (48%), Gaps = 3/172 (1%)
Frame = +2
Query: 35 ALLKIRDHFGGPKGRLDDWDNN-TECCDWSFVGCGRPYPGRVTVVTISRGWGLSGTLPAE 93
AL R+ P L WD C W V C RV+ + + GLSG+L +E
Sbjct: 263 ALHVFRNSLSDPNNVLQSWDPTLVNPCTWFHVTCDSN--NRVSRLDLGNA-GLSGSLGSE 433
Query: 94 FGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKLKRLKEVD 153
G+L +L L L + G IP KL+ L ++DL N L G IP GKLK L+ +
Sbjct: 434 LGHLHHLQYLELYGND-LRGKIPKELGKLKELISMDLYYNKLEGKIPKSFGKLKSLRFLR 610
Query: 154 LSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIP--AGLKKFNKNVFEHNK 203
L+NN L+G+IP L L L F+VS N LCG IP F FE+N+
Sbjct: 611 LNNNNLTGSIPRELTRLTHLEVFDVSNNDLCGTIPVDGNFGSFPIKSFENNR 766
>TC92543 weakly similar to GP|6041804|gb|AAF02124.1| putative protein kinase
{Arabidopsis thaliana}, partial (10%)
Length = 685
Score = 93.2 bits (230), Expect = 7e-20
Identities = 50/105 (47%), Positives = 66/105 (62%)
Frame = +1
Query: 85 GLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLG 144
GL G +P G + L LSLA ++G IP S ++ LQ LDL +NSL+G IP F+
Sbjct: 37 GLQGQIPTSLGQMKVLKFLSLAGN-NLSGSIPTSLGQMYSLQVLDLSTNSLTGEIPKFIE 213
Query: 145 KLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPA 189
++ L V L+NN LSG IPA L N+ +LS FNVSFN L G +P+
Sbjct: 214 NMRNLTNVLLNNNNLSGHIPAGLVNVTTLSAFNVSFNNLSGYLPS 348
Score = 66.6 bits (161), Expect = 7e-12
Identities = 34/79 (43%), Positives = 52/79 (65%)
Frame = +1
Query: 113 GPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQS 172
G IP S +++ L+ L L N+LSG IP+ LG++ L+ +DLS N L+G IP + N+++
Sbjct: 46 GQIPTSLGQMKVLKFLSLAGNNLSGSIPTSLGQMYSLQVLDLSTNSLTGEIPKFIENMRN 225
Query: 173 LSQFNVSFNQLCGAIPAGL 191
L+ ++ N L G IPAGL
Sbjct: 226 LTNVLLNNNNLSGHIPAGL 282
Score = 63.2 bits (152), Expect = 7e-11
Identities = 37/88 (42%), Positives = 51/88 (57%), Gaps = 2/88 (2%)
Frame = +1
Query: 122 LQRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFN 181
L L +L+L N L G IP+ LG++K LK + L+ N LSG+IP SLG + SL ++S N
Sbjct: 1 LVSLVSLNLSRNGLQGQIPTSLGQMKVLKFLSLAGNNLSGSIPTSLGQMYSLQVLDLSTN 180
Query: 182 QLCGAIPAGLKKFNK--NVFEHNKCLCG 207
L G IP ++ NV +N L G
Sbjct: 181 SLTGEIPKFIENMRNLTNVLLNNNNLSG 264
Score = 53.9 bits (128), Expect = 4e-08
Identities = 34/114 (29%), Positives = 57/114 (49%)
Frame = +1
Query: 83 GWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSF 142
G LSG++P G + L +L L+ +TG IP ++ L N+ L +N+LSG IP+
Sbjct: 103 GNNLSGSIPTSLGQMYSLQVLDLSTN-SLTGEIPKFIENMRNLTNVLLNNNNLSGHIPAG 279
Query: 143 LGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKKFNK 196
L + L ++S N LSG +P++ ++ S F C + + N+
Sbjct: 280 LVNVTTLSAFNVSFNNLSGYLPSNSSLIKCSSAVGNPFLSSCRGLSLTVPSANQ 441
>TC86633 weakly similar to SP|Q00874|D100_ARATH DNA-damage-repair/toleration
protein DRT100 precursor. [Mouse-ear cress] {Arabidopsis
thaliana}, partial (93%)
Length = 1407
Score = 90.5 bits (223), Expect = 4e-19
Identities = 64/219 (29%), Positives = 104/219 (47%), Gaps = 16/219 (7%)
Frame = +3
Query: 7 LTLLFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPK-GRLDDWDNNTECCDWSFV 65
LTLL L+ I KS C D+AALL + P G + W C W V
Sbjct: 45 LTLLLLTVISTVKS-------CPPSDRAALLAFKSALTEPNLGIFNSWSGYDCCRGWHGV 203
Query: 66 GCGRPYPGRVTVVTI------------SRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTG 113
C P RVT + + + ++G + E L L+ L +A+ ++G
Sbjct: 204 SCN-PTTWRVTDINLRGDSEDPIFQNLTHSGDMTGEISPEVCKLDELTTLVVADWKSISG 380
Query: 114 PIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSL 173
IP+ + L L+ LDL N +SG IP +GKL+ L ++L++N +SG IP S+ + L
Sbjct: 381 EIPSCITSLSSLRILDLTGNKISGNIPGNIGKLQHLTVLNLADNAISGEIPMSIVRISGL 560
Query: 174 SQFNVSFNQLCGAIPAG---LKKFNKNVFEHNKCLCGAP 209
+++ NQ+ G +P+ L++ ++ +F N+ P
Sbjct: 561 MHLDLAGNQISGELPSDIGKLRRLSRALFSRNQLTGSIP 677
Score = 84.7 bits (208), Expect = 2e-17
Identities = 42/108 (38%), Positives = 70/108 (63%)
Frame = +3
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
+SG +P G L +L++L+LA+ ++G IP S ++ L +LDL N +SG +PS +GK
Sbjct: 444 ISGNIPGNIGKLQHLTVLNLADNA-ISGEIPMSIVRISGLMHLDLAGNQISGELPSDIGK 620
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKK 193
L+RL S N+L+G+IP S+ + L+ ++S N++ G+IPA + K
Sbjct: 621 LRRLSRALFSRNQLTGSIPDSVLKMNRLADLDLSMNRITGSIPARIGK 764
Score = 81.3 bits (199), Expect = 3e-16
Identities = 46/130 (35%), Positives = 66/130 (50%), Gaps = 2/130 (1%)
Frame = +3
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
++G++PA G + LS L L + +TG IP++ + L+L N G IP G
Sbjct: 732 ITGSIPARIGKMRVLSTLKL-DGNSMTGQIPSTLLSNTGMGILNLSRNGFEGTIPDVFGS 908
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAG--LKKFNKNVFEHNK 203
+DLS NKL+G IP SL + + + ++S N LCG IP G + F +N
Sbjct: 909 KSYFMVLDLSFNKLTGRIPGSLSSAKFMGHLDISNNHLCGTIPIGSPFDHLDAASFSNND 1088
Query: 204 CLCGAPLAPC 213
C CG PL C
Sbjct: 1089CSCGNPLKTC 1118
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.320 0.139 0.436
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,518,199
Number of Sequences: 36976
Number of extensions: 108780
Number of successful extensions: 1501
Number of sequences better than 10.0: 264
Number of HSP's better than 10.0 without gapping: 899
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1209
length of query: 214
length of database: 9,014,727
effective HSP length: 92
effective length of query: 122
effective length of database: 5,612,935
effective search space: 684778070
effective search space used: 684778070
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 56 (26.2 bits)
Medicago: description of AC146750.9


