

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146750.10 + phase: 0
(221 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
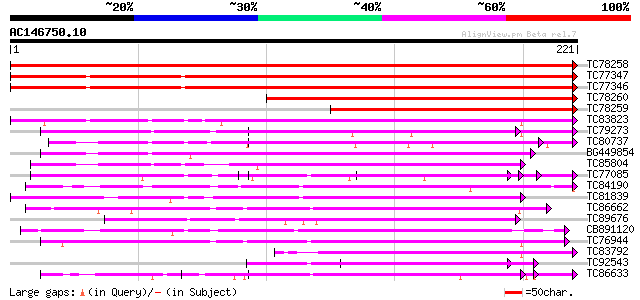
Score E
Sequences producing significant alignments: (bits) Value
TC78258 similar to GP|21593869|gb|AAM65836.1 polygalacturonase i... 451 e-127
TC77347 weakly similar to GP|22324851|gb|AAM95647.1 polygalactur... 374 e-104
TC77346 weakly similar to GP|18542897|gb|AAL75739.1 Putative rec... 369 e-103
TC78260 weakly similar to PIR|T00712|T00712 protein kinase homol... 244 2e-65
TC78259 weakly similar to PIR|T00712|T00712 protein kinase homol... 193 5e-50
TC83823 weakly similar to GP|21593869|gb|AAM65836.1 polygalactur... 155 9e-39
TC79273 weakly similar to PIR|S47965|S47965 polygalacturonase in... 137 3e-33
TC80737 weakly similar to GP|1143381|emb|CAA88846.1 polygalactur... 122 8e-29
BG449854 weakly similar to GP|5579315|gb| polygalacturonase inhi... 118 2e-27
TC85804 weakly similar to GP|8778050|gb|AAF79181.1| polygalactur... 110 3e-25
TC77085 similar to GP|21536600|gb|AAM60932.1 putative disease re... 98 3e-21
TC84190 weakly similar to GP|4008006|gb|AAC95351.1| receptor-lik... 97 4e-21
TC81839 similar to GP|21593619|gb|AAM65586.1 receptor protein ki... 97 6e-21
TC86662 weakly similar to GP|4756646|emb|CAB42250.1 unnamed prot... 95 2e-20
TC89676 similar to PIR|D84889|D84889 probable receptor-like prot... 94 5e-20
CB891120 similar to GP|17979065|gb putative receptor kinase homo... 93 9e-20
TC76944 similar to GP|13899097|gb|AAK48970.1 Unknown protein {Ar... 93 9e-20
TC83792 similar to GP|9280288|dbj|BAB01743.1 receptor protein ki... 92 2e-19
TC92543 weakly similar to GP|6041804|gb|AAF02124.1| putative pro... 91 5e-19
TC86633 weakly similar to SP|Q00874|D100_ARATH DNA-damage-repair... 90 6e-19
>TC78258 similar to GP|21593869|gb|AAM65836.1 polygalacturonase inhibiting
protein 1 {Arabidopsis thaliana}, partial (18%)
Length = 885
Score = 451 bits (1160), Expect = e-127
Identities = 217/221 (98%), Positives = 220/221 (99%)
Frame = +2
Query: 1 MKTTPNKFLLLHLTLLFLSSIYFSTSNADDVEFSCNANDKATLLKIRDHFGGPNGRLSDW 60
MKTTPNKFLLLHLTLLFLSSIYFSTSNADDVEFSCNANDKATLLKIRDHFGGPNGRLSDW
Sbjct: 50 MKTTPNKFLLLHLTLLFLSSIYFSTSNADDVEFSCNANDKATLLKIRDHFGGPNGRLSDW 229
Query: 61 DNGTDCCSDWSFVGCGKPLPGRITVVTISRGWGLSGTLPAEFGDLPYLNFLSLAEMPKVT 120
DNGTDCCSDWSFVGCGKPLPGRITVVTISRGWGLSGTLPAEFGDLPYLNFLSLAEMPKVT
Sbjct: 230 DNGTDCCSDWSFVGCGKPLPGRITVVTISRGWGLSGTLPAEFGDLPYLNFLSLAEMPKVT 409
Query: 121 GPIPNSFSKLQRLQKLDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPS 180
GPIPNSFSKL+RLQKLDLGSNSLSG IPTFLGQLKGLQEFDLSNNKLTGVIPASFGS+PS
Sbjct: 410 GPIPNSFSKLKRLQKLDLGSNSLSGPIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSIPS 589
Query: 181 LSQFNVSFNQLCGAIPTGLSKFAKSSFDHNKCLCGAPLAAC 221
LSQFNVSFNQLCGAIPTGL+KFAKSSFDHNKCLCGAPLAAC
Sbjct: 590 LSQFNVSFNQLCGAIPTGLNKFAKSSFDHNKCLCGAPLAAC 712
>TC77347 weakly similar to GP|22324851|gb|AAM95647.1 polygalacturonase
inhibitory protein {Brassica napus}, partial (21%)
Length = 926
Score = 374 bits (961), Expect = e-104
Identities = 182/221 (82%), Positives = 197/221 (88%)
Frame = +3
Query: 1 MKTTPNKFLLLHLTLLFLSSIYFSTSNADDVEFSCNANDKATLLKIRDHFGGPNGRLSDW 60
MKT+PN FLLLHLTLLFLSSIYFS SNA D +F CNA+DKA LLKIRDHFGGP GRL DW
Sbjct: 102 MKTSPNMFLLLHLTLLFLSSIYFSKSNASD-DFFCNADDKAALLKIRDHFGGPKGRLDDW 278
Query: 61 DNGTDCCSDWSFVGCGKPLPGRITVVTISRGWGLSGTLPAEFGDLPYLNFLSLAEMPKVT 120
DN T+CC DWSFVGCG+P PGR+TVVTISRGWGLSGTLPAEFG+LPYL+ LSLAEMPKVT
Sbjct: 279 DNNTECC-DWSFVGCGRPYPGRVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVT 455
Query: 121 GPIPNSFSKLQRLQKLDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPS 180
GPIPNSFSKLQRLQ LDLGSNSLSG IP+FLG+LK L+E DLSNNKL+G IPAS G+L S
Sbjct: 456 GPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQS 635
Query: 181 LSQFNVSFNQLCGAIPTGLSKFAKSSFDHNKCLCGAPLAAC 221
LSQFNVSFNQLCGAIP GL KF K+ F+HNKCLCGAPLA C
Sbjct: 636 LSQFNVSFNQLCGAIPAGLKKFNKNVFEHNKCLCGAPLAPC 758
>TC77346 weakly similar to GP|18542897|gb|AAL75739.1 Putative receptor-like
protein kinase {Oryza sativa}, partial (6%)
Length = 865
Score = 369 bits (946), Expect = e-103
Identities = 180/221 (81%), Positives = 194/221 (87%)
Frame = +2
Query: 1 MKTTPNKFLLLHLTLLFLSSIYFSTSNADDVEFSCNANDKATLLKIRDHFGGPNGRLSDW 60
MKT PN F+LL LTL FLSSIYFS SNA D +F CNA+DKA LLKIRDHFGGPNGRL DW
Sbjct: 38 MKTPPNMFMLLSLTLFFLSSIYFSKSNASD-DFFCNADDKAALLKIRDHFGGPNGRLDDW 214
Query: 61 DNGTDCCSDWSFVGCGKPLPGRITVVTISRGWGLSGTLPAEFGDLPYLNFLSLAEMPKVT 120
DN T+CC DW+FVGCG+P PGR+T VTI RGWGLSGTLPAEFGDLPYL+ LSLAEMPKVT
Sbjct: 215 DNNTECC-DWNFVGCGRPYPGRVTGVTIGRGWGLSGTLPAEFGDLPYLSMLSLAEMPKVT 391
Query: 121 GPIPNSFSKLQRLQKLDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPS 180
GPIPNSFSKL+RL+ LDLGSNSLSG IP FLG+LK LQE DLSNNKL+G IPAS +L S
Sbjct: 392 GPIPNSFSKLKRLENLDLGSNSLSGPIPEFLGKLKRLQEVDLSNNKLSGAIPASLDNLQS 571
Query: 181 LSQFNVSFNQLCGAIPTGLSKFAKSSFDHNKCLCGAPLAAC 221
LSQFNVSFNQLCGAIPTGLSKFAKSSF+HNKCLC APLA C
Sbjct: 572 LSQFNVSFNQLCGAIPTGLSKFAKSSFEHNKCLCDAPLAPC 694
>TC78260 weakly similar to PIR|T00712|T00712 protein kinase homolog F22O13.7
- Arabidopsis thaliana, partial (7%)
Length = 563
Score = 244 bits (623), Expect = 2e-65
Identities = 120/121 (99%), Positives = 120/121 (99%)
Frame = +3
Query: 101 EFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQRLQKLDLGSNSLSGSIPTFLGQLKGLQEF 160
EFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQRLQKLDLGSNSLSGSIPTFLGQLKGLQEF
Sbjct: 3 EFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQRLQKLDLGSNSLSGSIPTFLGQLKGLQEF 182
Query: 161 DLSNNKLTGVIPASFGSLPSLSQFNVSFNQLCGAIPTGLSKFAKSSFDHNKCLCGAPLAA 220
DLSNNKLTGVIPASFGSLPSLSQFNVSFNQLCGAIPTGLSKFAKSSFDHNKCLCGAPL A
Sbjct: 183 DLSNNKLTGVIPASFGSLPSLSQFNVSFNQLCGAIPTGLSKFAKSSFDHNKCLCGAPLTA 362
Query: 221 C 221
C
Sbjct: 363 C 365
>TC78259 weakly similar to PIR|T00712|T00712 protein kinase homolog F22O13.7
- Arabidopsis thaliana, partial (6%)
Length = 522
Score = 193 bits (490), Expect = 5e-50
Identities = 95/96 (98%), Positives = 96/96 (99%)
Frame = +2
Query: 126 SFSKLQRLQKLDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFN 185
+FSKLQRLQKLDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFN
Sbjct: 32 AFSKLQRLQKLDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFN 211
Query: 186 VSFNQLCGAIPTGLSKFAKSSFDHNKCLCGAPLAAC 221
VSFNQLCGAIPTGLSKFAKSSFDHNKCLCGAPLAAC
Sbjct: 212 VSFNQLCGAIPTGLSKFAKSSFDHNKCLCGAPLAAC 319
>TC83823 weakly similar to GP|21593869|gb|AAM65836.1 polygalacturonase
inhibiting protein 1 {Arabidopsis thaliana}, partial
(38%)
Length = 798
Score = 155 bits (393), Expect = 9e-39
Identities = 99/232 (42%), Positives = 125/232 (53%), Gaps = 11/232 (4%)
Frame = +1
Query: 1 MKTTPNKFLLLH----LTLLFLSSIYFSTSNADDVEFSCNANDKATLLKIRDHFGGPNGR 56
M+ P+ L+L L LLFL S+ FS S C+ +D+ LLKI+DHF +
Sbjct: 10 MERNPSMSLILFPSFLLLLLFLPSLTFSLSPPPR-RHHCHPDDEKVLLKIKDHFHNTS-L 183
Query: 57 LSDWDNGTDCCSDWSFVGCGKPLPG----RITVVTISRGWGLSGTLPAEFGDLPYLNFLS 112
S W TDCC W V C K +P R+ + I L GT+P DLPYL L
Sbjct: 184 FSTWTPKTDCCK-WRIVLC-KKIPKTTIYRVNFLEIDGADDLVGTIPPLIADLPYLETLI 357
Query: 113 LAEMPKVTGPIPNSFSKLQRLQKLDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIP 172
+P +TGPIP + +KL L + L NSL+G IP + +L L L+NN LTG +P
Sbjct: 358 FRNLPNLTGPIPQAIAKLPHLNFVLLNWNSLTGPIPDYFSKLTNLATLGLNNNHLTGRVP 537
Query: 173 ASFGSLPSLSQFNVSFNQLCGAIPTG---LSKFAKSSFDHNKCLCGAPLAAC 221
A G LP L +VS NQLCG IP L F S F++NKCLCGAPLA C
Sbjct: 538 AYLGRLPKLVGLDVSHNQLCGPIPKDGGKLQGFDPSWFENNKCLCGAPLAPC 693
>TC79273 weakly similar to PIR|S47965|S47965 polygalacturonase inhibitor
protein - tomato, partial (37%)
Length = 1399
Score = 137 bits (345), Expect = 3e-33
Identities = 76/187 (40%), Positives = 102/187 (53%)
Frame = +2
Query: 13 LTLLFLSSIYFSTSNADDVEFSCNANDKATLLKIRDHFGGPNGRLSDWDNGTDCCSDWSF 72
L+L+F ++ S+ N CN NDK LL + F + + WD TDCC +WS
Sbjct: 23 LSLIFTFILFISSLNLSFSTTICNTNDKNVLLGXKSQFNNASV-FTTWDPITDCCKNWSG 199
Query: 73 VGCGKPLPGRITVVTISRGWGLSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQR 132
+ C GR+T++ +S + G +P +LP+L F + A P V+G IP + +KL
Sbjct: 200 IECNSN--GRVTMLAVSDTNDVIGEIPTSVVNLPFLQFFTFAVFPGVSGTIPPAIAKLTN 373
Query: 133 LQKLDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQLC 192
L LD +SL+G IP FLGQLK L DLS N+ TG IPAS G L L N+ NQL
Sbjct: 374 LVHLDFSLDSLTGPIPDFLGQLKNLDVIDLSGNRFTGQIPASLGRLTKLRSANLGSNQLS 553
Query: 193 GAIPTGL 199
G IP L
Sbjct: 554 GPIPASL 574
Score = 95.5 bits (236), Expect = 1e-20
Identities = 64/178 (35%), Positives = 84/178 (46%), Gaps = 50/178 (28%)
Frame = +2
Query: 94 LSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQR-LQKLDLGSNSLSGSIPTFLG 152
LSG +PA LP LN LSL + ++TG IP SF + +DL SN+LSG IP+ G
Sbjct: 620 LSGPIPASLAQLPKLNELSLFQN-QLTGSIPESFGSFKNPALNIDLSSNNLSGPIPSSFG 796
Query: 153 QLK-----------------------------------------------GLQEFDLSNN 165
+ K GL+ D+S+N
Sbjct: 797 KAKITALVLSKNKFSGDASFLFGKDKTVLTTMDLSNNIFKFDFSNVDLSPGLKNLDISHN 976
Query: 166 KLTGVIPASFGSLPSLSQFNVSFNQLCGAIPTG--LSKFAKSSFDHNKCLCGAPLAAC 221
+ G +P G LP L + +VSFNQLCG IPTG L +F+ + F +N CLCG PL AC
Sbjct: 977 MIFGSLPEKLGQLP-LQRLDVSFNQLCGQIPTGRRLKQFSPTKFSNNTCLCGLPLPAC 1147
>TC80737 weakly similar to GP|1143381|emb|CAA88846.1 polygalacturonase
inhibitor {Actinidia deliciosa}, partial (77%)
Length = 1176
Score = 122 bits (307), Expect = 8e-29
Identities = 77/196 (39%), Positives = 101/196 (51%), Gaps = 3/196 (1%)
Frame = +2
Query: 16 LFLSSIYFSTSNADDVEFSCNANDKATLLKIRDHFGGPNGRLSDWDNGTDCCSDWSFVGC 75
L LS I FS CN DK LL+I+ P L+ WD TDCC W V C
Sbjct: 50 LLLSPIAFSEK--------CNPQDKRALLRIKKELNNPY-LLASWDPQTDCCG-WYCVKC 199
Query: 76 GKPLPGRITVVTISRG---WGLSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQR 132
+ RIT + + LSGT+P GDLPYL L ++P++ GPI + +KL +
Sbjct: 200 DL-ITHRITALIMQSSVPDTNLSGTIPPSVGDLPYLENLEFHKLPRLKGPIQPTIAKLTK 376
Query: 133 LQKLDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQLC 192
L+ L + ++SG IP FL QLK LQ LS N L+G IP+S LP+L ++ N+L
Sbjct: 377 LKYLFIEYTNVSGPIPPFLAQLKNLQLLHLSTNNLSGPIPSSLSQLPNLESLHLDRNKLT 556
Query: 193 GAIPTGLSKFAKSSFD 208
G IP F K D
Sbjct: 557 GPIPESFGSFKKPGPD 604
Score = 76.3 bits (186), Expect = 9e-15
Identities = 59/181 (32%), Positives = 77/181 (41%), Gaps = 53/181 (29%)
Frame = +2
Query: 94 LSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQRL-QKLDLGSNSLSGSIPTFLG 152
LSG +P+ LP L L L + K+TGPIP SF ++ + L N LSG IP LG
Sbjct: 479 LSGPIPSSLSQLPNLESLHL-DRNKLTGPIPESFGSFKKPGPDIILSHNQLSGPIPASLG 655
Query: 153 QL--------------------------------KGLQEFDLS---------------NN 165
Q+ + L FD S +N
Sbjct: 656 QIDPERIDLSRNKLEGDASVLFGSQKRTQILDVSRNLLSFDFSKVDFPKQSLIWLDINHN 835
Query: 166 KLTGVIPASFGSLPSLSQFNVSFNQLCGAIPTGLSKFAKSSFD-----HNKCLCGAPLAA 220
K+ G IP + + +L QFNVS+N+L G IP G + + FD HNK LCG PL
Sbjct: 836 KIYGKIPVTLTKVENLQQFNVSYNKLSGEIPQGGTGRLQDRFDVYAYLHNKGLCGPPLPK 1015
Query: 221 C 221
C
Sbjct: 1016C 1018
>BG449854 weakly similar to GP|5579315|gb| polygalacturonase inhibitor
protein {Glycine max}, partial (46%)
Length = 635
Score = 118 bits (296), Expect = 2e-27
Identities = 73/194 (37%), Positives = 103/194 (52%), Gaps = 1/194 (0%)
Frame = +3
Query: 13 LTLLFLSSIYFSTSNADDVEFSCNANDKATLLKIRDHFGGPNGRLSDWDNGTDCCSD-WS 71
L LL L++ F S ++ CN +DK TL +I+ FG P +LS WD+ TDCC+ W
Sbjct: 27 LLLLILTTHLFIPSLSEQ----CNPHDKKTLFQIKKQFGNPT-KLSSWDSTTDCCNGTWY 191
Query: 72 FVGCGKPLPGRITVVTISRGWGLSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQ 131
V C K T+ +P +LP+L +L+L P + G IP S S L
Sbjct: 192 GVFCDKQTYRVSTLDLQDLDLPHPVPIPPSIFNLPFLLYLTLMNTPNLVGTIPPSISNLT 371
Query: 132 RLQKLDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQL 191
+L + + ++SG IP L ++K L D+S+NKLTG +PAS +LP+L+ NQL
Sbjct: 372 KLNSIYIIQTNISGEIPNTLSEIKTLVTIDISDNKLTGPLPASLSTLPNLAGIFFYRNQL 551
Query: 192 CGAIPTGLSKFAKS 205
GAIP F KS
Sbjct: 552 TGAIPKSYGSFPKS 593
>TC85804 weakly similar to GP|8778050|gb|AAF79181.1| polygalacturonase
inhibiting protein {Prunus mahaleb}, partial (51%)
Length = 1094
Score = 110 bits (276), Expect = 3e-25
Identities = 75/212 (35%), Positives = 101/212 (47%), Gaps = 19/212 (8%)
Frame = +1
Query: 9 LLLHLTLLFLSSIYFSTSNADDVEFSCNANDKATLLKIRDHFGGPNGRLSDWDNGTDCCS 68
LL +LL L + S CN DK LL+I+ P LS W+ +CC
Sbjct: 28 LLCFFSLLLLFPVAISQK--------CNPQDKKALLQIKKELNNPTS-LSSWNPRKNCC- 177
Query: 69 DWSFVGCGKPLPGRITVVTISRGWGLS-------------------GTLPAEFGDLPYLN 109
DW F+ C VT SR L+ G + GDL Y+
Sbjct: 178 DWVFIHCD---------VTTSRVIWLAIQFSSPDQFTTPFPNPEFIGHISPSVGDLSYVE 330
Query: 110 FLSLAEMPKVTGPIPNSFSKLQRLQKLDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLTG 169
L ++P VTG IP++ SKL+ L+ L + S+SG IP+FLGQ K L+ DL +NKLTG
Sbjct: 331 RLEFNQLPNVTGQIPSTISKLKNLKYLTISGTSVSGPIPSFLGQFKNLELLDLYSNKLTG 510
Query: 170 VIPASFGSLPSLSQFNVSFNQLCGAIPTGLSK 201
IP+S L +L Q + N+L G IP L +
Sbjct: 511 SIPSSLSQLTNLKQLFLHENKLSGHIPASLGQ 606
>TC77085 similar to GP|21536600|gb|AAM60932.1 putative disease resistance
protein {Arabidopsis thaliana}, partial (88%)
Length = 1662
Score = 97.8 bits (242), Expect = 3e-21
Identities = 64/205 (31%), Positives = 101/205 (49%), Gaps = 6/205 (2%)
Frame = +2
Query: 9 LLLHLTLLFLSSIYFSTSNADDVEFSCNANDKATLLKIRDHF-GGPNGRLSDWDNGTDCC 67
L L TLL L +I + + C+ +D++ LL + P L W GT+CC
Sbjct: 50 LFLTTTLLILVTISTTLLPHNAYGAKCDPDDESGLLAFKSGIKSDPTSMLKSWIPGTNCC 229
Query: 68 SDWSFVGCGKPLPGRITVVTISRGWG-----LSGTLPAEFGDLPYLNFLSLAEMPKVTGP 122
+ W VGC R+T ++++ LSGT+ L +L+ + L + K++GP
Sbjct: 230 T-WVGVGCLDNK--RVTSLSLTGDTENPKSFLSGTISPSLSKLKFLDGIYLINLLKISGP 400
Query: 123 IPNSFSKLQRLQKLDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPSLS 182
P+ KL L+ + + +N+LSG IP +G + L+ F L NK TG IP+S +L L+
Sbjct: 401 FPDFLFKLPNLKYIYIENNTLSGPIPQNIGSMNQLEAFSLQENKFTGPIPSSISALTKLT 580
Query: 183 QFNVSFNQLCGAIPTGLSKFAKSSF 207
Q + N L G IP L ++
Sbjct: 581 QLKLGNNFLTGTIPVSLKNLTNLTY 655
Score = 77.0 bits (188), Expect = 5e-15
Identities = 46/107 (42%), Positives = 60/107 (55%)
Frame = +2
Query: 94 LSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQRLQKLDLGSNSLSGSIPTFLGQ 153
LSG +P G + L SL E K TGPIP+S S L +L +L LG+N L+G+IP L
Sbjct: 461 LSGPIPQNIGSMNQLEAFSLQEN-KFTGPIPSSISALTKLTQLKLGNNFLTGTIPVSLKN 637
Query: 154 LKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQLCGAIPTGLS 200
L L L N+L+G IP F SL +L +S N+ G IP +S
Sbjct: 638 LTNLTYLSLQGNQLSGNIPDIFTSLKNLIILQLSHNKFSGNIPLSIS 778
Score = 72.0 bits (175), Expect = 2e-13
Identities = 46/108 (42%), Positives = 59/108 (54%), Gaps = 1/108 (0%)
Frame = +2
Query: 90 RGWGLSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQ-RLQKLDLGSNSLSGSIP 148
+G LSG +P F L L L L+ K +G IP S S L L+ L+LG NSLSG IP
Sbjct: 665 QGNQLSGNIPDIFTSLKNLIILQLSHN-KFSGNIPLSISSLYPTLRYLELGHNSLSGKIP 841
Query: 149 TFLGQLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQLCGAIP 196
FLG+ K L DLS N+ G +P SF +L + ++S N L P
Sbjct: 842 DFLGKFKALDTLDLSKNQFKGTVPKSFANLTKIFNLDLSDNFLVDPFP 985
Score = 48.5 bits (114), Expect = 2e-06
Identities = 35/109 (32%), Positives = 48/109 (43%), Gaps = 23/109 (21%)
Frame = +2
Query: 136 LDLGSNSLSGSIPTFLGQLKGLQEF-----------------------DLSNNKLTGVIP 172
+DL N +SGS L + + L EF DLS+N + G +
Sbjct: 1163 IDLSGNEISGSAVGLLNKTEYLIEFRGSENLLKFDLESLKFGNRLKYLDLSHNLVFGKVT 1342
Query: 173 ASFGSLPSLSQFNVSFNQLCGAIPTGLSKFAKSSFDHNKCLCGAPLAAC 221
S + + + NVS+N+LCG IP + F S F CLCG PL C
Sbjct: 1343 KS---VVGIQKLNVSYNRLCGEIPK--NNFPASVFVGXDCLCGPPLMPC 1474
>TC84190 weakly similar to GP|4008006|gb|AAC95351.1| receptor-like protein
kinase {Arabidopsis thaliana}, partial (21%)
Length = 833
Score = 97.4 bits (241), Expect = 4e-21
Identities = 85/238 (35%), Positives = 111/238 (45%), Gaps = 23/238 (9%)
Frame = +3
Query: 7 KFLLLHLTLLFLSSIYFSTSNADDVEFSCNANDKATLLKIRDHFGGPNGRLSDWDNGTDC 66
K +LL+ T + +I S AD A+D+A+LL +R GG R W++
Sbjct: 99 KTVLLYFTACLIITI---VSGAD------LASDRASLLTLRATVGG---RTLLWNSTETN 242
Query: 67 CSDWSFVGCGKPLPGRITVVTISRGWGLSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNS 126
W+ V C R+T + + GLSG LP+ G+L L LSL +TGPIP
Sbjct: 243 PCLWTGVICNNK---RVTALRLP-AMGLSGNLPSGIGNLTELQTLSL-RYNALTGPIPMD 407
Query: 127 FSKLQRLQKLDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSL-------- 178
F+KL L+ L L SN SG +P FL L+ L +L N +G I F +L
Sbjct: 408 FAKLVSLRNLYLHSNFFSGEVPEFLYGLQNLVRLNLGKNNFSGEISQHFNNLTRLDTLFL 587
Query: 179 --------------PSLSQFNVSFNQLCGAIPTGLSKFAKSSFDHNKCLCGAPL-AAC 221
P L QFNVSFN L G IP S+ S+F N LCG PL AC
Sbjct: 588 EQNMFTGSVPDLNIPPLHQFNVSFNNLTGQIPKRFSRLNISAFSGNS-LCGNPLQVAC 758
>TC81839 similar to GP|21593619|gb|AAM65586.1 receptor protein kinase-like
protein {Arabidopsis thaliana}, partial (34%)
Length = 960
Score = 96.7 bits (239), Expect = 6e-21
Identities = 67/202 (33%), Positives = 96/202 (47%), Gaps = 1/202 (0%)
Frame = +2
Query: 1 MKTTPNKFLLLHLTLLFLSSIYFSTSNADDVEFSCNANDKATLLKIRDHFGGPNGRLSDW 60
MK + L +TL S + + + F A L+ I++ P+G +W
Sbjct: 131 MKMFRGEVFLCFVTLFMFWSCANALLSPKGINFEVQA-----LVSIKESLMDPHGIFENW 295
Query: 61 D-NGTDCCSDWSFVGCGKPLPGRITVVTISRGWGLSGTLPAEFGDLPYLNFLSLAEMPKV 119
D + D CS W+ V C P + V LSGTL + G+L L + L + +
Sbjct: 296 DGDAVDPCS-WNMVTCS---PENLVVSLGIPSQNLSGTLSSSIGNLTNLQTVVL-QNNNI 460
Query: 120 TGPIPNSFSKLQRLQKLDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSLP 179
TGPIP+ KL LQ LDL N G IP LG L+ LQ L+NN +G P S ++
Sbjct: 461 TGPIPSELGKLSMLQTLDLSDNLFHGKIPPSLGHLRNLQYLRLNNNSFSGECPESLANMA 640
Query: 180 SLSQFNVSFNQLCGAIPTGLSK 201
L+ ++SFN L G +P L+K
Sbjct: 641 QLAFLDLSFNNLTGNVPRILAK 706
>TC86662 weakly similar to GP|4756646|emb|CAB42250.1 unnamed protein product
{unidentified}, partial (76%)
Length = 1161
Score = 95.1 bits (235), Expect = 2e-20
Identities = 74/211 (35%), Positives = 100/211 (47%), Gaps = 6/211 (2%)
Frame = +2
Query: 7 KFLLLHLTLLFLSSIYFSTSNADDVEF---SCNANDKATLLKI-RDHFGGPNGRLSDWDN 62
K LL H+T + SI FS ++F + AN + L + R+ PN L WD
Sbjct: 149 KDLLTHMTKM-APSIPFSVLFILLLQFPFQTITANSEGNALHVFRNSLSDPNNVLQSWDP 325
Query: 63 GTDCCSDWSFVGCGKPLPGRITVVTISRGWGLSGTLPAEFGDLPYLNFLSLAEMPKVTGP 122
W V C R++ + + GLSG+L +E G L +L +L L + G
Sbjct: 326 TLVNPCTWFHVTCDSN--NRVSRLDLGNA-GLSGSLGSELGHLHHLQYLELYGND-LRGK 493
Query: 123 IPNSFSKLQRLQKLDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPSLS 182
IP KL+ L +DL N L G IP G+LK L+ L+NN LTG IP L L
Sbjct: 494 IPKELGKLKELISMDLYYNKLEGKIPKSFGKLKSLRFLRLNNNNLTGSIPRELTRLTHLE 673
Query: 183 QFNVSFNQLCGAIPT--GLSKFAKSSFDHNK 211
F+VS N LCG IP F SF++N+
Sbjct: 674 VFDVSNNDLCGTIPVDGNFGSFPIKSFENNR 766
>TC89676 similar to PIR|D84889|D84889 probable receptor-like protein kinase
[imported] - Arabidopsis thaliana, partial (28%)
Length = 774
Score = 93.6 bits (231), Expect = 5e-20
Identities = 67/185 (36%), Positives = 95/185 (51%), Gaps = 23/185 (12%)
Frame = +1
Query: 38 NDKATLLKIRDHFGGPNGRLSDWDNGTDCCSDWSFVGCGKPLPGRITVVTISRGWGLSGT 97
N+ TL+ I+D N L W+N +D CS +F G G +T +++ +G GLSG
Sbjct: 67 NELDTLMLIKDSLDPENHVLLSWNNHSDPCSG-TFDGVACNEQGLVTNISL-QGKGLSGE 240
Query: 98 LPAEFGDLP-----YLNFLSL-AEMPK-----------------VTGPIPNSFSKLQRLQ 134
+P+ G L YL+F +L +PK ++G IP+ + LQ
Sbjct: 241 IPSVIGKLKSLTGLYLHFNALNGILPKEIAGLTQLSDLYLNVNNLSGFIPHEIGNMSNLQ 420
Query: 135 KLDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQLCGA 194
L L N L+GSIPT LG+LK L L N L+G IPAS G L +L + ++SFN L G
Sbjct: 421 VLQLCHNELNGSIPTELGKLKRLSVLALQYNHLSGAIPASLGELETLERLDLSFNTLLGP 600
Query: 195 IPTGL 199
IP +
Sbjct: 601 IPVNI 615
Score = 29.6 bits (65), Expect = 0.96
Identities = 13/24 (54%), Positives = 19/24 (79%)
Frame = +2
Query: 132 RLQKLDLGSNSLSGSIPTFLGQLK 155
+L+ LD+ +NSLSGS+PT L + K
Sbjct: 629 KLETLDIRNNSLSGSVPTDLKKTK 700
>CB891120 similar to GP|17979065|gb putative receptor kinase homolog
{Arabidopsis thaliana}, partial (32%)
Length = 845
Score = 92.8 bits (229), Expect = 9e-20
Identities = 72/215 (33%), Positives = 104/215 (47%), Gaps = 1/215 (0%)
Frame = +2
Query: 5 PNKFLLLHLTLLFLSSIYFSTSNADDVEFSCNANDKATLLKIRDHFGGPNGRLSDWDN-G 63
P FLLL L LFLS FS+++ + L+ I++ P+ LS+WD
Sbjct: 35 PLNFLLL-LFFLFLSHQPFSSASEP------RNPEVVALMSIKEALNDPHNVLSNWDEFS 193
Query: 64 TDCCSDWSFVGCGKPLPGRITVVTISRGWGLSGTLPAEFGDLPYLNFLSLAEMPKVTGPI 123
D CS W+ + C + + LSGTL + +L L + L + ++G I
Sbjct: 194 VDPCS-WAMITCSSD---SFVIGLGAPSQSLSGTLSSSIANLTNLKQV-LLQNNNISGKI 358
Query: 124 PNSFSKLQRLQKLDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPSLSQ 183
P L +LQ LDL +N SG IP+ L QL LQ L+NN L+G P S ++ L+
Sbjct: 359 PPELGNLPKLQTLDLSNNRFSGFIPSSLNQLNSLQYMRLNNNSLSGPFPVSLSNITQLAF 538
Query: 184 FNVSFNQLCGAIPTGLSKFAKSSFDHNKCLCGAPL 218
++SFN L G +P KF SF+ + G PL
Sbjct: 539 LDLSFNNLTGPLP----KFPARSFN----IVGNPL 619
>TC76944 similar to GP|13899097|gb|AAK48970.1 Unknown protein {Arabidopsis
thaliana}, partial (92%)
Length = 1248
Score = 92.8 bits (229), Expect = 9e-20
Identities = 68/209 (32%), Positives = 103/209 (48%), Gaps = 3/209 (1%)
Frame = +1
Query: 13 LTLLFLS-SIYFSTSNADDVEFSCNANDKATLLKIRDHFGGPNGRLSDWDNGTDCCSDWS 71
+T +F S S++ T ++F+ ++ L ++ P+ L WD W
Sbjct: 130 MTPIFTSISLFTITLTLTLIQFTHANSEGDALYTLKRSLTDPDNVLQSWDPTLVSPCTWF 309
Query: 72 FVGCGKPLPGRITVVTISRGWGLSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQ 131
V C + R+T V + LSG L E G L +L +L L + + G IP L+
Sbjct: 310 HVTCNQD--NRVTRVDLGNS-NLSGHLVPELGKLEHLQYLELYKN-NIQGTIPKELGNLK 477
Query: 132 RLQKLDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQL 191
L LDL +N++SG+IP LG+LK L L++N+LTG IP ++ SL +VS N L
Sbjct: 478 SLVSLDLYNNNISGTIPPSLGKLKNLVFLRLNDNRLTGPIPRELIAVTSLKVVDVSNNDL 657
Query: 192 CGAIPTG--LSKFAKSSFDHNKCLCGAPL 218
CG IPT ++F++N L G L
Sbjct: 658 CGTIPTSGPFEHIPLNNFENNPRLEGPEL 744
>TC83792 similar to GP|9280288|dbj|BAB01743.1 receptor protein kinase
{Arabidopsis thaliana}, partial (21%)
Length = 906
Score = 91.7 bits (226), Expect = 2e-19
Identities = 57/120 (47%), Positives = 67/120 (55%), Gaps = 2/120 (1%)
Frame = +2
Query: 104 DLPYLNFLSLAEMPKVTGPIPNSFSKLQRLQKLDLGSNSLSGSIPTFLGQLKGLQEFDLS 163
DL Y NFLS G IP F + LQ L+LG N L+G IP LG LK + DLS
Sbjct: 326 DLSY-NFLS--------GTIPEKFGAMAYLQVLNLGHNRLNGKIPESLGALKPIGVLDLS 478
Query: 164 NNKLTGVIPASFGSLPSLSQFNVSFNQLCGAIPTG--LSKFAKSSFDHNKCLCGAPLAAC 221
+N L G IP S SL LS F+VS N L G IP+G L+ F S + +N LCG PL C
Sbjct: 479 HNNLQGFIPGSLQSLSFLSDFDVSNNNLSGLIPSGGQLTTFPASRYQNNSNLCGVPLPTC 658
>TC92543 weakly similar to GP|6041804|gb|AAF02124.1| putative protein kinase
{Arabidopsis thaliana}, partial (10%)
Length = 685
Score = 90.5 bits (223), Expect = 5e-19
Identities = 50/114 (43%), Positives = 68/114 (58%)
Frame = +1
Query: 93 GLSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQRLQKLDLGSNSLSGSIPTFLG 152
GL G +P G + L FLSLA ++G IP S ++ LQ LDL +NSL+G IP F+
Sbjct: 37 GLQGQIPTSLGQMKVLKFLSLAGN-NLSGSIPTSLGQMYSLQVLDLSTNSLTGEIPKFIE 213
Query: 153 QLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQLCGAIPTGLSKFAKSS 206
++ L L+NN L+G IPA ++ +LS FNVSFN L G +P+ S SS
Sbjct: 214 NMRNLTNVLLNNNNLSGHIPAGLVNVTTLSAFNVSFNNLSGYLPSNSSLIKCSS 375
Score = 57.4 bits (137), Expect = 4e-09
Identities = 31/67 (46%), Positives = 39/67 (57%)
Frame = +1
Query: 130 LQRLQKLDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFN 189
L L L+L N L G IPT LGQ+K L+ L+ N L+G IP S G + SL ++S N
Sbjct: 1 LVSLVSLNLSRNGLQGQIPTSLGQMKVLKFLSLAGNNLSGSIPTSLGQMYSLQVLDLSTN 180
Query: 190 QLCGAIP 196
L G IP
Sbjct: 181 SLTGEIP 201
>TC86633 weakly similar to SP|Q00874|D100_ARATH DNA-damage-repair/toleration
protein DRT100 precursor. [Mouse-ear cress] {Arabidopsis
thaliana}, partial (93%)
Length = 1407
Score = 90.1 bits (222), Expect = 6e-19
Identities = 63/207 (30%), Positives = 102/207 (48%), Gaps = 13/207 (6%)
Frame = +3
Query: 13 LTLLFLSSIYFSTSNADDVEFSCNANDKATLLKIRDHFGGPN-GRLSDWDNGTDCCSDWS 71
LTLL L+ I ST SC +D+A LL + PN G + W +G DCC W
Sbjct: 45 LTLLLLTVI--STVK------SCPPSDRAALLAFKSALTEPNLGIFNSW-SGYDCCRGWH 197
Query: 72 FVGCGKPLPGRITVV------------TISRGWGLSGTLPAEFGDLPYLNFLSLAEMPKV 119
V C P R+T + ++ ++G + E L L L +A+ +
Sbjct: 198 GVSC-NPTTWRVTDINLRGDSEDPIFQNLTHSGDMTGEISPEVCKLDELTTLVVADWKSI 374
Query: 120 TGPIPNSFSKLQRLQKLDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSLP 179
+G IP+ + L L+ LDL N +SG+IP +G+L+ L +L++N ++G IP S +
Sbjct: 375 SGEIPSCITSLSSLRILDLTGNKISGNIPGNIGKLQHLTVLNLADNAISGEIPMSIVRIS 554
Query: 180 SLSQFNVSFNQLCGAIPTGLSKFAKSS 206
L +++ NQ+ G +P+ + K + S
Sbjct: 555 GLMHLDLAGNQISGELPSDIGKLRRLS 635
Score = 75.9 bits (185), Expect = 1e-14
Identities = 46/139 (33%), Positives = 75/139 (53%), Gaps = 5/139 (3%)
Frame = +3
Query: 68 SDWSFVGCGKPLPGRITVVTISR-----GWGLSGTLPAEFGDLPYLNFLSLAEMPKVTGP 122
+DW + +P IT ++ R G +SG +P G L +L L+LA+ ++G
Sbjct: 357 ADWKSIS--GEIPSCITSLSSLRILDLTGNKISGNIPGNIGKLQHLTVLNLADNA-ISGE 527
Query: 123 IPNSFSKLQRLQKLDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPSLS 182
IP S ++ L LDL N +SG +P+ +G+L+ L S N+LTG IP S + L+
Sbjct: 528 IPMSIVRISGLMHLDLAGNQISGELPSDIGKLRRLSRALFSRNQLTGSIPDSVLKMNRLA 707
Query: 183 QFNVSFNQLCGAIPTGLSK 201
++S N++ G+IP + K
Sbjct: 708 DLDLSMNRITGSIPARIGK 764
Score = 73.6 bits (179), Expect = 6e-14
Identities = 55/178 (30%), Positives = 78/178 (42%), Gaps = 50/178 (28%)
Frame = +3
Query: 94 LSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQRLQKLDLGSNSLSGSIPTFLGQ 153
+SG LP++ G L L+ +L ++TG IP+S K+ RL LDL N ++GSIP +G+
Sbjct: 588 ISGELPSDIGKLRRLS-RALFSRNQLTGSIPDSVLKMNRLADLDLSMNRITGSIPARIGK 764
Query: 154 LKGLQEFDLSNNKLTGVIPAS------------------------FGSLPSLSQFNVSFN 189
++ L L N +TG IP++ FGS ++SFN
Sbjct: 765 MRVLSTLKLDGNSMTGQIPSTLLSNTGMGILNLSRNGFEGTIPDVFGSKSYFMVLDLSFN 944
Query: 190 QLCGAIPTGLS--KFA------------------------KSSFDHNKCLCGAPLAAC 221
+L G IP LS KF +SF +N C CG PL C
Sbjct: 945 KLTGRIPGSLSSAKFMGHLDISNNHLCGTIPIGSPFDHLDAASFSNNDCSCGNPLKTC 1118
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.320 0.138 0.429
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,757,819
Number of Sequences: 36976
Number of extensions: 116338
Number of successful extensions: 1288
Number of sequences better than 10.0: 247
Number of HSP's better than 10.0 without gapping: 874
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1082
length of query: 221
length of database: 9,014,727
effective HSP length: 92
effective length of query: 129
effective length of database: 5,612,935
effective search space: 724068615
effective search space used: 724068615
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 56 (26.2 bits)
Medicago: description of AC146750.10


