

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146747.7 - phase: 0 /pseudo
(305 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
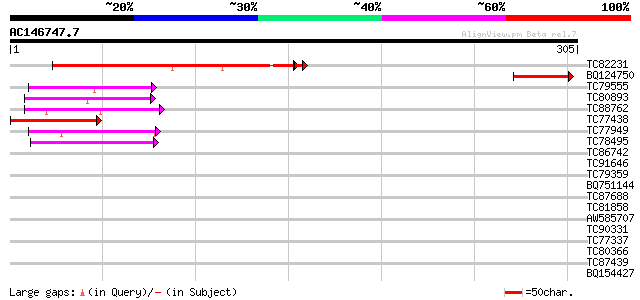
Score E
Sequences producing significant alignments: (bits) Value
TC82231 similar to GP|7658344|gb|AAF66134.1| unknown protein; 15... 177 6e-45
BQ124750 similar to PIR|G86446|G864 unknown protein [imported] -... 64 6e-11
TC79555 homologue to GP|10567859|gb|AAG18593.1 L8019.5 {Leishman... 46 2e-05
TC80893 similar to GP|4557063|gb|AAD22502.1| expressed protein {... 45 3e-05
TC88762 similar to GP|22136928|gb|AAM91808.1 unknown protein {Ar... 41 5e-04
TC77438 homologue to PIR|T06807|T06807 nucleosome assembly prote... 41 7e-04
TC77949 weakly similar to GP|11762218|gb|AAG40387.1 AT4g27520 {A... 40 9e-04
TC78495 homologue to PIR|S53504|S53504 extensin-like protein S3 ... 40 9e-04
TC86742 similar to PIR|T01541|T01541 hypothetical protein A_IG00... 40 0.001
TC91646 weakly similar to GP|6523547|emb|CAB62280.1 hydroxyproli... 39 0.002
TC79359 weakly similar to PIR|S49915|S49915 extensin-like protei... 39 0.002
BQ751144 weakly similar to PIR|PQ0479|PQ04 pistil extensin-like ... 39 0.003
TC87688 similar to GP|10177535|dbj|BAB10930. gene_id:K1F13.21~un... 39 0.003
TC81858 similar to GP|18252179|gb|AAL61922.1 unknown protein {Ar... 39 0.003
AW585707 weakly similar to GP|13605599|gb| AT4g32910/F26P21_30 {... 39 0.003
TC90331 weakly similar to GP|21751020|dbj|BAC03887. unnamed prot... 39 0.003
TC77337 weakly similar to GP|4019275|gb|AAC95573.1| orf 48 {Atel... 38 0.004
TC80366 similar to GP|13877617|gb|AAK43886.1 protein kinase-like... 38 0.004
TC87439 weakly similar to GP|5139695|dbj|BAA81686.1 expressed in... 38 0.004
BQ154427 weakly similar to GP|13277220|emb ZF-HD homeobox protei... 35 0.047
>TC82231 similar to GP|7658344|gb|AAF66134.1| unknown protein; 15741-13972
{Arabidopsis thaliana}, partial (48%)
Length = 494
Score = 177 bits (449), Expect(2) = 6e-45
Identities = 106/144 (73%), Positives = 108/144 (74%), Gaps = 12/144 (8%)
Frame = +2
Query: 24 APSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPNSKKLRTPSNTPPALK 83
APSRSQSPT TSS PAAQAAS PSPT PP M HVSPSWHDP NSKKL+T S TPP K
Sbjct: 14 APSRSQSPTATSSLPAAQAASVLPSPTAPPPMVHVSPSWHDPFKNSKKLKTLSKTPPV*K 193
Query: 84 *GY-------LMR*RKWLMKLDPLMCCC*IMECFMRW-----N*VMSSLRLMLI*WVV*I 131
* Y +M *RK LMKLDPLMCC *IM CFMRW N*VM SLRLML *WVV*I
Sbjct: 194 *RYSQRM*EIMML*RKRLMKLDPLMCC**IMVCFMRWNWRKRN*VM*SLRLMLT*WVV*I 373
Query: 132 *LKLLCLI*RRIGKILYLLLLLCF 155
*LKLLC * RIGKILYLLLLL F
Sbjct: 374 *LKLLCRK*-RIGKILYLLLLLLF 442
Score = 21.2 bits (43), Expect(2) = 6e-45
Identities = 7/7 (100%), Positives = 7/7 (100%)
Frame = +3
Query: 154 CFITSWS 160
CFITSWS
Sbjct: 438 CFITSWS 458
>BQ124750 similar to PIR|G86446|G864 unknown protein [imported] - Arabidopsis
thaliana, partial (13%)
Length = 658
Score = 64.3 bits (155), Expect = 6e-11
Identities = 26/32 (81%), Positives = 28/32 (87%)
Frame = +3
Query: 272 AAGIMRIAALCFQWNWYGSIEKWHKQRKCKCL 303
AAGIM I ALC QWNWYGSIEKWHKQRKC+ +
Sbjct: 3 AAGIMCIVALCLQWNWYGSIEKWHKQRKCQLM 98
>TC79555 homologue to GP|10567859|gb|AAG18593.1 L8019.5 {Leishmania major},
partial (1%)
Length = 1410
Score = 46.2 bits (108), Expect = 2e-05
Identities = 30/75 (40%), Positives = 40/75 (53%), Gaps = 6/75 (8%)
Frame = +2
Query: 11 SSSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAAS-----ASPSPTVPPLMAHVSPSWHDP 65
+SS PS+T S+P + P T+SSPA A S ASPS +VPP S S P
Sbjct: 1010 ASSGPSNTCPSSPRRRSGRLPAATTSSPATNANSPPFPPASPSSSVPPAKPPTSKSARSP 1189
Query: 66 SPN-SKKLRTPSNTP 79
+P S+ PS++P
Sbjct: 1190 APV*SRPSPPPSSSP 1234
Score = 39.3 bits (90), Expect = 0.002
Identities = 28/71 (39%), Positives = 34/71 (47%), Gaps = 2/71 (2%)
Frame = +2
Query: 12 SSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPNSKK 71
S PSST SS PSR SP T+ P ++ PSPT P + S PS +S
Sbjct: 479 SPQPSSTRSSPAPPSRPSSPATTTRPP----PTSKPSPTQTP--SRSPSSARTPSTSSTP 640
Query: 72 LRTP--SNTPP 80
R P S+ PP
Sbjct: 641 TRAPLLSSPPP 673
Score = 33.5 bits (75), Expect = 0.10
Identities = 26/79 (32%), Positives = 35/79 (43%), Gaps = 1/79 (1%)
Frame = +2
Query: 1 MADAFFIFSLSSSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTV-PPLMAHVS 59
MA + SP T P S P+ T SSPA + +SP+ T PP + S
Sbjct: 410 MASYLLVLHHHQPSPKPTL----VPP*SPQPSSTRSSPAPPSRPSSPATTTRPPPTSKPS 577
Query: 60 PSWHDPSPNSKKLRTPSNT 78
P+ PS + RTPS +
Sbjct: 578 PT-QTPSRSPSSARTPSTS 631
>TC80893 similar to GP|4557063|gb|AAD22502.1| expressed protein {Arabidopsis
thaliana}, partial (39%)
Length = 883
Score = 45.1 bits (105), Expect = 3e-05
Identities = 31/72 (43%), Positives = 43/72 (59%), Gaps = 2/72 (2%)
Frame = -1
Query: 9 SLSSSSPSSTSSSAPAPSRSQSPTVTSSSPAA--QAASASPSPTVPPLMAHVSPSWHDPS 66
S SSSS SS+SSS+P+ S S SP+ +SSSP++ A AS S + P + SP S
Sbjct: 559 SSSSSSSSSSSSSSPSSSSSSSPSSSSSSPSSLPPAPLASGSSSGSPSSSPSSPE*SSSS 380
Query: 67 PNSKKLRTPSNT 78
P+S T S++
Sbjct: 379 PSSSSSCTSSSS 344
Score = 28.5 bits (62), Expect = 3.4
Identities = 18/46 (39%), Positives = 26/46 (56%)
Frame = -1
Query: 11 SSSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMA 56
S SS S SSS +P S S +SSS + ++S+S S +V L +
Sbjct: 442 SGSSSGSPSSSPSSPE*SSSSPSSSSSCTSSSSSSSSSSSVSLLFS 305
>TC88762 similar to GP|22136928|gb|AAM91808.1 unknown protein {Arabidopsis
thaliana}, partial (3%)
Length = 805
Score = 41.2 bits (95), Expect = 5e-04
Identities = 27/87 (31%), Positives = 45/87 (51%), Gaps = 12/87 (13%)
Frame = +3
Query: 9 SLSSSSPSST--------SSSAPAPSRSQSPTVTSSSPAAQAASASP----SPTVPPLMA 56
S ++S P+ST ++S PA + + SP T++SP A ++SP SPT P +
Sbjct: 126 STTTSPPASTPASSPVSTTTSPPASTPASSPVPTTTSPPAPTPASSPVSTNSPTASPAGS 305
Query: 57 HVSPSWHDPSPNSKKLRTPSNTPPALK 83
+ + PS + +PS+TPP +
Sbjct: 306 LPAAATPSPSSTTAGSPSPSSTPPGTR 386
Score = 32.3 bits (72), Expect = 0.23
Identities = 19/66 (28%), Positives = 32/66 (47%), Gaps = 2/66 (3%)
Frame = +3
Query: 17 STSSSAPAPSRSQSPTVTSSSPAAQA--ASASPSPTVPPLMAHVSPSWHDPSPNSKKLRT 74
S + AP+P + +PT SSSP+A + ++ + P P + VS + P+
Sbjct: 42 SGNGIAPSPYQFNNPTSPSSSPSASSPVSTTTSPPASTPASSPVSTTTSPPASTPASSPV 221
Query: 75 PSNTPP 80
P+ T P
Sbjct: 222 PTTTSP 239
Score = 28.5 bits (62), Expect = 3.4
Identities = 16/42 (38%), Positives = 26/42 (61%), Gaps = 4/42 (9%)
Frame = +3
Query: 11 SSSSPSSTSSSAPAPSR----SQSPTVTSSSPAAQAASASPS 48
++ SPSST++ +P+PS ++S T+ + ASASPS
Sbjct: 318 ATPSPSSTTAGSPSPSSTPPGTRSSNETTPGRSIGVASASPS 443
Score = 27.3 bits (59), Expect = 7.5
Identities = 22/71 (30%), Positives = 29/71 (39%)
Frame = +3
Query: 9 SLSSSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPN 68
SL + S + APS Q TS S + A+S + T PP S SP
Sbjct: 12 SLPPGVAQTPSGNGIAPSPYQFNNPTSPSSSPSASSPVSTTTSPPASTPAS------SPV 173
Query: 69 SKKLRTPSNTP 79
S P++TP
Sbjct: 174STTTSPPASTP 206
>TC77438 homologue to PIR|T06807|T06807 nucleosome assembly protein 1 - garden
pea, partial (93%)
Length = 1607
Score = 40.8 bits (94), Expect = 7e-04
Identities = 22/49 (44%), Positives = 33/49 (66%)
Frame = -3
Query: 1 MADAFFIFSLSSSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSP 49
+AD FF+ +SSSS SS+SSS+ + S S S +SSS ++ + S + SP
Sbjct: 1161 LADDFFLVLVSSSSSSSSSSSSSSSSSSSSSASSSSSSSSSSRSPNSSP 1015
>TC77949 weakly similar to GP|11762218|gb|AAG40387.1 AT4g27520 {Arabidopsis
thaliana}, partial (34%)
Length = 1297
Score = 40.4 bits (93), Expect = 9e-04
Identities = 28/77 (36%), Positives = 38/77 (48%), Gaps = 6/77 (7%)
Frame = +3
Query: 11 SSSSPSSTSSSAPAPS---RSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSP 67
SS SPS+ S P+PS + SP+ + SP A ++SP PT P + SP P
Sbjct: 504 SSPSPSAGGLSPPSPSPTTTTPSPSGSPPSPVAIPPASSPVPTSGPTASSPSPVVSTPPA 683
Query: 68 NSKKLRTPS---NTPPA 81
+PS +TPPA
Sbjct: 684 GGPMASSPSPVVSTPPA 734
Score = 28.1 bits (61), Expect = 4.4
Identities = 16/38 (42%), Positives = 18/38 (47%)
Frame = +3
Query: 30 SPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSP 67
SP SSP+ A SP P+ P SPS PSP
Sbjct: 486 SPRTPKSSPSPSAGGLSP-PSPSPTTTTPSPSGSPPSP 596
>TC78495 homologue to PIR|S53504|S53504 extensin-like protein S3 - alfalfa,
partial (97%)
Length = 1049
Score = 40.4 bits (93), Expect = 9e-04
Identities = 20/69 (28%), Positives = 32/69 (45%)
Frame = +3
Query: 12 SSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPNSKK 71
+++P +T ++P P +S P V SS P Q++ T PP+ + P P P
Sbjct: 288 ANTPPTTPQASPPPVQSSPPPVQSSPPPLQSSPPPAQSTPPPVQSSPPPVQQSPPPTPLT 467
Query: 72 LRTPSNTPP 80
+TPP
Sbjct: 468 PPPVQSTPP 494
Score = 38.9 bits (89), Expect = 0.002
Identities = 25/68 (36%), Positives = 30/68 (43%)
Frame = +3
Query: 13 SSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPNSKKL 72
SSP SS P P++S P V SS P Q S P+P PP + P P +
Sbjct: 357 SSPPPLQSSPP-PAQSTPPPVQSSPPPVQ-QSPPPTPLTPPPVQSTPPPASPPPASPPPF 530
Query: 73 RTPSNTPP 80
P TPP
Sbjct: 531 SPPPATPP 554
Score = 32.7 bits (73), Expect = 0.18
Identities = 21/70 (30%), Positives = 31/70 (44%), Gaps = 1/70 (1%)
Frame = +3
Query: 12 SSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASP-SPTVPPLMAHVSPSWHDPSPNSK 70
S+SP+S S + P P + PT +SP +S P + PPL + P+ P P
Sbjct: 246 STSPNS-SPAPPTPPANTPPTTPQASPPPVQSSPPPVQSSPPPLQSSPPPAQSTPPPVQS 422
Query: 71 KLRTPSNTPP 80
+PP
Sbjct: 423 SPPPVQQSPP 452
Score = 31.6 bits (70), Expect = 0.40
Identities = 19/73 (26%), Positives = 32/73 (43%)
Frame = +3
Query: 11 SSSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPNSK 70
+ +PS++ +S+PAP PT P A +P + PP+ + P P P
Sbjct: 231 AQQAPSTSPNSSPAP-----PT-----PPANTPPTTPQASPPPVQSSPPPVQSSPPPLQS 380
Query: 71 KLRTPSNTPPALK 83
+TPP ++
Sbjct: 381 SPPPAQSTPPPVQ 419
Score = 30.8 bits (68), Expect = 0.68
Identities = 26/84 (30%), Positives = 33/84 (38%), Gaps = 12/84 (14%)
Frame = +3
Query: 10 LSSSSPSSTSSSAPA----PSRSQSPTVT--------SSSPAAQAASASPSPTVPPLMAH 57
L SS P + S+ P P QSP T S+ P A ASP P PP
Sbjct: 372 LQSSPPPAQSTPPPVQSSPPPVQQSPPPTPLTPPPVQSTPPPASPPPASPPPFSPPPATP 551
Query: 58 VSPSWHDPSPNSKKLRTPSNTPPA 81
+ +P TP ++PPA
Sbjct: 552 PPATPPPATPPPALTPTPLSSPPA 623
Score = 27.3 bits (59), Expect = 7.5
Identities = 16/62 (25%), Positives = 31/62 (49%)
Frame = +3
Query: 9 SLSSSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPN 68
+L+ + SS ++ PAP+ ++ + + + S +P+P + L +SPS D S
Sbjct: 588 ALTPTPLSSPPATTPAPAPAKLKSKAPALAPVLSPSDAPAPGLSSLSPSISPSGTDDSGA 767
Query: 69 SK 70
K
Sbjct: 768 EK 773
>TC86742 similar to PIR|T01541|T01541 hypothetical protein A_IG005I10.16 -
Arabidopsis thaliana, partial (19%)
Length = 2073
Score = 39.7 bits (91), Expect = 0.001
Identities = 32/79 (40%), Positives = 39/79 (48%), Gaps = 1/79 (1%)
Frame = +3
Query: 9 SLSSSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTV-PPLMAHVSPSWHDPSP 67
SLSSSS S S +P P+ S T TSS+PA S PS TV PL + P PSP
Sbjct: 690 SLSSSSSPSPPSPSP-PTSSHPSTTTSSTPA--PPSPPPSTTVFSPLNTLLFPPPPIPSP 860
Query: 68 NSKKLRTPSNTPPALK*GY 86
+ K P + P G+
Sbjct: 861 SQTKTHFPIHLPSTTPLGF 917
>TC91646 weakly similar to GP|6523547|emb|CAB62280.1 hydroxyproline-rich
glycoprotein DZ-HRGP {Volvox carteri f. nagariensis},
partial (15%)
Length = 984
Score = 38.9 bits (89), Expect = 0.002
Identities = 25/74 (33%), Positives = 34/74 (45%), Gaps = 5/74 (6%)
Frame = +1
Query: 12 SSSPSSTSS----SAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDP-S 66
++SPS T+S S P P P S P S SP P PP ++ +P+ P S
Sbjct: 331 TTSPSPTASPPPRSPPRPPPRSPPRQLMSPPRPLPRSPSPPPRNPPELSRTTPTPRPPMS 510
Query: 67 PNSKKLRTPSNTPP 80
P+ P+ TPP
Sbjct: 511 PSPSPTSPPAATPP 552
Score = 33.9 bits (76), Expect = 0.080
Identities = 23/70 (32%), Positives = 32/70 (44%), Gaps = 1/70 (1%)
Frame = +1
Query: 14 SPSSTSSSAPAPSRSQSPTVTSSSPAAQAA-SASPSPTVPPLMAHVSPSWHDPSPNSKKL 72
SP +P+P P ++ ++P + S SPSPT PP A S P P ++L
Sbjct: 415 SPPRPLPRSPSPPPRNPPELSRTTPTPRPPMSPSPSPTSPPA-ATPPVSRPPPRPALRRL 591
Query: 73 RTPSNTPPAL 82
P PP L
Sbjct: 592 LPPPPLPPTL 621
Score = 27.7 bits (60), Expect = 5.7
Identities = 24/70 (34%), Positives = 30/70 (42%), Gaps = 2/70 (2%)
Frame = +1
Query: 15 PSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPP--LMAHVSPSWHDPSPNSKKL 72
PS + P PS + SP+ T+S P + P P PP LM+ P PSP
Sbjct: 295 PSRLPLTTP-PSLTTSPSPTASPPPR--SPPRPPPRSPPRQLMSPPRPLPRSPSP----- 450
Query: 73 RTPSNTPPAL 82
P PP L
Sbjct: 451 --PPRNPPEL 474
Score = 26.9 bits (58), Expect = 9.8
Identities = 22/70 (31%), Positives = 27/70 (38%), Gaps = 12/70 (17%)
Frame = +1
Query: 22 APAPSRSQSP------------TVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPNS 69
AP+P S SP T S + + +ASP P PP SP SP
Sbjct: 247 APSPRPSPSPLSPLRFPSRLPLTTPPSLTTSPSPTASPPPRSPPRPPPRSPPRQLMSPPR 426
Query: 70 KKLRTPSNTP 79
R+PS P
Sbjct: 427 PLPRSPSPPP 456
>TC79359 weakly similar to PIR|S49915|S49915 extensin-like protein - maize,
partial (3%)
Length = 1170
Score = 38.9 bits (89), Expect = 0.002
Identities = 26/80 (32%), Positives = 38/80 (47%), Gaps = 3/80 (3%)
Frame = +3
Query: 6 FIFSLSSSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPT---VPPLMAHVSPSW 62
F + +SSP S+ ++ P S S SPT + +SPA + PS T P A P
Sbjct: 213 FAQNAPASSPKSSVTAKPPSSVSVSPTNSPASPAKSPTLSPPSQTPAVSPSGSASTPPPA 392
Query: 63 HDPSPNSKKLRTPSNTPPAL 82
P S ++ PS+ PA+
Sbjct: 393 TSPPAKSPAVQPPSSVSPAI 452
Score = 28.5 bits (62), Expect = 3.4
Identities = 23/84 (27%), Positives = 39/84 (46%), Gaps = 19/84 (22%)
Frame = +3
Query: 14 SPSSTSSSAP------------------APSRSQSPTVTSSSPAAQAASASPSPTVPPLM 55
SPS ++S+ P +P+ S S V+S+ P + AS+ P+ V P+
Sbjct: 357 SPSGSASTPPPATSPPAKSPAVQPPSSVSPAISPSNNVSSTPPVSSPASSPPTAAVSPVS 536
Query: 56 AHV-SPSWHDPSPNSKKLRTPSNT 78
+ V +PS P P + PS++
Sbjct: 537 SPVEAPSVSSP-PEASSAGIPSSS 605
>BQ751144 weakly similar to PIR|PQ0479|PQ04 pistil extensin-like protein
(clone pMG14) - common tobacco (fragment), partial (11%)
Length = 632
Score = 38.5 bits (88), Expect = 0.003
Identities = 25/71 (35%), Positives = 37/71 (51%), Gaps = 1/71 (1%)
Frame = +1
Query: 13 SSPSSTSSSAPAP-SRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPNSKK 71
S PSS S++P+P + +SPT ++P + AS+ SPT P + P PSP+ +
Sbjct: 181 SLPSSPPSTSPSPHALLRSPTAPPAAPLSPPASSPSSPTRPSSLFSPPP----PSPSRSR 348
Query: 72 LRTPSNTPPAL 82
PS P L
Sbjct: 349 PPPPSTPSPLL 381
Score = 30.8 bits (68), Expect = 0.68
Identities = 25/78 (32%), Positives = 39/78 (49%), Gaps = 8/78 (10%)
Frame = +1
Query: 13 SSPSSTSSSAPA-PSRSQ---SPTVTSSSPAAQAASASPSPTVP----PLMAHVSPSWHD 64
++P S +S+P+ P+R SP S S + ++PSP +P PL SP +
Sbjct: 253 AAPLSPPASSPSSPTRPSSLFSPPPPSPSRSRPPPPSTPSPLLPLPLLPLATEPSP-FPP 429
Query: 65 PSPNSKKLRTPSNTPPAL 82
SP + +PS +PP L
Sbjct: 430 SSPPPRPASSPSLSPPVL 483
Score = 30.4 bits (67), Expect = 0.89
Identities = 24/71 (33%), Positives = 29/71 (40%)
Frame = +1
Query: 12 SSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPNSKK 71
S P S+ PA S S SP V P+A SPS L PS P+S +
Sbjct: 415 SPFPPSSPPPRPASSPSLSPPVLLFPPSASPPPWSPSRPPSALAPPSRPSLPASPPSSPR 594
Query: 72 LRTPSNTPPAL 82
T + PAL
Sbjct: 595 SSTRPSRFPAL 627
>TC87688 similar to GP|10177535|dbj|BAB10930. gene_id:K1F13.21~unknown
protein {Arabidopsis thaliana}, partial (46%)
Length = 2077
Score = 38.5 bits (88), Expect = 0.003
Identities = 29/80 (36%), Positives = 47/80 (58%), Gaps = 1/80 (1%)
Frame = -2
Query: 4 AFFIFSLSS-SSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSW 62
++ F +SS SS SS+SSS+ + S S SP +SSS ++ ++SAS ++ + +S S+
Sbjct: 786 SYLTFPISSASSSSSSSSSSSSSSISNSPAFSSSSSSSLSSSAS---SLSSPFS*LSSSF 616
Query: 63 HDPSPNSKKLRTPSNTPPAL 82
S NS + S+ PP L
Sbjct: 615 SKYSVNSSIFKNLSSIPPPL 556
Score = 27.7 bits (60), Expect = 5.7
Identities = 20/50 (40%), Positives = 31/50 (62%), Gaps = 7/50 (14%)
Frame = -2
Query: 6 FIFSLSSSSPS--STSSSAPAPSRSQSPTVTS-----SSPAAQAASASPS 48
F FS SSSS S +SSS+ + S+S SP++ S+P+ ++S+S S
Sbjct: 552 FSFSSSSSSSSSLPSSSSSSSSSKSSSPSLPPFAFFFSTPSKSSSSSSNS 403
Score = 26.9 bits (58), Expect = 9.8
Identities = 18/47 (38%), Positives = 26/47 (55%)
Frame = -2
Query: 7 IFSLSSSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPP 53
IF SS P + S + S S S +SSS ++ + S+SPS +PP
Sbjct: 591 IFKNLSSIPPPLTFSFSSSSSSSSSLPSSSSSSSSSKSSSPS--LPP 457
>TC81858 similar to GP|18252179|gb|AAL61922.1 unknown protein {Arabidopsis
thaliana}, partial (9%)
Length = 748
Score = 38.5 bits (88), Expect = 0.003
Identities = 24/89 (26%), Positives = 47/89 (51%), Gaps = 12/89 (13%)
Frame = +1
Query: 5 FFIFSL---------SSSSPSST---SSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVP 52
FF+F+L S+SP++ S +APAP + +P ++++P +A++ S P
Sbjct: 97 FFVFALLATTAFGEAPSTSPTAAPKASHAAPAPKATATPPSSTTTPPKSSATSPTSSPAP 276
Query: 53 PLMAHVSPSWHDPSPNSKKLRTPSNTPPA 81
+ + SP+ P+ + +P+ +PPA
Sbjct: 277 KVSSPPSPT---PTSAEAPVESPTESPPA 354
Score = 37.4 bits (85), Expect = 0.007
Identities = 23/71 (32%), Positives = 33/71 (46%), Gaps = 1/71 (1%)
Frame = +1
Query: 12 SSSPSSTSSSAPAPSRSQSPTVTSS-SPAAQAASASPSPTVPPLMAHVSPSWHDPSPNSK 70
SS+ S TSS AP S SPT TS+ +P + P+P P + SP P+ +
Sbjct: 241 SSATSPTSSPAPKVSSPPSPTPTSAEAPVESPTESPPAPVSPTVSPATSPVASGPAVSDA 420
Query: 71 KLRTPSNTPPA 81
P+ + A
Sbjct: 421 PAEAPAGSSAA 453
>AW585707 weakly similar to GP|13605599|gb| AT4g32910/F26P21_30 {Arabidopsis
thaliana}, partial (6%)
Length = 452
Score = 38.5 bits (88), Expect = 0.003
Identities = 30/85 (35%), Positives = 39/85 (45%)
Frame = +2
Query: 11 SSSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPNSK 70
S++ P S S + P SQS T+ SP A SP P P VSPS SPN
Sbjct: 116 STTEP*SHSPAKPTTILSQSTLSTTLSPVQFLALPSPGPAATP---SVSPS----SPNLL 274
Query: 71 KLRTPSNTPPALK*GYLMR*RKWLM 95
K P P +LK ++ R+ +M
Sbjct: 275 KPPMPETVPKSLKLSSVVEIRRLMM 349
>TC90331 weakly similar to GP|21751020|dbj|BAC03887. unnamed protein product
{Homo sapiens}, partial (25%)
Length = 557
Score = 38.5 bits (88), Expect = 0.003
Identities = 25/70 (35%), Positives = 38/70 (53%)
Frame = +1
Query: 9 SLSSSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPN 68
S SS+S SSTSSS+ +PS SP+ S++P+ + +S T P S S SP+
Sbjct: 340 SSSSTSSSSTSSSSSSPSSFSSPSSMSAAPSLRGSSCPTG*TRYPAPKSSSSSSSSNSPS 519
Query: 69 SKKLRTPSNT 78
S + S++
Sbjct: 520 SSSPSSSSSS 549
Score = 32.7 bits (73), Expect = 0.18
Identities = 29/93 (31%), Positives = 39/93 (41%), Gaps = 19/93 (20%)
Frame = +1
Query: 6 FIFSLSSSSPSSTSSS-----------------APAPSRSQSPTVTSSSPAAQAASASPS 48
F+ + SSSS SS SSS A + S S S T +SSS + +S S
Sbjct: 238 FVNAASSSSSSSASSSLSLFLLGCEEAGGNTS*ASSSSTSSSSTSSSSSSPSSFSSPSSM 417
Query: 49 PTVPPLMAHVSPS--WHDPSPNSKKLRTPSNTP 79
P L P+ P+P S + SN+P
Sbjct: 418 SAAPSLRGSSCPTG*TRYPAPKSSSSSSSSNSP 516
>TC77337 weakly similar to GP|4019275|gb|AAC95573.1| orf 48 {Ateline
herpesvirus 3}, partial (8%)
Length = 986
Score = 38.1 bits (87), Expect = 0.004
Identities = 29/72 (40%), Positives = 41/72 (56%), Gaps = 7/72 (9%)
Frame = -2
Query: 5 FFIF------SLSSSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHV 58
FF+F S SSSS SS+ SS+ PS S S +SSS ++ ++S+S S + P +
Sbjct: 775 FFLFFGGCKASSSSSSSSSSLSSSSTPSSSSSSPPSSSSSSSSSSSSSSSWSSPSSSSSS 596
Query: 59 SP-SWHDPSPNS 69
SP S P+P S
Sbjct: 595 SPFSSAPPAPPS 560
Score = 34.7 bits (78), Expect = 0.047
Identities = 28/81 (34%), Positives = 42/81 (51%), Gaps = 10/81 (12%)
Frame = -2
Query: 12 SSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPT------VPPLMAHVSPSWHD- 64
SSS SS+SSS+ + S S + +SSSP + A A PS + L+ + PS D
Sbjct: 670 SSSSSSSSSSSSSSSWSSPSSSSSSSPFSSAPPAPPSGAFFELLLLLELLPYPPPSSDDN 491
Query: 65 ---PSPNSKKLRTPSNTPPAL 82
PSPN+ + S++ +L
Sbjct: 490 SSSPSPNAPSPSSSSSSSSSL 428
Score = 34.7 bits (78), Expect = 0.047
Identities = 28/85 (32%), Positives = 40/85 (46%), Gaps = 12/85 (14%)
Frame = -2
Query: 9 SLSSSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASA------------SPSPTVPPLMA 56
S SSSS SS+SSS +PS S S + SS+P A + A P P+ +
Sbjct: 664 SSSSSSSSSSSSSWSSPSSSSSSSPFSSAPPAPPSGAFFELLLLLELLPYPPPSSDDNSS 485
Query: 57 HVSPSWHDPSPNSKKLRTPSNTPPA 81
SP+ PS +S + ++P A
Sbjct: 484 SPSPNAPSPSSSSSSSSSLPSSPSA 410
>TC80366 similar to GP|13877617|gb|AAK43886.1 protein kinase-like protein
{Arabidopsis thaliana}, partial (16%)
Length = 814
Score = 38.1 bits (87), Expect = 0.004
Identities = 25/69 (36%), Positives = 32/69 (46%), Gaps = 1/69 (1%)
Frame = +3
Query: 14 SPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASP-SPTVPPLMAHVSPSWHDPSPNSKKL 72
+PS+ S PA S P+ T S+P S SP SP P ++ S PSP
Sbjct: 69 TPSAPPPSTPATPSSPPPSTTPSAPPPSTPSNSPPSPPTTPAISPPSGGGTTPSP----- 233
Query: 73 RTPSNTPPA 81
PS TPP+
Sbjct: 234 --PSRTPPS 254
Score = 36.2 bits (82), Expect = 0.016
Identities = 25/73 (34%), Positives = 37/73 (50%), Gaps = 4/73 (5%)
Frame = +3
Query: 13 SSPSSTSSSAPAPSRSQSP---TVTSSSPAAQAASASPSP-TVPPLMAHVSPSWHDPSPN 68
SSP + S+P PS +P T ++ P+ A +SP P T P +PS PSP
Sbjct: 3 SSPPPATPSSPPPSTPANPPPVTPSAPPPSTPATPSSPPPSTTPSAPPPSTPSNSPPSPP 182
Query: 69 SKKLRTPSNTPPA 81
+ TP+ +PP+
Sbjct: 183T----TPAISPPS 209
>TC87439 weakly similar to GP|5139695|dbj|BAA81686.1 expressed in cucumber
hypocotyls {Cucumis sativus}, partial (57%)
Length = 897
Score = 38.1 bits (87), Expect = 0.004
Identities = 23/59 (38%), Positives = 29/59 (48%), Gaps = 3/59 (5%)
Frame = +3
Query: 26 SRSQSPTVTSSSPAAQAASASP---SPTVPPLMAHVSPSWHDPSPNSKKLRTPSNTPPA 81
S S +PT + S+PAA +SP PT P A V+ P S K P+ TPPA
Sbjct: 105 SPSAAPTTSPSTPAATTPVSSPVAAPPTTPTTPAPVASPKSSPPATSPKAAAPTATPPA 281
Score = 29.3 bits (64), Expect = 2.0
Identities = 22/74 (29%), Positives = 34/74 (45%), Gaps = 2/74 (2%)
Frame = +3
Query: 11 SSSSPSST--SSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPN 68
+SS P+ T S+ PAP +SP + + A + +PT P + +PS N
Sbjct: 282 ASSPPAVTPVSTPPPAPVPVKSPPTPAPVSSPPAVTPVAAPTTTPAVPAPAPS--KGKKN 455
Query: 69 SKKLRTPSNTPPAL 82
KK P+ +P L
Sbjct: 456 KKKHGAPAPSPALL 497
Score = 28.5 bits (62), Expect = 3.4
Identities = 16/68 (23%), Positives = 31/68 (45%)
Frame = +3
Query: 12 SSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPNSKK 71
+S SS +++P + + +SSP A ++P P P+ + +P+ P
Sbjct: 213 ASPKSSPPATSPKAAAPTATPPAASSPPAVTPVSTPPPAPVPVKSPPTPAPVSSPPAVTP 392
Query: 72 LRTPSNTP 79
+ P+ TP
Sbjct: 393 VAAPTTTP 416
>BQ154427 weakly similar to GP|13277220|emb ZF-HD homeobox protein
{Flaveria bidentis}, partial (28%)
Length = 645
Score = 34.7 bits (78), Expect = 0.047
Identities = 23/72 (31%), Positives = 33/72 (44%), Gaps = 6/72 (8%)
Frame = +2
Query: 15 PSSTSSSAPAPSR---SQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPNSKK 71
P P P R SQS + + S+ + + S SP+ PP ++H+ PS+H S
Sbjct: 23 PPQPPPPLPLPHRGPISQSTSPSQSTSPSHSPSPIXSPSSPPPLSHLPPSYHHSSAPHML 202
Query: 72 LRTPSN---TPP 80
L N TPP
Sbjct: 203LALGGNAYSTPP 238
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.337 0.143 0.471
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,900,649
Number of Sequences: 36976
Number of extensions: 204823
Number of successful extensions: 6385
Number of sequences better than 10.0: 487
Number of HSP's better than 10.0 without gapping: 4136
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 5366
length of query: 305
length of database: 9,014,727
effective HSP length: 96
effective length of query: 209
effective length of database: 5,465,031
effective search space: 1142191479
effective search space used: 1142191479
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.8 bits)
S2: 58 (26.9 bits)
Medicago: description of AC146747.7


