

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146747.6 + phase: 0
(498 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
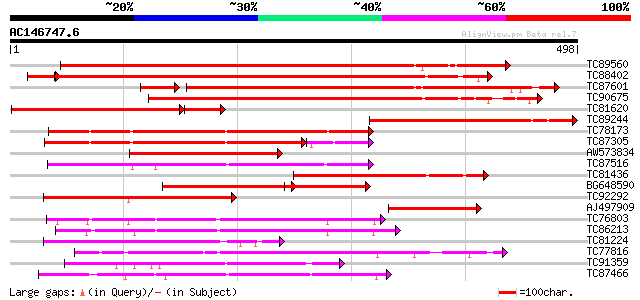
Score E
Sequences producing significant alignments: (bits) Value
TC89560 similar to GP|7671528|emb|CAB89490.1 CRK1 protein {Beta ... 594 e-170
TC88402 similar to GP|20804521|dbj|BAB92215. putative CRK1 prote... 488 e-143
TC87601 similar to GP|7671528|emb|CAB89490.1 CRK1 protein {Beta ... 426 e-130
TC90675 similar to GP|20804521|dbj|BAB92215. putative CRK1 prote... 366 e-101
TC81620 similar to GP|8953767|dbj|BAA98122.1 cyclin-dependent pr... 263 9e-84
TC89244 similar to GP|8953767|dbj|BAA98122.1 cyclin-dependent pr... 244 4e-65
TC78173 SP|P24923|CC21_MEDSA Cell division control protein 2 hom... 222 2e-58
TC87305 homologue to SP|Q05006|CC22_MEDSA Cell division control ... 199 1e-56
AW573834 similar to PIR|D86320|D86 hypothetical protein AAF27112... 210 1e-54
TC87516 PIR|T09591|T09591 probable cdc2-like protein kinase cdc2... 196 2e-50
TC81436 similar to PIR|A96571|A96571 hypothetical protein F8L10.... 196 2e-50
BG648590 similar to PIR|H96771|H96 hypothetical protein F1M20.1 ... 187 1e-47
TC92292 homologue to PIR|T09572|T09572 cdc2-like protein kinase ... 172 3e-43
AJ497909 similar to GP|8953767|dbj| cyclin-dependent protein kin... 169 3e-42
TC76803 PIR|T09622|T09622 protein kinase MMK4 (EC 2.7.1.-) cold... 165 3e-41
TC86213 homologue to PIR|S60121|S60121 mitogen-activated protein... 165 3e-41
TC81224 homologue to GP|17064846|gb|AAL32577.1 putative protein ... 162 2e-40
TC77816 similar to GP|14532760|gb|AAK64081.1 putative kinase {Ar... 162 4e-40
TC91359 homologue to PIR|T09586|T09586 probable cdc2-like protei... 161 6e-40
TC87466 homologue to GP|10185114|emb|CAC08564. wound-induced GSK... 159 3e-39
>TC89560 similar to GP|7671528|emb|CAB89490.1 CRK1 protein {Beta vulgaris},
partial (64%)
Length = 1266
Score = 594 bits (1531), Expect = e-170
Identities = 297/399 (74%), Positives = 338/399 (84%), Gaps = 3/399 (0%)
Frame = +2
Query: 45 TYSNVYKAKDLVTGKIVALKKVRFDNLEPESVKFMAREILVLRKLDHPNVVKLEGLVTSR 104
TYSNVYKA+D +TGK+VALKKVRFDNLEPESVKFMAREIL+LR+LDHPNVVKLEGLVTSR
Sbjct: 2 TYSNVYKARDTLTGKVVALKKVRFDNLEPESVKFMAREILILRRLDHPNVVKLEGLVTSR 181
Query: 105 MSCSLYLVFEYMEHDLAGLSAGQGVKFTEPQVKCFMKQLLSGLEHCHSRGVLHRDIKGSN 164
MSCSLYLVFEYM HDLAGL+ +KFTEPQVKC+M QL SGLEHCH+R VLHRDIKGSN
Sbjct: 182 MSCSLYLVFEYMAHDLAGLATNPAIKFTEPQVKCYMHQLFSGLEHCHNRHVLHRDIKGSN 361
Query: 165 LLIDNEGILKIADFGLATFYNPNKKQSMTSRVVTLWYRPPELLLGATFYGVGIDLWSAGC 224
LLIDN+G+LKIADFGLA+F++P+ K MTSRVVTLWYRPPELLLGAT YGVG+DLWSAGC
Sbjct: 362 LLIDNDGVLKIADFGLASFFDPDHKHPMTSRVVTLWYRPPELLLGATEYGVGVDLWSAGC 541
Query: 225 ILAELLAGKPIMPGRTEVEQLHKIFKLCGSPAEEYWRKHKLPNATIFKPQQPYKRCISET 284
ILAELLAGKPIMPGRTEVEQLHKIFKLCGSP+E+YW+K KLP+ATIFKPQQ YKRCI+ET
Sbjct: 542 ILAELLAGKPIMPGRTEVEQLHKIFKLCGSPSEDYWKKSKLPHATIFKPQQSYKRCIAET 721
Query: 285 FKDFPPSSLPLIDSLLAIDPDRRGTASAALNHEFFTTEPYACEPSSLPKYPPSKELDVKM 344
FK+FPPSSLPLI++LLAIDPD R TA+AAL+ EFFTT+PYAC+PSSLPKYPPSKE+D K+
Sbjct: 722 FKNFPPSSLPLIETLLAIDPDERLTATAALHSEFFTTKPYACDPSSLPKYPPSKEMDAKL 901
Query: 345 RDEEARRQKALNGKANA---VDGAKRVRARERGRAIPAPEANAEIQTNLDRWRVVTHANA 401
RDEEARR +A G++NA + ++E P N + Q +D ++THANA
Sbjct: 902 RDEEARRLRAA-GRSNA*WCQEITST*ASKEGDFPFQMPMLNCK-QILID-GGLITHANA 1072
Query: 402 KSKSEKFPPPHQDGAVGYPQDESSKGPVSFGASDTSFSS 440
KSKSEKFPPPHQDG +GYP S F ASD FSS
Sbjct: 1073KSKSEKFPPPHQDGDLGYPLGSSQHMDPVFDASDVPFSS 1189
>TC88402 similar to GP|20804521|dbj|BAB92215. putative CRK1 protein {Oryza
sativa (japonica cultivar-group)}, partial (46%)
Length = 2184
Score = 488 bits (1256), Expect(2) = e-143
Identities = 236/387 (60%), Positives = 299/387 (76%), Gaps = 2/387 (0%)
Frame = +1
Query: 40 KIGQGTYSNVYKAKDLVTGKIVALKKVRFDNLEPESVKFMAREILVLRKLDHPNVVKLEG 99
K+G+GTYS VYKA+D+ KIVALK+VRFDNL+PESVKFMAREI +LR+LDHPN++KLEG
Sbjct: 991 KLGKGTYSTVYKARDVTNQKIVALKRVRFDNLDPESVKFMAREIHILRRLDHPNIIKLEG 1170
Query: 100 LVTSRMSCSLYLVFEYMEHDLAGLSAGQGVKFTEPQVKCFMKQLLSGLEHCHSRGVLHRD 159
L+TS S SLYLVFEYMEHDL GL++ +KF+EPQ+KC+M QLLSGL+HCHS GVLHRD
Sbjct: 1171 LITSETSRSLYLVFEYMEHDLTGLASNPSIKFSEPQLKCYMHQLLSGLDHCHSHGVLHRD 1350
Query: 160 IKGSNLLIDNEGILKIADFGLATFYNPNKKQSMTSRVVTLWYRPPELLLGATFYGVGIDL 219
IKGSNLLIDN G+LKIADFGLA ++ + +TSRVVTLWYRPPELLLGA YGV +DL
Sbjct: 1351 IKGSNLLIDNNGVLKIADFGLANVFDAHLNIPLTSRVVTLWYRPPELLLGANHYGVAVDL 1530
Query: 220 WSAGCILAELLAGKPIMPGRTEVEQLHKIFKLCGSPAEEYWRKHKLPNATIFKPQQPYKR 279
WS GCIL EL G+PI+PG+TEVEQLH+IFKLCGSP+E+YW K +LP++T+FKP Y+R
Sbjct: 1531 WSTGCILRELYTGRPILPGKTEVEQLHRIFKLCGSPSEDYWLKLRLPHSTVFKPPHHYRR 1710
Query: 280 CISETFKDFPPSSLPLIDSLLAIDPDRRGTASAALNHEFFTTEPYACEPSSLPKYPPSKE 339
C+++TFK++ ++L LI++LL++DP RGTA+AAL EFFT+EP C+PSSLPKYPPSKE
Sbjct: 1711 CVADTFKEYSSTALKLIETLLSVDPSNRGTAAAALKSEFFTSEPLPCDPSSLPKYPPSKE 1890
Query: 340 LDVKMRDEEARRQKALNGKANAVDGAKRVRARERGRAIPAPEANAEIQTNLDRWRVVTHA 399
+D KMRDE RRQ A+ K A VR + RA + NA++ ++ + ++
Sbjct: 1891 IDAKMRDEATRRQGAVGDKEQRSGSA--VRQEKGPRAAVLTKDNADLGASIQQKH---YS 2055
Query: 400 NAKSKSEKFPP--PHQDGAVGYPQDES 424
K++SE P H G GYP +S
Sbjct: 2056 ITKNRSELSYPHREHVSGTQGYPHKQS 2136
Score = 39.7 bits (91), Expect(2) = e-143
Identities = 18/29 (62%), Positives = 22/29 (75%)
Frame = +3
Query: 16 LMAVAGEAIGDWTPRRANSFEKLAKIGQG 44
L + AG+AI W PR AN+FE+L KIGQG
Sbjct: 918 LSSGAGDAIKGWIPRSANTFERLHKIGQG 1004
>TC87601 similar to GP|7671528|emb|CAB89490.1 CRK1 protein {Beta vulgaris},
partial (50%)
Length = 1493
Score = 426 bits (1096), Expect(2) = e-130
Identities = 215/335 (64%), Positives = 260/335 (77%), Gaps = 7/335 (2%)
Frame = +2
Query: 156 LHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQSMTSRVVTLWYRPPELLLGATFYGV 215
LHRDIKGSNLLIDNEGILKIADFGLA+F++P ++ MT+RVVTLWYRPPELLLGAT YGV
Sbjct: 125 LHRDIKGSNLLIDNEGILKIADFGLASFFDPTRRHPMTNRVVTLWYRPPELLLGATDYGV 304
Query: 216 GIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKLCGSPAEEYWRKHKLPNATIFKPQQ 275
G+DLWSAGCIL ELL GKPIMPGRTEVEQLHKI+KLCGSP++EYW+K KLPNAT+FKP++
Sbjct: 305 GVDLWSAGCILGELLYGKPIMPGRTEVEQLHKIYKLCGSPSDEYWKKSKLPNATLFKPRE 484
Query: 276 PYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGTASAALNHEFFTTEPYACEPSSLPKYP 335
PYKRCI + FKDFPPS+LPL+D+LLAIDP R +AS AL EFF TEPYAC+PSSLPKYP
Sbjct: 485 PYKRCIRDVFKDFPPSALPLVDTLLAIDPIERRSASDALRSEFFNTEPYACDPSSLPKYP 664
Query: 336 PSKELDVKMRDEEARRQKALNGKANAVDGAKRVRARERG-RAIPAPEANAEIQTNLDRWR 394
P+KE+DVK RD+EARR +A KA A DG K+ R R+R +A PAPEANAE+Q N+DR R
Sbjct: 665 PTKEMDVKRRDDEARRSRAA-AKARA-DGGKKHRTRDRAVKAAPAPEANAELQYNIDRRR 838
Query: 395 VVTHANAKSKSEKFPPPHQDGAVGYPQDESSKGPVSFGASDTSFS--SGTFNVKP----S 448
++THANAKSKSEKFPPPH+DG +G+P S+ D S S TF+ +P S
Sbjct: 839 LITHANAKSKSEKFPPPHEDGQLGFPLGASNHIDPDIVPHDVSLDSMSYTFSKEPFQAYS 1018
Query: 449 GPTRSHDGTGLHKGTKTKKEESQMASSWKFMRPFK 483
GP + G TK++++ + +P+K
Sbjct: 1019GPIGNTTSNGF-----TKRKKNTANDALDLSKPYK 1108
Score = 58.2 bits (139), Expect(2) = e-130
Identities = 26/34 (76%), Positives = 29/34 (84%)
Frame = +3
Query: 116 MEHDLAGLSAGQGVKFTEPQVKCFMKQLLSGLEH 149
M HDLAGL+A +KFTEPQVKC+M QLLSGLEH
Sbjct: 3 MVHDLAGLAASPDIKFTEPQVKCYMHQLLSGLEH 104
>TC90675 similar to GP|20804521|dbj|BAB92215. putative CRK1 protein {Oryza
sativa (japonica cultivar-group)}, partial (33%)
Length = 1069
Score = 366 bits (939), Expect = e-101
Identities = 183/354 (51%), Positives = 249/354 (69%), Gaps = 8/354 (2%)
Frame = +2
Query: 123 LSAGQGVKFTEPQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLAT 182
L + G+KF+EPQ+KC+MKQLLSGL+HCHSRG+LHRDIKGSNLLIDN+GILKIADFGLA
Sbjct: 8 LVSAPGIKFSEPQIKCYMKQLLSGLDHCHSRGILHRDIKGSNLLIDNKGILKIADFGLAN 187
Query: 183 FYNPNKKQSMTSRVVTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEV 242
F++PN+ +TSRVVTLWYRPPELLLG++ YGV +D+WS GCIL EL G+PI+PG+TEV
Sbjct: 188 FFDPNRSAPLTSRVVTLWYRPPELLLGSSNYGVAVDMWSTGCILGELYCGRPILPGKTEV 367
Query: 243 EQLHKIFKLCGSPAEEYWRKHKLPNATIFKPQQPYKRCISETFKDFPPSSLPLIDSLLAI 302
EQLH+IFKLCGSP+ +YWRK +LP++T F+P Y+ C+ +TFK+ P +++ L+++LL++
Sbjct: 368 EQLHRIFKLCGSPSVDYWRKLRLPHSTAFRPPHHYRNCVKDTFKEVPSAAVRLMETLLSL 547
Query: 303 DPDRRGTASAALNHEFFTTEPYACEPSSLPKYPPSKELDVKMRDEEARRQKALNGKANAV 362
DP RGTA+ AL +EFF+ EP AC+PS+LPKYPP+KE+D KMR+E R++ AL GK V
Sbjct: 548 DPRLRGTAATALKNEFFSAEPVACDPSTLPKYPPTKEIDAKMREESTRKEGALGGKEQKV 727
Query: 363 DGAKRVRARERGRAIPAPEANAEIQTNLDRWRVVTHANAKSKSEKFPPPHQDGAVGY--- 419
RVR + ++ P+ANA ++ + + N S++E F PH++ A G+
Sbjct: 728 --GSRVRQEKEPQSFVLPKANAVSHISMQQGQ--HFPNMTSRNESF-NPHREPASGFMVL 892
Query: 420 PQDESSKGPVSFGASDTSFSSGTFNVKPSGPTRSHD-----GTGLHKGTKTKKE 468
P +S D ++ F+ P P SH G+G HK K E
Sbjct: 893 PHKQS---------EDVKETANYFS-GPLYPRSSHSGLLVPGSGRHKIGKEASE 1024
>TC81620 similar to GP|8953767|dbj|BAA98122.1 cyclin-dependent protein
kinase-like {Arabidopsis thaliana}, partial (30%)
Length = 810
Score = 263 bits (672), Expect(2) = 9e-84
Identities = 129/152 (84%), Positives = 139/152 (90%)
Frame = +3
Query: 2 LDPLSVNQQGWPSWLMAVAGEAIGDWTPRRANSFEKLAKIGQGTYSNVYKAKDLVTGKIV 61
L+ L+ QQ WP WLM VAG+AI DWTPRRAN+FEKLAKIG+GTYSNVYKAKDLVTGKIV
Sbjct: 240 LNSLAATQQSWPPWLMEVAGDAIRDWTPRRANTFEKLAKIGKGTYSNVYKAKDLVTGKIV 419
Query: 62 ALKKVRFDNLEPESVKFMAREILVLRKLDHPNVVKLEGLVTSRMSCSLYLVFEYMEHDLA 121
ALKKVR DNL+ ESVKFMAREILVLRKLDHPNV+KLEGLVTSR+S SLYLVFEYMEHDLA
Sbjct: 420 ALKKVRIDNLDAESVKFMAREILVLRKLDHPNVIKLEGLVTSRISSSLYLVFEYMEHDLA 599
Query: 122 GLSAGQGVKFTEPQVKCFMKQLLSGLEHCHSR 153
GL AG GVKF+ PQVKC+MKQLLSGLEHCHSR
Sbjct: 600 GLIAGLGVKFSLPQVKCYMKQLLSGLEHCHSR 695
Score = 65.9 bits (159), Expect(2) = 9e-84
Identities = 31/36 (86%), Positives = 34/36 (94%)
Frame = +1
Query: 154 GVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKK 189
GVLHRDIKGSNLLID+EGILKIADFGLATFY+ +K
Sbjct: 697 GVLHRDIKGSNLLIDDEGILKIADFGLATFYDSKQK 804
>TC89244 similar to GP|8953767|dbj|BAA98122.1 cyclin-dependent protein
kinase-like {Arabidopsis thaliana}, partial (19%)
Length = 809
Score = 244 bits (624), Expect = 4e-65
Identities = 130/184 (70%), Positives = 152/184 (81%), Gaps = 2/184 (1%)
Frame = +2
Query: 317 EFFTTEPYACEPSSLPKYPPSKELDVKMRDEEARRQKALNGKANAVDGAKRVRARERGRA 376
EFFTTEPYACEPSSLPKYPPSKELDVK+RDEEARRQ+AL+GK+NAVDGA++ RARER A
Sbjct: 14 EFFTTEPYACEPSSLPKYPPSKELDVKLRDEEARRQRALSGKSNAVDGARQSRARERSYA 193
Query: 377 IPAPEANAEIQTNLDRWRVVTHANAKSKSEKFPPPHQDGAVGYPQDESSKGPVSFGASD- 435
IPAPEANAEIQTNLDR RVVT+ N KSKSEKFPPPH+DGAVGYP D S+KG VSFG ++
Sbjct: 194 IPAPEANAEIQTNLDRLRVVTNGNGKSKSEKFPPPHEDGAVGYPGDGSNKGAVSFGTTEK 373
Query: 436 TSFSSGTFNVKPSGPTRSHDGTGLHKGTKTKKEESQMASSWKFMR-PFKPSTIGLSMDLL 494
TSFSS N S S+ G+ +K K+ + +S++ SSWKFMR FKPST+GLS +LL
Sbjct: 374 TSFSSILSNSIHSKSVGSYAGSS-YKRRKSNEADSRI-SSWKFMRSSFKPSTVGLSFNLL 547
Query: 495 FRSK 498
FRS+
Sbjct: 548 FRSR 559
>TC78173 SP|P24923|CC21_MEDSA Cell division control protein 2 homolog 1 (EC
2.7.1.-) (Fragment). [Alfalfa] {Medicago sativa},
complete
Length = 1268
Score = 222 bits (566), Expect = 2e-58
Identities = 126/289 (43%), Positives = 183/289 (62%), Gaps = 4/289 (1%)
Frame = +2
Query: 35 FEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDNLEPESVKFMA-REILVLRKLDHPN 93
+EK+ KIG+GTY VYKA+D VT + +ALKK+R + E E V A REI +L+++ H N
Sbjct: 137 YEKVEKIGEGTYGVVYKARDRVTNETIALKKIRLEQ-EDEGVPSTAIREISLLKEMQHRN 313
Query: 94 VVKLEGLVTSRMSCSLYLVFEYMEHDLAGLSAGQGVKFTEP-QVKCFMKQLLSGLEHCHS 152
+V+L+ +V S LYLVFEY++ DL +P QVK F+ Q+L G+ +CHS
Sbjct: 314 IVRLQDVVHSDKR--LYLVFEYLDLDLKKHMDSSPEFIKDPRQVKMFLYQMLCGIAYCHS 487
Query: 153 RGVLHRDIKGSNLLIDNE-GILKIADFGLATFYNPNKKQSMTSRVVTLWYRPPELLLGAT 211
VLHRD+K NLLID LK+ADFGLA + + + T VVTLWYR PE+LLG+
Sbjct: 488 HRVLHRDLKPQNLLIDRRTNSLKLADFGLARAFGIPVR-TFTHEVVTLWYRAPEILLGSR 664
Query: 212 FYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKLCGSPAEEYW-RKHKLPNATI 270
Y +D+WS GCI AE+ +P+ PG +E+++L KIF++ G+P E+ W LP+
Sbjct: 665 HYSTPVDVWSVGCIFAEMANRRPLFPGDSEIDELFKIFRILGTPNEDTWPGVTSLPDFKS 844
Query: 271 FKPQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGTASAALNHEFF 319
P+ P K ++ + P+ L L++S+L +DP +R TA +A+ HE+F
Sbjct: 845 TFPRWPSKD-LATVVPNLEPAGLDLLNSMLCLDPTKRITARSAVEHEYF 988
>TC87305 homologue to SP|Q05006|CC22_MEDSA Cell division control protein 2
homolog 2 (EC 2.7.1.-). [Alfalfa] {Medicago sativa},
complete
Length = 1378
Score = 199 bits (507), Expect(2) = 1e-56
Identities = 106/233 (45%), Positives = 153/233 (65%), Gaps = 3/233 (1%)
Frame = +2
Query: 31 RANSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDNLEPESVKFMA-REILVLRKL 89
R +EK+ KIG+GTY VYKA+D T + +ALKK+R + E E V A REI +L+++
Sbjct: 98 RMEQYEKVEKIGEGTYGVVYKARDRATNETIALKKIRLEQ-EDEGVPSTAIREISLLKEM 274
Query: 90 DHPNVVKLEGLVTSRMSCSLYLVFEYMEHDLAG-LSAGQGVKFTEPQVKCFMKQLLSGLE 148
H N+V+L+ +V S LYLVFEY++ DL + + + Q+K F+ Q+L G+
Sbjct: 275 QHRNIVRLQDVVHSEKR--LYLVFEYLDLDLKKFMDSSPEFAKDQRQIKMFLYQILCGIA 448
Query: 149 HCHSRGVLHRDIKGSNLLID-NEGILKIADFGLATFYNPNKKQSMTSRVVTLWYRPPELL 207
+CHS VLHRD+K NLLID + LK+ADFGLA + + + T VVTLWYR PE+L
Sbjct: 449 YCHSHRVLHRDLKPQNLLIDRSSNALKLADFGLARAFGIPVR-TFTHEVVTLWYRAPEIL 625
Query: 208 LGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKLCGSPAEEYW 260
LG+ Y +D+WS GCI AE++ +P+ PG +E+++L KIF++ G+P EE W
Sbjct: 626 LGSRHYSTPVDVWSVGCIFAEMINQRPLFPGDSEIDELFKIFRITGTPNEETW 784
Score = 38.5 bits (88), Expect(2) = 1e-56
Identities = 23/64 (35%), Positives = 34/64 (52%), Gaps = 5/64 (7%)
Frame = +3
Query: 261 RKH-----KLPNATIFKPQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGTASAALN 315
RKH LP+ P+ P K ++ + P+ L L+ ++L +DP RR TA AL
Sbjct: 774 RKHGPGVTSLPDFKSAFPKWPAKDLATQV-PNLEPAGLDLLSNMLCLDPTRRITARGALE 950
Query: 316 HEFF 319
HE+F
Sbjct: 951 HEYF 962
>AW573834 similar to PIR|D86320|D86 hypothetical protein AAF27112.1
[imported] - Arabidopsis thaliana, partial (20%)
Length = 402
Score = 210 bits (534), Expect = 1e-54
Identities = 97/134 (72%), Positives = 113/134 (83%)
Frame = +1
Query: 106 SCSLYLVFEYMEHDLAGLSAGQGVKFTEPQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNL 165
+C +YLVFEYME D+ GL + + FTE Q+KC+MKQLLSGLEHCHSRGV+HRDIKGSNL
Sbjct: 1 ACIIYLVFEYMELDVTGLWSKPEISFTESQIKCYMKQLLSGLEHCHSRGVMHRDIKGSNL 180
Query: 166 LIDNEGILKIADFGLATFYNPNKKQSMTSRVVTLWYRPPELLLGATFYGVGIDLWSAGCI 225
L++NEGILK+ADFGLA F N KKQ +TSRVVTLWYRPPELLLG+T YG +DLWS GC+
Sbjct: 181 LVNNEGILKVADFGLANFTNSGKKQPLTSRVVTLWYRPPELLLGSTDYGPSVDLWSVGCV 360
Query: 226 LAELLAGKPIMPGR 239
AELL GKPI+ GR
Sbjct: 361 FAELLVGKPIL*GR 402
>TC87516 PIR|T09591|T09591 probable cdc2-like protein kinase cdc2MsF -
alfalfa, complete
Length = 1251
Score = 196 bits (497), Expect = 2e-50
Identities = 119/296 (40%), Positives = 167/296 (56%), Gaps = 10/296 (3%)
Frame = +2
Query: 34 SFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDNLEPESVKFMAREILVLRKLDH-P 92
+FEKL K+G+GTY VY+A++ TGKIVALKK R + RE+ +LR L P
Sbjct: 107 AFEKLEKVGEGTYGKVYRAREKATGKIVALKKTRLHEDDEGVPPTTLREVSILRMLSRDP 286
Query: 93 NVVKLEGLVTSRMS---CSLYLVFEYMEHDLAGLSAG---QGVKFTEPQVKCFMKQLLSG 146
+VV+L + + LYLVFEYM+ DL G P +K M QL G
Sbjct: 287 HVVRLLDVKQGQNKEGKTVLYLVFEYMDTDLKKFIRSFRQTGQNIPPPTIKGLMYQLCKG 466
Query: 147 LEHCHSRGVLHRDIKGSNLLIDNEGI-LKIADFGLA-TFYNPNKKQSMTSRVVTLWYRPP 204
+ CH G+LHRD+K NLL+D + + LKIAD GLA F P KK T ++TLWYR P
Sbjct: 467 VAFCHGHGILHRDLKPHNLLMDRKTMMLKIADLGLARAFTVPLKKY--THEILTLWYRAP 640
Query: 205 ELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKLCGSPAEEYW-RKH 263
E+LLGAT Y + +D+WS CI AEL+ + PG +E++QL IF+L G+P E+ W
Sbjct: 641 EVLLGATHYSMAVDMWSVACIFAELVTKTALFPGDSELQQLLHIFRLLGTPNEDVWPGVS 820
Query: 264 KLPNATIFKPQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGTASAALNHEFF 319
KL N + P + +S+ + + L+ +L +P +R +A A+ H +F
Sbjct: 821 KLMNWHEYPQWGP--QSLSKAVPGLEETGVDLLSQMLQYEPSKRLSAKKAMEHPYF 982
>TC81436 similar to PIR|A96571|A96571 hypothetical protein F8L10.9
[imported] - Arabidopsis thaliana, partial (34%)
Length = 1060
Score = 196 bits (497), Expect = 2e-50
Identities = 94/171 (54%), Positives = 127/171 (73%)
Frame = +1
Query: 250 KLCGSPAEEYWRKHKLPNATIFKPQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGT 309
KLCGSP+E+YWRK KLP+ATIFKPQQPY+RC++ETFK+FP ++ LI++LL+IDP RGT
Sbjct: 1 KLCGSPSEDYWRKSKLPHATIFKPQQPYRRCVAETFKEFPAPAIELIETLLSIDPADRGT 180
Query: 310 ASAALNHEFFTTEPYACEPSSLPKYPPSKELDVKMRDEEARRQKALNGKANAVDGAKRVR 369
+++AL EFF+T+P C+PSSLPKYPPSKE D K+RDEEARRQ A K D + R
Sbjct: 181 SASALISEFFSTKPLPCDPSSLPKYPPSKEFDAKVRDEEARRQGAAGSKGQRHDPER--R 354
Query: 370 ARERGRAIPAPEANAEIQTNLDRWRVVTHANAKSKSEKFPPPHQDGAVGYP 420
RA+PAP+ANAE+ ++ + + + ++S+SEKF P +D G+P
Sbjct: 355 GVRESRAVPAPDANAELVVSMQKRQGQNY--SQSRSEKFNPHPEDAGSGFP 501
>BG648590 similar to PIR|H96771|H96 hypothetical protein F1M20.1 [imported] -
Arabidopsis thaliana, partial (40%)
Length = 805
Score = 187 bits (474), Expect = 1e-47
Identities = 88/118 (74%), Positives = 100/118 (84%)
Frame = +1
Query: 135 QVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQSMTS 194
Q+KC+MKQLLSGLEHCHSRGV+HRDIKGSNLL++NEGILK+ADFGLA F N KKQ +TS
Sbjct: 85 QIKCYMKQLLSGLEHCHSRGVMHRDIKGSNLLVNNEGILKVADFGLANFTNSGKKQPLTS 264
Query: 195 RVVTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKLC 252
RVVTLWYRPPELLLG+T YG +DLWS GC+ AELL GKPI+ GRTEV + IF C
Sbjct: 265 RVVTLWYRPPELLLGSTDYGPSVDLWSVGCVFAELLVGKPILKGRTEV--VIPIFSKC 432
Score = 113 bits (283), Expect = 1e-25
Identities = 50/76 (65%), Positives = 66/76 (86%)
Frame = +2
Query: 242 VEQLHKIFKLCGSPAEEYWRKHKLPNATIFKPQQPYKRCISETFKDFPPSSLPLIDSLLA 301
VEQLHKIFKLCGSP +EYW+K +LP+AT+FKPQQPY C+ ETFKDF +S+ L+ +LL+
Sbjct: 431 VEQLHKIFKLCGSPPDEYWKKTRLPHATLFKPQQPYDSCLRETFKDFSATSVNLLQNLLS 610
Query: 302 IDPDRRGTASAALNHE 317
I+P++RGTAS+AL+ E
Sbjct: 611 IEPNKRGTASSALSLE 658
>TC92292 homologue to PIR|T09572|T09572 cdc2-like protein kinase cdc2MsC -
alfalfa, partial (38%)
Length = 711
Score = 172 bits (435), Expect = 3e-43
Identities = 90/185 (48%), Positives = 124/185 (66%), Gaps = 15/185 (8%)
Frame = +1
Query: 30 RRANSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDNLEPESVKFMA-REILVLRK 88
R + FEKL +IG+GTY VY A+++ TG+IVALKK+ + A +I +L+K
Sbjct: 157 RSVDCFEKLEQIGEGTYGMVYMAREIETGEIVALKKIPIG**KEXGFPITAIPQIKILKK 336
Query: 89 LDHPNVVKLEGLVTS--------------RMSCSLYLVFEYMEHDLAGLSAGQGVKFTEP 134
L H NV+KL+ +VTS + +Y+VFEYM+HDL GL+ G++FT P
Sbjct: 337 LHHENVIKLKEIVTSPGPEKDDQGRPDGNKYKGGIYMVFEYMDHDLTGLADRPGMRFTVP 516
Query: 135 QVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQSMTS 194
Q+KC+M+QLL+GL +CH VLHRDIKGSNLLIDNEG LK+ADFGLA ++ ++T+
Sbjct: 517 QIKCYMRQLLTGLHYCHVNQVLHRDIKGSNLLIDNEGNLKLADFGLARSFSNEHNANLTN 696
Query: 195 RVVTL 199
RV+TL
Sbjct: 697 RVITL 711
>AJ497909 similar to GP|8953767|dbj| cyclin-dependent protein kinase-like
{Arabidopsis thaliana}, partial (13%)
Length = 263
Score = 169 bits (427), Expect = 3e-42
Identities = 82/82 (100%), Positives = 82/82 (100%)
Frame = +3
Query: 333 KYPPSKELDVKMRDEEARRQKALNGKANAVDGAKRVRARERGRAIPAPEANAEIQTNLDR 392
KYPPSKELDVKMRDEEARRQKALNGKANAVDGAKRVRARERGRAIPAPEANAEIQTNLDR
Sbjct: 18 KYPPSKELDVKMRDEEARRQKALNGKANAVDGAKRVRARERGRAIPAPEANAEIQTNLDR 197
Query: 393 WRVVTHANAKSKSEKFPPPHQD 414
WRVVTHANAKSKSEKFPPPHQD
Sbjct: 198 WRVVTHANAKSKSEKFPPPHQD 263
>TC76803 PIR|T09622|T09622 protein kinase MMK4 (EC 2.7.1.-) cold- and
drought-induced - alfalfa, complete
Length = 1709
Score = 165 bits (418), Expect = 3e-41
Identities = 115/319 (36%), Positives = 163/319 (51%), Gaps = 21/319 (6%)
Frame = +2
Query: 33 NSFEKLAK-------IGQGTYSNVYKAKDLVTGKIVALKKVR--FDNLEPESVKFMAREI 83
N FE AK IG+G Y V + T ++VA+KK+ FDN K REI
Sbjct: 203 NLFEVTAKYRPPIMPIGRGAYGIVCSLLNTETNELVAVKKIANAFDN--HMDAKRTLREI 376
Query: 84 LVLRKLDHPNVVKLEGLVTS---RMSCSLYLVFEYMEHDLAGLSAGQGVKFTEPQVKCFM 140
+LR LDH NV+ L ++ R +Y+ E M+ DL + ++ + F+
Sbjct: 377 KLLRHLDHENVIGLRDVIPPPLRREFNDVYITTELMDTDLHQIIRSNQ-NLSDEHCQYFL 553
Query: 141 KQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQS-MTSRVVTL 199
Q+L GL + HS ++HRD+K SNLL++ LKI DFGLA P + MT VVT
Sbjct: 554 YQILRGLRYIHSANIIHRDLKPSNLLLNANCDLKIIDFGLA---RPTMESDFMTEYVVTR 724
Query: 200 WYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKLCGSPAEEY 259
WYR PELLL ++ Y ID+WS GCI EL+ KP+ PG+ V Q+ + +L G+P +
Sbjct: 725 WYRAPELLLNSSDYTSAIDVWSVGCIFMELMNKKPLFPGKDHVHQMRLLTELLGTPTDAD 904
Query: 260 WRKHKLPNATIFKPQQPY--KRCISETFKDFPPSSLPLIDSLLAIDPDRRGTASAALNHE 317
K +A + Q P ++ ++ F P ++ L+D +L IDP RR T AL H
Sbjct: 905 VGLVKNDDARRYIRQLPQYPRQPLNRVFPHVHPLAIDLVDKMLTIDPTRRITVEEALAHP 1084
Query: 318 FF------TTEPYACEPSS 330
+ EP EP S
Sbjct: 1085YLEKLHDVADEPICMEPFS 1141
>TC86213 homologue to PIR|S60121|S60121 mitogen-activated protein kinase
MMK2 (EC 2.7.1.-) - alfalfa, complete
Length = 1596
Score = 165 bits (418), Expect = 3e-41
Identities = 113/317 (35%), Positives = 163/317 (50%), Gaps = 14/317 (4%)
Frame = +2
Query: 41 IGQGTYSNVYKAKDLVTGKIVALKKV--RFDNLEPESVKFMAREILVLRKLDHPNVVKLE 98
+G+G Y V A + T + VA+KK+ FDN K REI +LR +DH NV+ ++
Sbjct: 368 VGRGAYGIVCAAVNAETREEVAIKKIGNAFDNRI--DAKRTLREIKLLRHMDHENVMSIK 541
Query: 99 GLVTSRMSCS---LYLVFEYMEHDLAGLSAGQGVKFTEPQVKCFMKQLLSGLEHCHSRGV 155
++ + +Y+V E M+ DL + T+ + F+ QLL GL++ HS V
Sbjct: 542 DIIRPPQKENFNDVYIVSELMDTDLHQIIRSNQ-PMTDDHCRYFVYQLLRGLKYVHSANV 718
Query: 156 LHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQSMTSRVVTLWYRPPELLLGATFYGV 215
LHRD+K SNLL++ LKI DFGLA ++ MT VVT WYR PELLL + Y
Sbjct: 719 LHRDLKPSNLLLNANCDLKIGDFGLAR--TTSETDFMTEYVVTRWYRAPELLLNCSEYTA 892
Query: 216 GIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKLCGSPAEEYWRKHKLPNATIFKPQQ 275
ID+WS GCIL E++ +P+ PGR V QL + +L GSP + + NA + Q
Sbjct: 893 AIDIWSVGCILGEIVTRQPLFPGRDYVHQLRLVTELIGSPDDASLGFLRSENARRYVRQL 1072
Query: 276 PY--KRCISETFKDFPPSSLPLIDSLLAIDPDRRGTASAALNHEF------FTTEPYACE 327
P ++ S F P ++ L++ +L DP +R AL H + EP
Sbjct: 1073PQYPQQNFSTRFPSMSPGAVDLLEKMLIFDPSKRIRVDEALCHPYMAPLHDINEEPICAR 1252
Query: 328 PSSLP-KYPPSKELDVK 343
P S + P E D+K
Sbjct: 1253PFSFDFEEPMFTEEDIK 1303
>TC81224 homologue to GP|17064846|gb|AAL32577.1 putative protein kinase
{Arabidopsis thaliana}, partial (34%)
Length = 868
Score = 162 bits (411), Expect = 2e-40
Identities = 95/222 (42%), Positives = 130/222 (57%), Gaps = 10/222 (4%)
Frame = +3
Query: 30 RRANSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDNLEPESVKFMAREILVLRKL 89
R + FE+L KI +GTY VY+AKD TG++VALKKV+ + + REI +L
Sbjct: 138 RSVDEFERLNKIDEGTYGVVYRAKDKKTGEVVALKKVKMEKEKEGFPLTSLREINILLSF 317
Query: 90 DHPNVVKLEGLVTSRMSCSLYLVFEYMEHDLAGLSAGQGVKFTEPQVKCFMKQLLSGLEH 149
HP +V ++ +V S+++V EYMEHDL GL F++ +VKC M QLL G+++
Sbjct: 318 HHPYIVDVKEVVVGSSLDSIFMVMEYMEHDLKGLMESMKQPFSQSEVKCLMIQLLEGVKY 497
Query: 150 CHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQSMTSRVVTLWY-----RPP 204
H VLHRD+K SNLL++N G LKI DFGLA Y S ++ W+ P
Sbjct: 498 LHDNWVLHRDLKTSNLLLNNRGELKICDFGLARQYG-----SPFETIILTWWLLFGTGAP 662
Query: 205 ELLLGATFYG-----VGIDLWSAGCILAELLAGKPIMPGRTE 241
ELLLGA Y VG+ GCI+AELL+ +P+ G+TE
Sbjct: 663 ELLLGAKQYSNSY*YVGL----LGCIMAELLSKEPLFNGKTE 776
>TC77816 similar to GP|14532760|gb|AAK64081.1 putative kinase {Arabidopsis
thaliana}, partial (75%)
Length = 2695
Score = 162 bits (409), Expect = 4e-40
Identities = 120/390 (30%), Positives = 192/390 (48%), Gaps = 10/390 (2%)
Frame = +3
Query: 58 GKIVALKKVRFDNLEPESVKFMAREILVLRKLDHPNVVKLEGLVTSRMSCSLYLVFEYME 117
G++VA+KK++ E + RE+ LRK++H NVVKL+ ++ R S LYLVFEYME
Sbjct: 1083 GEVVAIKKMKKKYYSWEECVNL-REVKSLRKMNHSNVVKLKEVI--RESDILYLVFEYME 1253
Query: 118 HDLAGLSAGQGVKFTEPQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIAD 177
+L L + F++ +V+ Q+ GL + H RG HRD+K NLL+ + I+K++D
Sbjct: 1254 CNLYQLMKKREKLFSDDEVRNLCFQVFQGLAYMHQRGYFHRDLKPENLLVTKD-IIKVSD 1430
Query: 178 FGLATFYNPNKKQSMTSRVVTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMP 237
FGL + + T V T WYR PE+LL ++ Y +D+W+ G I+AEL +P+ P
Sbjct: 1431 FGLVR--EISSQPPYTEYVSTRWYRAPEVLLQSSLYSSKVDMWAMGAIMAELFTLRPLFP 1604
Query: 238 GRTEVEQLHKIFKLCGSPAEEYWRKH-KLPNATIFKPQQPYKRCISETFKDFPPSSLPLI 296
G +E ++++KI + GSP E W KL ++ Q +S ++ LI
Sbjct: 1605 GTSEADEIYKICGVIGSPTAESWADGLKLSRDINYQFPQVATADLSVLIPSRSDDAINLI 1784
Query: 297 DSLLAIDPDRRGTASAALNHEFFTT---EPYACEPSSLPKYPPSKELDVKMRDEEARRQK 353
SL + DP +R +A+ AL H FF + P + + + PPS + + + +R
Sbjct: 1785 KSLCSWDPCKRPSAAEALQHPFFKSCFYIPPSLRTKGVTRTPPSAGIRGSVDRQGVKRYS 1964
Query: 354 A--LNGKANAVDGAKRVRARERGRAIPAPEANAEIQTNLDRWRVVTHANAK----SKSEK 407
LN K + ++ P +Q LD N K ++ K
Sbjct: 1965 GGLLNSKLTTNFSSNKLH----------PALAPGVQRKLDMANEDGTKNKKTVKTTQPSK 2114
Query: 408 FPPPHQDGAVGYPQDESSKGPVSFGASDTS 437
+ PP +D +KG + G S+T+
Sbjct: 2115 YRPPGKDSPTSI-----NKGRTARGVSETA 2189
>TC91359 homologue to PIR|T09586|T09586 probable cdc2-like protein kinase
cdc2MsD - alfalfa, partial (83%)
Length = 833
Score = 161 bits (407), Expect = 6e-40
Identities = 101/263 (38%), Positives = 143/263 (53%), Gaps = 17/263 (6%)
Frame = +3
Query: 49 VYKAKDLVTGKIVALKKVRFDNLEPESVKFMAREILVLRKLDHP----NVVKLEGLVTSR 104
VYKAK++ TG+IVALKK R + E RE+ +L+ L ++ +E +
Sbjct: 42 VYKAKEISTGQIVALKKTRLEMDEEGVPPTALREVSLLQMLSQSLYIVRLLNVEHIDKPP 221
Query: 105 MSCS-------LYLVFEYMEHDLAGL--SAGQGVK---FTEPQVKCFMKQLLSGLEHCHS 152
+ + LYLVFEY++ DL + +GV V+ F+ QL G+ HCHS
Sbjct: 222 KNATHTPAKPLLYLVFEYLDTDLKKFIDTFRKGVNPRPLPNTLVQSFLFQLCKGVAHCHS 401
Query: 153 RGVLHRDIKGSNLLIDN-EGILKIADFGLATFYNPNKKQSMTSRVVTLWYRPPELLLGAT 211
GVLHRD+K NLL+D +GILKIAD GL + K S T +VTLWYR PE+LLG++
Sbjct: 402 HGVLHRDLKPQNLLLDQAKGILKIADLGLGRAFTVPLK-SYTHEIVTLWYRAPEVLLGSS 578
Query: 212 FYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKLCGSPAEEYWRKHKLPNATIF 271
Y +D+WS GCI AE++ + + PG +E +QL IFKL G+P ++ W P +
Sbjct: 579 TYSTSVDIWSVGCIFAEMVRRQALFPGDSEFQQLLNIFKLLGTPTDQQW-----PGVSSL 743
Query: 272 KPQQPYKRCISETFKDFPPSSLP 294
+ Y R + PS P
Sbjct: 744 RDWHVYPRWEPQNLARAVPSLSP 812
>TC87466 homologue to GP|10185114|emb|CAC08564. wound-induced GSK-3-like
protein {Medicago sativa}, complete
Length = 1934
Score = 159 bits (401), Expect = 3e-39
Identities = 116/322 (36%), Positives = 167/322 (51%), Gaps = 12/322 (3%)
Frame = +3
Query: 26 DWTPRRANSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDNLEPESVKFMAREILV 85
D P++ S+ +G G++ V++AK L T + VA+KKV D ++ RE+ V
Sbjct: 411 DGQPKQTISYMAERVVGTGSFGVVFQAKCLETNEAVAIKKVLQDK------RYKNRELQV 572
Query: 86 LRKLDHPNVVKLEGL---VTSRMSCSLYLVFEYMEHDLAGLSAGQGVKFTEPQ----VKC 138
+R +DHPN+VKL+ T + L LV EY+ + +S ++ + V+
Sbjct: 573 MRMVDHPNIVKLKHCFYSTTEKDELYLNLVLEYVPETVYKVSKNY-IRMHQHMPIIHVQL 749
Query: 139 FMKQLLSGLEHCHSR-GVLHRDIKGSNLLIDNEGI-LKIADFGLATFYNPNKKQSMTSRV 196
+ Q+L GL + H GV HRDIK NLL++ + LKI DFG A P + S +
Sbjct: 750 YTYQILRGLNYLHEVIGVCHRDIKPQNLLVNAQTHQLKICDFGSAKMLVPGEPN--ISYI 923
Query: 197 VTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKLCGSPA 256
+ +YR PEL+ GAT Y ID+WS GC+LAELL G+ + PG + V+QL +I K+ G+P
Sbjct: 924 CSRYYRAPELIFGATEYTTAIDMWSVGCVLAELLLGQAMFPGESGVDQLVEIIKVLGTPT 1103
Query: 257 EEYWRKHKLPNATIFKPQQPYKRCISETF-KDFPPSSLPLIDSLLAIDPDRRGTASAALN 315
E R PN FK Q + F K P ++ L+ LL P R TA AA
Sbjct: 1104REEIRCMN-PNYNEFKFPQIKAHPWHKLFHKRMPSEAVDLVSRLLQYSPHLRCTALAACA 1280
Query: 316 HEFFT--TEPYACEPSSLPKYP 335
H FF +P A P+ P P
Sbjct: 1281HPFFNDLRDPNASLPNGQPLPP 1346
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.317 0.134 0.403
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,769,872
Number of Sequences: 36976
Number of extensions: 224041
Number of successful extensions: 1973
Number of sequences better than 10.0: 546
Number of HSP's better than 10.0 without gapping: 1632
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1666
length of query: 498
length of database: 9,014,727
effective HSP length: 100
effective length of query: 398
effective length of database: 5,317,127
effective search space: 2116216546
effective search space used: 2116216546
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 60 (27.7 bits)
Medicago: description of AC146747.6


