

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
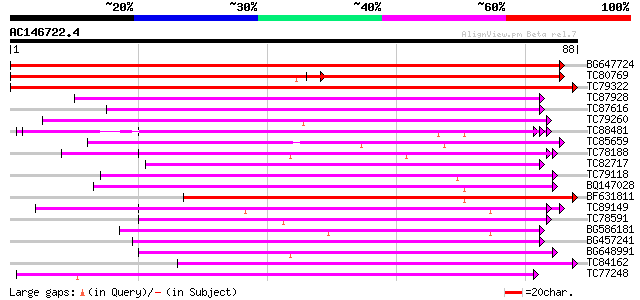
Query= AC146722.4 - phase: 1
(88 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BG647724 similar to GP|11141605|gb LEUNIG {Arabidopsis thaliana}... 175 2e-45
TC80769 similar to GP|11141605|gb|AAG32022.1 LEUNIG {Arabidopsis... 78 4e-31
TC79322 similar to GP|13605815|gb|AAK32893.1 At2g32700/F24L7.16 ... 120 1e-28
TC87928 similar to PIR|A96839|A96839 F23A5.2(form2) [imported] -... 57 9e-10
TC87616 similar to PIR|A96839|A96839 F23A5.2(form2) [imported] -... 55 4e-09
TC79260 weakly similar to GP|9759350|dbj|BAB10005.1 WD-repeat pr... 49 3e-07
TC88481 similar to PIR|T00806|T02445 probable U4/U6 small nuclea... 48 6e-07
TC85659 SP|O24076|GBLP_MEDSA Guanine nucleotide-binding protein ... 48 6e-07
TC78188 similar to PIR|C84870|C84870 probable splicing factor [i... 47 2e-06
TC82717 weakly similar to GP|21539583|gb|AAM53344.1 putative coa... 45 4e-06
TC79118 similar to GP|16323188|gb|AAL15328.1 AT5g67320/K8K14_4 {... 44 1e-05
BQ147028 similar to GP|16323188|gb| AT5g67320/K8K14_4 {Arabidops... 44 1e-05
BF631811 similar to GP|9759350|dbj WD-repeat protein-like {Arabi... 43 2e-05
TC89149 similar to GP|20466231|gb|AAM20433.1 cell cycle switch p... 43 2e-05
TC78591 similar to GP|9759350|dbj|BAB10005.1 WD-repeat protein-l... 42 4e-05
BG586181 similar to GP|4689316|gb| peroxisomal targeting signal ... 42 4e-05
BG457241 homologue to PIR|F84600|F8 coatomer alpha subunit [impo... 41 7e-05
BG648991 similar to PIR|C86239|C86 protein T10O24.21 [imported] ... 41 9e-05
TC84162 similar to GP|15810485|gb|AAL07130.1 putative WD40-repea... 40 1e-04
TC77248 similar to GP|13489182|gb|AAK27816.1 putative WD-repeat ... 40 1e-04
>BG647724 similar to GP|11141605|gb LEUNIG {Arabidopsis thaliana}, partial
(25%)
Length = 757
Score = 175 bits (444), Expect = 2e-45
Identities = 86/86 (100%), Positives = 86/86 (100%)
Frame = +2
Query: 1 FTFSELNSVRASTNKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITD 60
FTFSELNSVRASTNKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITD
Sbjct: 395 FTFSELNSVRASTNKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITD 574
Query: 61 VRFSPSMPRLATSSYDKTVRVWDVDN 86
VRFSPSMPRLATSSYDKTVRVWDVDN
Sbjct: 575 VRFSPSMPRLATSSYDKTVRVWDVDN 652
>TC80769 similar to GP|11141605|gb|AAG32022.1 LEUNIG {Arabidopsis thaliana},
partial (40%)
Length = 1362
Score = 78.2 bits (191), Expect(2) = 4e-31
Identities = 37/50 (74%), Positives = 43/50 (86%), Gaps = 1/50 (2%)
Frame = +2
Query: 1 FTFSELNSVRASTNKVVCSHFSSDGKLLASGGHDKKVVLWYTD-SLKQKA 49
FTFS++NSVRAS++K+ C HFSSDGKLLASGGHDKK V+WY LKQKA
Sbjct: 326 FTFSDVNSVRASSSKIACCHFSSDGKLLASGGHDKKAVIWYRRIHLKQKA 475
Score = 71.2 bits (173), Expect(2) = 4e-31
Identities = 34/40 (85%), Positives = 36/40 (90%)
Frame = +3
Query: 47 QKATLEEHSLLITDVRFSPSMPRLATSSYDKTVRVWDVDN 86
+K LEEHS LITDVRFS SMPRLATSS+DKTVRVWDVDN
Sbjct: 468 KKPILEEHSALITDVRFSASMPRLATSSFDKTVRVWDVDN 587
Score = 26.2 bits (56), Expect = 2.3
Identities = 12/33 (36%), Positives = 20/33 (60%)
Frame = +3
Query: 50 TLEEHSLLITDVRFSPSMPRLATSSYDKTVRVW 82
TL H LIT + S +A++S+DK +++W
Sbjct: 1095 TLSAHDGLITALAVSTVNGLVASASHDKFIKLW 1193
>TC79322 similar to GP|13605815|gb|AAK32893.1 At2g32700/F24L7.16
{Arabidopsis thaliana}, partial (59%)
Length = 1824
Score = 120 bits (300), Expect = 1e-28
Identities = 56/88 (63%), Positives = 72/88 (81%)
Frame = +2
Query: 1 FTFSELNSVRASTNKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITD 60
F+FSE++S+R S KVVC HFSSDGKLLAS GHDKKVVLW ++LK ++T EEH+++ITD
Sbjct: 764 FSFSEVSSIRKSNGKVVCCHFSSDGKLLASAGHDKKVVLWNMETLKTQSTPEEHTVIITD 943
Query: 61 VRFSPSMPRLATSSYDKTVRVWDVDNVS 88
VRF P+ +LATSS+D TVR+WD + S
Sbjct: 944 VRFRPNSTQLATSSFDTTVRLWDAADPS 1027
Score = 32.7 bits (73), Expect = 0.025
Identities = 20/65 (30%), Positives = 35/65 (53%)
Frame = +2
Query: 18 CSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSYDK 77
C S LL GG+ + + LW+ K T+ H +I+ + SP +A++S+DK
Sbjct: 1442 CVFHPSYSNLLVIGGY-QSLELWHMAENKCM-TIPAHEGVISALAQSPVTGMVASASHDK 1615
Query: 78 TVRVW 82
+V++W
Sbjct: 1616 SVKIW 1630
>TC87928 similar to PIR|A96839|A96839 F23A5.2(form2) [imported] -
Arabidopsis thaliana, partial (95%)
Length = 1519
Score = 57.4 bits (137), Expect = 9e-10
Identities = 29/73 (39%), Positives = 41/73 (55%)
Frame = +2
Query: 11 ASTNKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRL 70
A + V+CS + DG + SGG DK+V +W S Q T+ H I D+ + P M L
Sbjct: 374 AHDHPVLCSAWKDDGTTVFSGGCDKQVKMWPLLSGGQPITVAMHDAPIKDIAWIPEMNLL 553
Query: 71 ATSSYDKTVRVWD 83
AT S+DK ++ WD
Sbjct: 554 ATGSWDKNIKYWD 592
Score = 26.9 bits (58), Expect = 1.4
Identities = 10/27 (37%), Positives = 16/27 (59%)
Frame = +2
Query: 58 ITDVRFSPSMPRLATSSYDKTVRVWDV 84
++ + FSP L +S+D VR W+V
Sbjct: 245 VSSLNFSPKSNLLVATSWDNQVRCWEV 325
>TC87616 similar to PIR|A96839|A96839 F23A5.2(form2) [imported] -
Arabidopsis thaliana, partial (96%)
Length = 1471
Score = 55.5 bits (132), Expect = 4e-09
Identities = 28/68 (41%), Positives = 38/68 (55%)
Frame = +3
Query: 16 VVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSY 75
V+CS + DG + SGG DK+ +W S Q T+ H I ++ + P M LAT S
Sbjct: 348 VLCSAWKDDGTTVFSGGCDKQAKMWPLLSGGQPVTVAMHDAPIKEIAWIPEMSLLATGSL 527
Query: 76 DKTVRVWD 83
DKTV+ WD
Sbjct: 528 DKTVKYWD 551
Score = 26.2 bits (56), Expect = 2.3
Identities = 10/27 (37%), Positives = 16/27 (59%)
Frame = +3
Query: 58 ITDVRFSPSMPRLATSSYDKTVRVWDV 84
I+ + FSP L +S+D VR W++
Sbjct: 207 ISSLSFSPKANFLVATSWDNQVRCWEI 287
>TC79260 weakly similar to GP|9759350|dbj|BAB10005.1 WD-repeat protein-like
{Arabidopsis thaliana}, partial (50%)
Length = 1227
Score = 48.9 bits (115), Expect = 3e-07
Identities = 27/82 (32%), Positives = 42/82 (50%), Gaps = 3/82 (3%)
Frame = +2
Query: 6 LNSVRASTNKVVCSHFSSDGKLLASGGHDKKVVLWYTDS---LKQKATLEEHSLLITDVR 62
L + A ++V FS DGK LA+ +D+ ++W D+ L K L H ++ V
Sbjct: 524 LQILDAHDDEVWFVQFSHDGKYLATASNDRTAIIWEVDTNDGLSMKHKLSGHQKSVSSVS 703
Query: 63 FSPSMPRLATSSYDKTVRVWDV 84
+SP+ L T ++ VR WDV
Sbjct: 704 WSPNGQELLTCGVEEAVRRWDV 769
Score = 29.6 bits (65), Expect = 0.21
Identities = 21/60 (35%), Positives = 31/60 (51%), Gaps = 8/60 (13%)
Frame = +2
Query: 34 DKKVVLWYTDSLKQKATLEEHSLLITD--------VRFSPSMPRLATSSYDKTVRVWDVD 85
DK++ L Y+D K + +L I D V+FS LAT+S D+T +W+VD
Sbjct: 461 DKEMSL-YSDHHCGKTQIPSRTLQILDAHDDEVWFVQFSHDGKYLATASNDRTAIIWEVD 637
>TC88481 similar to PIR|T00806|T02445 probable U4/U6 small nuclear
ribonucleoprotein [imported] - Arabidopsis thaliana,
partial (60%)
Length = 1836
Score = 48.1 bits (113), Expect = 6e-07
Identities = 32/82 (39%), Positives = 45/82 (54%), Gaps = 1/82 (1%)
Frame = +1
Query: 3 FSELNSVRASTNKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVR 62
FSE+ R T CS FS DGK LA+ LW ++K+ +TL+ H+ TDV
Sbjct: 931 FSEIGDDRPLTG---CS-FSRDGKGLATCSFTGATKLWSMPNVKKVSTLKGHTQRATDVA 1098
Query: 63 FSP-SMPRLATSSYDKTVRVWD 83
+SP LAT+S D+T + W+
Sbjct: 1099 YSPVHKNHLATASADRTAKYWN 1164
Score = 47.4 bits (111), Expect = 1e-06
Identities = 26/63 (41%), Positives = 34/63 (53%), Gaps = 1/63 (1%)
Frame = +1
Query: 21 FSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPR-LATSSYDKTV 79
FS +G LA+GG D +W K T+ HS LI+ V+F P L T+SYD T
Sbjct: 1477 FSPNGYHLATGGEDNTCRIWDLRKKKSLYTIPAHSNLISQVKFEPQEGYFLVTASYDMTA 1656
Query: 80 RVW 82
+VW
Sbjct: 1657 KVW 1665
Score = 43.9 bits (102), Expect = 1e-05
Identities = 28/83 (33%), Positives = 36/83 (42%)
Frame = +1
Query: 2 TFSELNSVRASTNKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDV 61
T EL + V F DG L AS G D +W + + LE H I +
Sbjct: 1294 TEEELLLQEGHSRSVYGLDFHHDGSLAASCGLDALARVWDLRTGRSVLALEGHVKSILGI 1473
Query: 62 RFSPSMPRLATSSYDKTVRVWDV 84
FSP+ LAT D T R+WD+
Sbjct: 1474 SFSPNGYHLATGGEDNTCRIWDL 1542
Score = 33.1 bits (74), Expect = 0.019
Identities = 18/78 (23%), Positives = 35/78 (44%), Gaps = 1/78 (1%)
Frame = +1
Query: 6 LNSVRASTNKVVCSHFS-SDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFS 64
L ++ A +N + F +G L + +D +W + K TL H +T +
Sbjct: 1558 LYTIPAHSNLISQVKFEPQEGYFLVTASYDMTAKVWSSRDFKPVKTLSGHEAKVTSLDVL 1737
Query: 65 PSMPRLATSSYDKTVRVW 82
+ T S+D+T+++W
Sbjct: 1738 GDGGYIVTVSHDRTIKLW 1791
>TC85659 SP|O24076|GBLP_MEDSA Guanine nucleotide-binding protein beta
subunit-like protein. [Alfalfa] {Medicago sativa},
complete
Length = 1261
Score = 48.1 bits (113), Expect = 6e-07
Identities = 28/79 (35%), Positives = 47/79 (59%), Gaps = 5/79 (6%)
Frame = +2
Query: 13 TNKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEE---HSLLITDVRFSPS--M 67
T V+ FS D + + S D+ + LW T + K T+++ HS ++ VRFSPS
Sbjct: 374 TKDVLSVAFSIDNRQIVSASRDRTIKLWNTLG-ECKYTIQDGDAHSDWVSCVRFSPSTLQ 550
Query: 68 PRLATSSYDKTVRVWDVDN 86
P + ++S+D+TV+VW++ N
Sbjct: 551 PTIVSASWDRTVKVWNLTN 607
Score = 37.4 bits (85), Expect = 0.001
Identities = 22/67 (32%), Positives = 40/67 (58%)
Frame = +2
Query: 22 SSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSYDKTVRV 81
S DG L ASGG D ++LW K+ +L+ S +I + FSP+ L ++ + ++++
Sbjct: 665 SPDGSLCASGGKDGVILLWDLAEGKRLYSLDAGS-IIHALCFSPNRYWLCAAT-ESSIKI 838
Query: 82 WDVDNVS 88
WD+++ S
Sbjct: 839 WDLESKS 859
Score = 35.8 bits (81), Expect = 0.003
Identities = 20/71 (28%), Positives = 36/71 (50%), Gaps = 6/71 (8%)
Frame = +2
Query: 19 SHF------SSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLAT 72
SHF SSDG+ SG D ++ LW ++ H+ + V FS ++ +
Sbjct: 248 SHFVQDVVLSSDGQFALSGSWDGELRLWDLNAGTSARRFVGHTKDVLSVAFSIDNRQIVS 427
Query: 73 SSYDKTVRVWD 83
+S D+T+++W+
Sbjct: 428 ASRDRTIKLWN 460
>TC78188 similar to PIR|C84870|C84870 probable splicing factor [imported] -
Arabidopsis thaliana, partial (95%)
Length = 1436
Score = 46.6 bits (109), Expect = 2e-06
Identities = 21/66 (31%), Positives = 38/66 (56%), Gaps = 1/66 (1%)
Frame = +1
Query: 21 FSSDGKLLASGGHDKKVVLWYT-DSLKQKATLEEHSLLITDVRFSPSMPRLATSSYDKTV 79
F+ G ++ASG HDK++ LW K L+ H + D+ ++ ++ ++S DKT+
Sbjct: 337 FNPTGSVVASGSHDKEIFLWNVHGDCKNFMVLKGHKNAVLDLHWTSDGTQIISASPDKTL 516
Query: 80 RVWDVD 85
R+WD +
Sbjct: 517 RLWDTE 534
Score = 40.0 bits (92), Expect = 2e-04
Identities = 24/79 (30%), Positives = 40/79 (50%), Gaps = 3/79 (3%)
Frame = +1
Query: 9 VRASTNKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITD---VRFSP 65
++ N V+ H++SDG + S DK + LW T++ KQ + EH + R P
Sbjct: 430 LKGHKNAVLDLHWTSDGTQIISASPDKTLRLWDTETGKQIKKMVEHLSYVNSCCPTRRGP 609
Query: 66 SMPRLATSSYDKTVRVWDV 84
P + + S D T ++WD+
Sbjct: 610 --PLVVSGSDDGTAKLWDM 660
Score = 26.2 bits (56), Expect = 2.3
Identities = 10/34 (29%), Positives = 21/34 (61%)
Frame = +1
Query: 51 LEEHSLLITDVRFSPSMPRLATSSYDKTVRVWDV 84
L H + ++F+P+ +A+ S+DK + +W+V
Sbjct: 301 LTGHQSAVYTMKFNPTGSVVASGSHDKEIFLWNV 402
>TC82717 weakly similar to GP|21539583|gb|AAM53344.1 putative coatomer
complex subunit {Arabidopsis thaliana}, partial (29%)
Length = 1118
Score = 45.4 bits (106), Expect = 4e-06
Identities = 23/62 (37%), Positives = 33/62 (53%)
Frame = +1
Query: 22 SSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSYDKTVRV 81
S+D + L SG D +W DS TLE H +T + P +P + T+S D TV++
Sbjct: 676 SNDKQYLLSGSDDYTAKVWDYDSKNCVQTLEGHKNNVTAICAHPEIPIIITASEDSTVKI 855
Query: 82 WD 83
WD
Sbjct: 856 WD 861
>TC79118 similar to GP|16323188|gb|AAL15328.1 AT5g67320/K8K14_4 {Arabidopsis
thaliana}, partial (44%)
Length = 1098
Score = 43.9 bits (102), Expect = 1e-05
Identities = 26/80 (32%), Positives = 39/80 (48%), Gaps = 9/80 (11%)
Frame = +2
Query: 15 KVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMP------ 68
+V C + G LLAS D +W K EHS I +R+SP+ P
Sbjct: 323 EVNCVKWDPTGTLLASCSDDITAKIWSVKQDKYLHDFREHSKEIYTIRWSPTGPGTNNPN 502
Query: 69 ---RLATSSYDKTVRVWDVD 85
LA++S+D TV++WD++
Sbjct: 503 KKLLLASASFDSTVKLWDIE 562
Score = 37.4 bits (85), Expect = 0.001
Identities = 20/58 (34%), Positives = 30/58 (51%)
Frame = +2
Query: 27 LLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSYDKTVRVWDV 84
LLAS D V LW + K +L H + V FSP+ +A+ S DK++ +W +
Sbjct: 512 LLASASFDSTVKLWDIELGKLIHSLNGHRHPVYSVAFSPNGEYIASGSLDKSLHIWSL 685
>BQ147028 similar to GP|16323188|gb| AT5g67320/K8K14_4 {Arabidopsis
thaliana}, partial (14%)
Length = 658
Score = 43.5 bits (101), Expect = 1e-05
Identities = 25/81 (30%), Positives = 39/81 (47%), Gaps = 9/81 (11%)
Frame = +3
Query: 14 NKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPR---- 69
++V C + G LLAS D +W K EHS + +R+SP+ P
Sbjct: 405 SEVNCIKWDPTGSLLASCSDDNTAKIWSMKQDKYIHDFREHSKEVYTIRWSPTGPGTNNP 584
Query: 70 -----LATSSYDKTVRVWDVD 85
LA++S+D TV++WD +
Sbjct: 585 NKKLVLASASFDSTVKLWDAE 647
>BF631811 similar to GP|9759350|dbj WD-repeat protein-like {Arabidopsis
thaliana}, partial (21%)
Length = 575
Score = 43.1 bits (100), Expect = 2e-05
Identities = 20/62 (32%), Positives = 39/62 (62%), Gaps = 1/62 (1%)
Frame = +3
Query: 28 LASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPR-LATSSYDKTVRVWDVDN 86
+ASG D +V +W+ S + L HS + V ++P+ P LA++S D+T+R+W +++
Sbjct: 216 IASGSEDSQVYIWHRSSGEPIEALPGHSGAVNCVSWNPANPHMLASASDDRTIRIWGLNS 395
Query: 87 VS 88
++
Sbjct: 396 LN 401
>TC89149 similar to GP|20466231|gb|AAM20433.1 cell cycle switch protein
{Arabidopsis thaliana}, partial (71%)
Length = 1283
Score = 42.7 bits (99), Expect = 2e-05
Identities = 21/69 (30%), Positives = 37/69 (53%), Gaps = 3/69 (4%)
Frame = +2
Query: 21 FSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATS---SYDK 77
+S D + LASGG+D ++++W S + L EH+ + + +SP L S + D+
Sbjct: 569 WSCDDRELASGGNDNQLLVWNQHSQQPTLRLTEHTAAVKAIAWSPHQSNLLVSGGGTADR 748
Query: 78 TVRVWDVDN 86
+R W+ N
Sbjct: 749 CIRFWNTTN 775
Score = 40.4 bits (93), Expect = 1e-04
Identities = 22/81 (27%), Positives = 45/81 (55%), Gaps = 1/81 (1%)
Frame = +2
Query: 5 ELNSVRASTNKVVCSHFSSDGKLLASGGHDK-KVVLWYTDSLKQKATLEEHSLLITDVRF 63
+LNSV + + + +L+++ G+ + ++++W SL + ATL HS+ + +
Sbjct: 782 QLNSVDTGSQVCNLAWSKNVNELVSTHGYSQNQIMVWKYPSLAKVATLTGHSMRVLYLAM 961
Query: 64 SPSMPRLATSSYDKTVRVWDV 84
SP + T + D+T+R W+V
Sbjct: 962 SPDGQTIVTGAGDETLRFWNV 1024
>TC78591 similar to GP|9759350|dbj|BAB10005.1 WD-repeat protein-like
{Arabidopsis thaliana}, partial (17%)
Length = 916
Score = 42.0 bits (97), Expect = 4e-05
Identities = 22/66 (33%), Positives = 36/66 (54%), Gaps = 2/66 (3%)
Frame = +2
Query: 21 FSSDGKLLASGGHDKKVVLWY--TDSLKQKATLEEHSLLITDVRFSPSMPRLATSSYDKT 78
FS +GK LAS +D+ ++W + + K L H I+ V +SP+ L T +++
Sbjct: 488 FSHNGKYLASASNDQTAIIWEVGVNGVCVKHRLSGHQKPISSVSWSPNDEELLTCGVEES 667
Query: 79 VRVWDV 84
+R WDV
Sbjct: 668 IRRWDV 685
>BG586181 similar to GP|4689316|gb| peroxisomal targeting signal type 2
receptor {Arabidopsis thaliana}, partial (56%)
Length = 804
Score = 42.0 bits (97), Expect = 4e-05
Identities = 26/68 (38%), Positives = 34/68 (49%), Gaps = 2/68 (2%)
Frame = +1
Query: 18 CSHFSSDGKLLASGGHDKKVVLWYTDSLKQK-ATLEEHSLLITDVRFSPSMPRLATS-SY 75
C D ++A+G DK V +W S + A L H + V+FSP + L S SY
Sbjct: 184 CDWNKYDECIIATGSVDKSVKVWDVRSYRVPVAVLNGHGYAVRKVKFSPHVRNLLVSCSY 363
Query: 76 DKTVRVWD 83
D TV VWD
Sbjct: 364 DMTVCVWD 387
Score = 25.4 bits (54), Expect = 4.0
Identities = 13/36 (36%), Positives = 22/36 (61%), Gaps = 1/36 (2%)
Frame = +1
Query: 50 TLEEHSLLITDVRFSPSMPRL-ATSSYDKTVRVWDV 84
T +EH+ + ++P + A++S D TVR+WDV
Sbjct: 22 TFKEHAYCVYSSVWNPRHADVFASASGDCTVRIWDV 129
>BG457241 homologue to PIR|F84600|F8 coatomer alpha subunit [imported] -
Arabidopsis thaliana, partial (15%)
Length = 654
Score = 41.2 bits (95), Expect = 7e-05
Identities = 21/64 (32%), Positives = 33/64 (50%)
Frame = +1
Query: 20 HFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSYDKTV 79
HF + L SGG D K+ +W + TL H I V+F P + ++S D+T+
Sbjct: 244 HFHNSQPLFVSGGDDYKIKVWNYKLHRCLFTLLGHLDYIRTVQFHHESPWIVSASDDQTI 423
Query: 80 RVWD 83
R+W+
Sbjct: 424 RIWN 435
Score = 35.4 bits (80), Expect = 0.004
Identities = 17/67 (25%), Positives = 32/67 (47%)
Frame = +1
Query: 21 FSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSYDKTVR 80
F + + S D+ + +W S + L H+ + F P + ++S D+TVR
Sbjct: 373 FHHESPWIVSASDDQTIRIWNWQSRTCISVLTGHNHYVMCALFHPKDDLVVSASLDQTVR 552
Query: 81 VWDVDNV 87
VWD+ ++
Sbjct: 553 VWDIGSL 573
Score = 25.4 bits (54), Expect = 4.0
Identities = 13/42 (30%), Positives = 21/42 (49%)
Frame = +1
Query: 16 VVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLL 57
V+C+ F L+ S D+ V +W SLK+K+ +L
Sbjct: 484 VMCALFHPKDDLVVSASLDQTVRVWDIGSLKRKSASXADDIL 609
>BG648991 similar to PIR|C86239|C86 protein T10O24.21 [imported] -
Arabidopsis thaliana, partial (35%)
Length = 702
Score = 40.8 bits (94), Expect = 9e-05
Identities = 20/66 (30%), Positives = 34/66 (51%), Gaps = 1/66 (1%)
Frame = +3
Query: 21 FSSDGKLLASGGHDKKVVLWYT-DSLKQKATLEEHSLLITDVRFSPSMPRLATSSYDKTV 79
F + G L+ S G D KV +W ++ K T HS + D+ F+ + ++ YDK +
Sbjct: 354 FPNSGHLILSAGMDTKVKIWDVFNTGKCMRTYMGHSKAVRDICFTNDGTKFLSAGYDKNI 533
Query: 80 RVWDVD 85
+ WD +
Sbjct: 534 KYWDTE 551
Score = 28.5 bits (62), Expect = 0.47
Identities = 17/68 (25%), Positives = 32/68 (47%), Gaps = 3/68 (4%)
Frame = +3
Query: 21 FSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPR---LATSSYDK 77
F++DG S G+DK + W T++ + +T + V+ +P + L DK
Sbjct: 483 FTNDGTKFLSAGYDKNIKYWDTETGQVISTFSTGKIPYV-VKLNPDEDKQNVLLAGMSDK 659
Query: 78 TVRVWDVD 85
+ WD++
Sbjct: 660 KIVQWDMN 683
>TC84162 similar to GP|15810485|gb|AAL07130.1 putative WD40-repeat protein
{Arabidopsis thaliana}, partial (29%)
Length = 628
Score = 40.4 bits (93), Expect = 1e-04
Identities = 20/62 (32%), Positives = 30/62 (48%)
Frame = +1
Query: 27 LLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSYDKTVRVWDVDN 86
L+ SG D+ +W L + H I V FSP + T+S DKT+R+W + +
Sbjct: 28 LVCSGSQDRTACVWRLPDLVSVVVFKGHKRGIWSVEFSPVDQCVVTASGDKTIRIWAISD 207
Query: 87 VS 88
S
Sbjct: 208GS 213
Score = 32.3 bits (72), Expect = 0.033
Identities = 21/77 (27%), Positives = 32/77 (41%)
Frame = +1
Query: 6 LNSVRASTNKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSP 65
L + T+ V+ + F + G + S G D V LW S + AT + H + +
Sbjct: 217 LKTFEGHTSSVLRALFVTRGTQIVSCGADGLVKLWTVKSNECVATYDNHEDKVWALAVGR 396
Query: 66 SMPRLATSSYDKTVRVW 82
LAT D V +W
Sbjct: 397 KTETLATGGSDAVVNLW 447
>TC77248 similar to GP|13489182|gb|AAK27816.1 putative WD-repeat containing
protein {Oryza sativa}, partial (92%)
Length = 2005
Score = 40.4 bits (93), Expect = 1e-04
Identities = 25/84 (29%), Positives = 42/84 (49%), Gaps = 3/84 (3%)
Frame = +1
Query: 2 TFSELNSV---RASTNKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLI 58
T+++++S + S ++ L+A+G D V++ S + ATL HS +
Sbjct: 718 TYTQISSHPLNKTSKQGIIALDILHSKDLIATGNIDTNAVIFDRPSGQVLATLTGHSKKV 897
Query: 59 TDVRFSPSMPRLATSSYDKTVRVW 82
T V+F + T S DKTVR+W
Sbjct: 898 TSVKFVGQGESIITGSADKTVRLW 969
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.315 0.128 0.369
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,722,444
Number of Sequences: 36976
Number of extensions: 29246
Number of successful extensions: 328
Number of sequences better than 10.0: 128
Number of HSP's better than 10.0 without gapping: 258
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 317
length of query: 88
length of database: 9,014,727
effective HSP length: 64
effective length of query: 24
effective length of database: 6,648,263
effective search space: 159558312
effective search space used: 159558312
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 51 (24.3 bits)
Medicago: description of AC146722.4


