

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146721.7 - phase: 0
(188 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
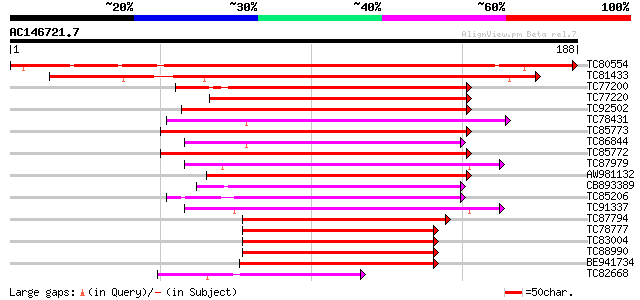
Score E
Sequences producing significant alignments: (bits) Value
TC80554 similar to GP|13430400|gb|AAK25822.1 bZip transcription ... 222 8e-59
TC81433 similar to GP|13430400|gb|AAK25822.1 bZip transcription ... 166 6e-42
TC77200 similar to GP|22597162|gb|AAN03468.1 bZIP transcription ... 80 5e-16
TC77220 similar to GP|22597162|gb|AAN03468.1 bZIP transcription ... 79 1e-15
TC92502 weakly similar to GP|22597162|gb|AAN03468.1 bZIP transcr... 79 1e-15
TC78431 similar to GP|16797791|gb|AAL27150.1 bZIP transcription ... 70 5e-13
TC85773 weakly similar to GP|9650828|emb|CAC00658.1 common plant... 69 1e-12
TC86844 similar to GP|16797791|gb|AAL27150.1 bZIP transcription ... 69 1e-12
TC85772 weakly similar to GP|9650828|emb|CAC00658.1 common plant... 69 1e-12
TC87979 weakly similar to GP|10954097|gb|AAG25728.1 bZIP protein... 65 2e-11
AW981132 similar to GP|4457221|gb|A putative bZIP DNA-binding pr... 65 2e-11
CB893389 similar to PIR|T48034|T480 bZIP transcription factor-li... 62 1e-10
TC85206 similar to GP|16580134|gb|AAK92215.1 bZIP transcription ... 61 3e-10
TC91337 weakly similar to GP|13365772|dbj|BAB39174. RISBZ4 {Oryz... 59 9e-10
TC87794 similar to GP|13775107|gb|AAK39130.1 bZIP transcription ... 57 4e-09
TC78777 similar to PIR|T10985|T10985 regulator protein ROM2 - ki... 57 4e-09
TC83004 similar to PIR|T10985|T10985 regulator protein ROM2 - ki... 55 2e-08
TC88990 similar to PIR|T11751|T11751 transcription repressor ROM... 55 2e-08
BE941734 similar to GP|7406677|emb| putative ripening-related bZ... 55 2e-08
TC82668 weakly similar to GP|13346157|gb|AAK19602.1 bZIP protein... 47 4e-06
>TC80554 similar to GP|13430400|gb|AAK25822.1 bZip transcription factor
{Phaseolus vulgaris}, partial (52%)
Length = 839
Score = 222 bits (565), Expect = 8e-59
Identities = 127/195 (65%), Positives = 146/195 (74%), Gaps = 7/195 (3%)
Frame = +1
Query: 1 MLP-SNTSPYLQNYNNMIQNNNISSYQLQKFSNQIYGNYLNTPHQQFPDFNYSPQSSCIS 59
+LP S+ SPY NMIQN I + Q K SNQ YG + P Q DF SP SSCIS
Sbjct: 85 LLPYSDPSPYPPCNYNMIQNT-IQTPQHPKLSNQFYG-FNQNPIQILQDF--SPPSSCIS 252
Query: 60 SNSTSDEADEQNLSLINERKHRRMISNRESARRSRMRKQKHLDELWSQVLWLRNENHQLI 119
SNSTSDEADEQNL LINERKHRRM+SNRESARRSRMRKQK LDELWSQV+WLRNENHQL+
Sbjct: 253 SNSTSDEADEQNLGLINERKHRRMVSNRESARRSRMRKQKQLDELWSQVVWLRNENHQLL 432
Query: 120 EKLNHVSENHDQVVQENAQLKEEALELRQMIKDMQIHSPLIPSFSPLDDT------YLRD 173
+KLN+ E HD+VVQEN QLKE+A ELRQM+ DMQ+HS +P SPL+D ++
Sbjct: 433 DKLNNFCETHDKVVQENVQLKEQASELRQMVCDMQLHSSCLP-LSPLEDVPSITSPNVKS 609
Query: 174 DSSNNSISSSMDLLG 188
DSSN I+SS+DLLG
Sbjct: 610 DSSN**ITSSVDLLG 654
>TC81433 similar to GP|13430400|gb|AAK25822.1 bZip transcription factor
{Phaseolus vulgaris}, partial (68%)
Length = 774
Score = 166 bits (419), Expect = 6e-42
Identities = 93/170 (54%), Positives = 115/170 (66%), Gaps = 7/170 (4%)
Frame = +2
Query: 14 NNMIQNNNISSYQLQKFSNQIYG-NYLNTPHQQFPDFNYSPQSSCISSNST-SDEADEQN 71
N ++ N I +Y L + +Y + H QF SC+SSNST SDEADE
Sbjct: 95 NFVLPQNEIPNYHLSNILGTLPNFHYPSAGHDQFAP------PSCLSSNSTTSDEADELQ 256
Query: 72 LSLINERKHRRMISNRESARRSRMRKQKHLDELWSQVLWLRNENHQLIEKLNHVSENHDQ 131
++I+ERKHRRMISNRESARRSRMRKQKHLDELWSQV+ LR ENH L++KLNHVSE+HD
Sbjct: 257 FNIIDERKHRRMISNRESARRSRMRKQKHLDELWSQVMKLRTENHNLVDKLNHVSESHDT 436
Query: 132 VVQENAQLKEEALELRQMIKDMQIHSPLIPSFS-----PLDDTYLRDDSS 176
VVQENA+LKEE +LRQM+ DMQI + + P + + +DDSS
Sbjct: 437 VVQENARLKEETFDLRQMVADMQIGNSFPCNMEDLCEIPCNTSQHKDDSS 586
>TC77200 similar to GP|22597162|gb|AAN03468.1 bZIP transcription factor ATB2
{Glycine max}, partial (84%)
Length = 1481
Score = 80.1 bits (196), Expect = 5e-16
Identities = 46/98 (46%), Positives = 67/98 (67%)
Frame = +1
Query: 56 SCISSNSTSDEADEQNLSLINERKHRRMISNRESARRSRMRKQKHLDELWSQVLWLRNEN 115
S + NS S+E D Q +L+++RK +RMISNRESARRSRMRKQKHLD+L SQV LR EN
Sbjct: 562 SSLLQNSGSEE-DLQ--ALMDQRKRKRMISNRESARRSRMRKQKHLDDLVSQVSKLRKEN 732
Query: 116 HQLIEKLNHVSENHDQVVQENAQLKEEALELRQMIKDM 153
+++ +N ++ + V EN+ L+ + EL ++ +
Sbjct: 733 QEILTSVNITTQKYLSVEAENSVLRAQMGELSNRLESL 846
>TC77220 similar to GP|22597162|gb|AAN03468.1 bZIP transcription factor ATB2
{Glycine max}, partial (65%)
Length = 1341
Score = 79.0 bits (193), Expect = 1e-15
Identities = 40/87 (45%), Positives = 59/87 (66%)
Frame = +3
Query: 67 ADEQNLSLINERKHRRMISNRESARRSRMRKQKHLDELWSQVLWLRNENHQLIEKLNHVS 126
++E + L+++RK +RMISNRESARRSRMRKQKHLD+L Q+ LRNEN Q++ +N +
Sbjct: 615 SEEDLMLLMDQRKRKRMISNRESARRSRMRKQKHLDDLAVQLSQLRNENQQILTSVNLTT 794
Query: 127 ENHDQVVQENAQLKEEALELRQMIKDM 153
+ V EN+ L+ + EL + +
Sbjct: 795 QRFLAVESENSVLRAQLNELNSRFESL 875
>TC92502 weakly similar to GP|22597162|gb|AAN03468.1 bZIP transcription
factor ATB2 {Glycine max}, partial (59%)
Length = 597
Score = 78.6 bits (192), Expect = 1e-15
Identities = 38/96 (39%), Positives = 66/96 (68%)
Frame = +3
Query: 58 ISSNSTSDEADEQNLSLINERKHRRMISNRESARRSRMRKQKHLDELWSQVLWLRNENHQ 117
+S+ S S ++E L+++RK +R S RESARRSRMRKQKH+D+L ++V LRNEN +
Sbjct: 12 VSTESQSYGSEEDLQRLMDQRKRKRKQSGRESARRSRMRKQKHMDDLIAEVERLRNENSE 191
Query: 118 LIEKLNHVSENHDQVVQENAQLKEEALELRQMIKDM 153
++ ++N ++++ ++ EN L+ + EL Q ++ +
Sbjct: 192 ILTRMNMTTQHYLKIEAENCVLRAQMCELNQRLQSL 299
>TC78431 similar to GP|16797791|gb|AAL27150.1 bZIP transcription factor
{Nicotiana tabacum}, partial (61%)
Length = 1546
Score = 70.1 bits (170), Expect = 5e-13
Identities = 43/120 (35%), Positives = 64/120 (52%), Gaps = 6/120 (5%)
Frame = +1
Query: 53 PQSSCISSNSTSDEADEQNLSLINE------RKHRRMISNRESARRSRMRKQKHLDELWS 106
P +S S + D+ E +I+ ++ RRM+SNRESARRSR RKQ HL EL +
Sbjct: 658 PSTSGSSREQSDDDEAEGETYMIDSTDPTDVKRVRRMLSNRESARRSRRRKQAHLTELET 837
Query: 107 QVLWLRNENHQLIEKLNHVSENHDQVVQENAQLKEEALELRQMIKDMQIHSPLIPSFSPL 166
QV LR EN L+++L V++ + +N LK + LR +K + + +PL
Sbjct: 838 QVSQLRGENSTLVKRLTDVNQKYTDSAVDNRVLKADVETLRAKVKMAEETVKRVTGLNPL 1017
>TC85773 weakly similar to GP|9650828|emb|CAC00658.1 common plant regulatory
factor 7 {Petroselinum crispum}, partial (62%)
Length = 945
Score = 68.6 bits (166), Expect = 1e-12
Identities = 33/103 (32%), Positives = 65/103 (63%)
Frame = +1
Query: 51 YSPQSSCISSNSTSDEADEQNLSLINERKHRRMISNRESARRSRMRKQKHLDELWSQVLW 110
YS +S + ++ + + + +++ +++K +RM SNRESARRSRM+KQ+H+++L +Q+
Sbjct: 187 YSSGTSSLQNSGSEGDHNHVQVNITDQKKRKRMQSNRESARRSRMKKQQHMEDLSNQIEQ 366
Query: 111 LRNENHQLIEKLNHVSENHDQVVQENAQLKEEALELRQMIKDM 153
L+ EN Q+ + ++ + V ENA L+ + EL ++ +
Sbjct: 367 LKKENIQISTNVGVTTQMYLNVESENAILRVQMAELSHRLQSL 495
>TC86844 similar to GP|16797791|gb|AAL27150.1 bZIP transcription factor
{Nicotiana tabacum}, partial (47%)
Length = 1609
Score = 68.6 bits (166), Expect = 1e-12
Identities = 39/98 (39%), Positives = 59/98 (59%), Gaps = 5/98 (5%)
Frame = +1
Query: 59 SSNSTSDEADEQNLSLINE-----RKHRRMISNRESARRSRMRKQKHLDELWSQVLWLRN 113
+S S+ DE + +++ + ++ RRM+SNRESARRSR RKQ HL EL +QV LR
Sbjct: 601 TSGSSDDEEGDGEINMNGDNPTDAKRVRRMLSNRESARRSRRRKQAHLTELETQVSELRG 780
Query: 114 ENHQLIEKLNHVSENHDQVVQENAQLKEEALELRQMIK 151
EN L+++L V++ + +N LK + LR +K
Sbjct: 781 ENSSLLKRLTDVTQKFNNSAVDNRILKADVETLRAKVK 894
>TC85772 weakly similar to GP|9650828|emb|CAC00658.1 common plant regulatory
factor 7 {Petroselinum crispum}, partial (64%)
Length = 1224
Score = 68.6 bits (166), Expect = 1e-12
Identities = 33/103 (32%), Positives = 65/103 (63%)
Frame = +1
Query: 51 YSPQSSCISSNSTSDEADEQNLSLINERKHRRMISNRESARRSRMRKQKHLDELWSQVLW 110
YS +S + ++ + + + +++ +++K +RM SNRESARRSRM+KQ+H+++L +Q+
Sbjct: 676 YSSGTSSLQNSGSEGDHNHVQVNITDQKKRKRMQSNRESARRSRMKKQQHMEDLSNQIEQ 855
Query: 111 LRNENHQLIEKLNHVSENHDQVVQENAQLKEEALELRQMIKDM 153
L+ EN Q+ + ++ + V ENA L+ + EL ++ +
Sbjct: 856 LKKENIQISTNVGVTTQMYLNVESENAILRVQMAELSHRLQSL 984
>TC87979 weakly similar to GP|10954097|gb|AAG25728.1 bZIP protein BZO2H2
{Arabidopsis thaliana}, partial (49%)
Length = 1346
Score = 65.1 bits (157), Expect = 2e-11
Identities = 43/113 (38%), Positives = 61/113 (53%), Gaps = 7/113 (6%)
Frame = +2
Query: 59 SSNSTSDEADE-----QNLSLINERKHRRMISNRESARRSRMRKQKHLDELWSQVLWLRN 113
SS SDE DE Q+ + ++ ++ RR +SNRESARRSR RKQ HL +L QV LR
Sbjct: 494 SSRDPSDEDDEAGPCEQSTNPVDMKRLRRKVSNRESARRSRRRKQAHLADLEVQVEQLRL 673
Query: 114 ENHQLIEKLNHVSENHDQVVQENAQLKEEALELRQMIK--DMQIHSPLIPSFS 164
EN L ++L S+ N LK + LR +K + + +P+F+
Sbjct: 674 ENASLFKQLTDASQQFRDANTNNRVLKSDVEALRAKVKLAEDMVSRGTLPTFN 832
>AW981132 similar to GP|4457221|gb|A putative bZIP DNA-binding protein
{Capsicum chinense}, partial (47%)
Length = 587
Score = 64.7 bits (156), Expect = 2e-11
Identities = 31/88 (35%), Positives = 56/88 (63%)
Frame = +2
Query: 66 EADEQNLSLINERKHRRMISNRESARRSRMRKQKHLDELWSQVLWLRNENHQLIEKLNHV 125
E+ L+++RK++R SNRESA+R RMRK KH+D++ SQ+ L +N +++ +N
Sbjct: 233 ESSGPEKELMDQRKNKRKQSNRESAKRCRMRKHKHVDDMMSQMSQLTKDNSEILNSINIT 412
Query: 126 SENHDQVVQENAQLKEEALELRQMIKDM 153
++++ V EN+ L+ + EL Q + +
Sbjct: 413 TQHYLNVEAENSILRAQIGELSQRFQSL 496
>CB893389 similar to PIR|T48034|T480 bZIP transcription factor-like protein -
Arabidopsis thaliana, partial (35%)
Length = 724
Score = 62.4 bits (150), Expect = 1e-10
Identities = 38/89 (42%), Positives = 51/89 (56%)
Frame = +1
Query: 63 TSDEADEQNLSLINERKHRRMISNRESARRSRMRKQKHLDELWSQVLWLRNENHQLIEKL 122
TS +D N ++ERK +RMISNRESARRSR RKQK L++ + LRNEN +L E +
Sbjct: 166 TSSGSDGVN-GAMDERKRKRMISNRESARRSRERKQKLLEDYQDEANRLRNENRRLSENI 342
Query: 123 NHVSENHDQVVQENAQLKEEALELRQMIK 151
E + N L+ + EL +K
Sbjct: 343 RVREEGFNANEAANGVLRAQTQELTDQLK 429
>TC85206 similar to GP|16580134|gb|AAK92215.1 bZIP transcription factor
BZI-4 {Nicotiana tabacum}, partial (31%)
Length = 1139
Score = 60.8 bits (146), Expect = 3e-10
Identities = 37/99 (37%), Positives = 58/99 (58%)
Frame = +2
Query: 53 PQSSCISSNSTSDEADEQNLSLINERKHRRMISNRESARRSRMRKQKHLDELWSQVLWLR 112
P SSC S++ SD D Q I+ERK +RM+SNRESARRSR+RKQ+ +++L + L+
Sbjct: 482 PVSSC-SASGGSDGMDLQ----IDERKRKRMLSNRESARRSRLRKQQQVEDLTGEAGKLK 646
Query: 113 NENHQLIEKLNHVSENHDQVVQENAQLKEEALELRQMIK 151
EN +L + E + ++ N ++ + EL +
Sbjct: 647 IENDRLARSIKATEEAYLKMEAANDVIRAQTRELEAQFR 763
>TC91337 weakly similar to GP|13365772|dbj|BAB39174. RISBZ4 {Oryza sativa},
partial (38%)
Length = 1112
Score = 59.3 bits (142), Expect = 9e-10
Identities = 42/117 (35%), Positives = 60/117 (50%), Gaps = 11/117 (9%)
Frame = +1
Query: 59 SSNSTSDEADEQNLS--------LINERKHRRMISNRESARRSRMRKQKHLDELWSQVLW 110
+++ +SD +DE N S ++ ++ RR SN ESARRSR RKQ HL EL +QV
Sbjct: 337 TASGSSDPSDEDNESGPCEQITNPVDMKRQRRKDSNCESARRSRWRKQAHLSELEAQVEK 516
Query: 111 LRNENHQLIEKLNHVSENHDQVVQENAQLKEEALELRQMIK---DMQIHSPLIPSFS 164
L+ EN L ++ S+ + N LK + LR +K DM S L S +
Sbjct: 517 LKLENATLYKQFTDTSQQFHEADTNNRVLKSDVEALRAKVKLAEDMVTRSSLTTSLN 687
>TC87794 similar to GP|13775107|gb|AAK39130.1 bZIP transcription factor 2
{Phaseolus vulgaris}, partial (50%)
Length = 1264
Score = 57.0 bits (136), Expect = 4e-09
Identities = 30/69 (43%), Positives = 42/69 (60%)
Frame = +2
Query: 78 RKHRRMISNRESARRSRMRKQKHLDELWSQVLWLRNENHQLIEKLNHVSENHDQVVQENA 137
++ RR SNRESARRSR+RKQ DEL + L EN L +L+ + ++++ ENA
Sbjct: 389 KRQRRKQSNRESARRSRLRKQAECDELAQRAEVLNQENASLRAELSPIKSEYEKIPSENA 568
Query: 138 QLKEEALEL 146
LKE E+
Sbjct: 569 SLKERLGEI 595
>TC78777 similar to PIR|T10985|T10985 regulator protein ROM2 - kidney bean,
partial (84%)
Length = 1822
Score = 57.0 bits (136), Expect = 4e-09
Identities = 30/65 (46%), Positives = 44/65 (67%)
Frame = +1
Query: 78 RKHRRMISNRESARRSRMRKQKHLDELWSQVLWLRNENHQLIEKLNHVSENHDQVVQENA 137
++ RR SNRESARRSR+RKQ +EL +V L E+ L ++N ++EN +++ ENA
Sbjct: 997 KRERRKQSNRESARRSRLRKQAEAEELARKVESLNAESASLRSEINRLAENSERLRMENA 1176
Query: 138 QLKEE 142
LKE+
Sbjct: 1177 ALKEK 1191
>TC83004 similar to PIR|T10985|T10985 regulator protein ROM2 - kidney bean,
partial (78%)
Length = 1250
Score = 55.1 bits (131), Expect = 2e-08
Identities = 31/65 (47%), Positives = 43/65 (65%)
Frame = +1
Query: 78 RKHRRMISNRESARRSRMRKQKHLDELWSQVLWLRNENHQLIEKLNHVSENHDQVVQENA 137
++ RR SNRESARRSR+RKQ +EL +V L EN L ++N ++EN ++ ENA
Sbjct: 778 KRERRKQSNRESARRSRLRKQAEAEELARRVDALTAENLALKSEMNELAENSAKLKIENA 957
Query: 138 QLKEE 142
LKE+
Sbjct: 958 TLKEK 972
>TC88990 similar to PIR|T11751|T11751 transcription repressor ROM1 - kidney
bean, partial (91%)
Length = 1528
Score = 54.7 bits (130), Expect = 2e-08
Identities = 28/65 (43%), Positives = 42/65 (64%)
Frame = +1
Query: 78 RKHRRMISNRESARRSRMRKQKHLDELWSQVLWLRNENHQLIEKLNHVSENHDQVVQENA 137
++ +R SNRESAR+SR+RKQ +EL +V L EN L E+L +SE +++ EN
Sbjct: 991 KRQKRKQSNRESARKSRLRKQAECEELLKRVEALGGENRTLREELQKLSEECEKLTSEND 1170
Query: 138 QLKEE 142
+KE+
Sbjct: 1171 SIKED 1185
>BE941734 similar to GP|7406677|emb| putative ripening-related bZIP protein
{Vitis vinifera}, partial (18%)
Length = 607
Score = 54.7 bits (130), Expect = 2e-08
Identities = 30/66 (45%), Positives = 40/66 (60%)
Frame = +2
Query: 77 ERKHRRMISNRESARRSRMRKQKHLDELWSQVLWLRNENHQLIEKLNHVSENHDQVVQEN 136
ER+ RRMI NRESA RSR RKQ + EL ++V L+ EN +L +K + E V+E
Sbjct: 8 ERRQRRMIKNRESAARSRARKQAYTMELEAEVAKLKEENEELQKKQEEIMELQKNQVKEM 187
Query: 137 AQLKEE 142
L+ E
Sbjct: 188 MNLQRE 205
>TC82668 weakly similar to GP|13346157|gb|AAK19602.1 bZIP protein DPBF4
{Arabidopsis thaliana}, partial (32%)
Length = 973
Score = 47.0 bits (110), Expect = 4e-06
Identities = 32/80 (40%), Positives = 44/80 (55%), Gaps = 11/80 (13%)
Frame = +1
Query: 50 NYSPQSSCISSNSTS-----------DEADEQNLSLINERKHRRMISNRESARRSRMRKQ 98
++ P +S +SS+S S D D +L ER+ RR I NRESA RSR RKQ
Sbjct: 178 SFQPNTSLVSSSSISSLSDAKPGRKRDAPDAYEKAL--ERRLRRKIKNRESAARSRARKQ 351
Query: 99 KHLDELWSQVLWLRNENHQL 118
+ +EL ++V L +N QL
Sbjct: 352 AYHNELVTKVTLLEQQNMQL 411
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.311 0.126 0.352
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,248,786
Number of Sequences: 36976
Number of extensions: 88300
Number of successful extensions: 617
Number of sequences better than 10.0: 85
Number of HSP's better than 10.0 without gapping: 608
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 614
length of query: 188
length of database: 9,014,727
effective HSP length: 91
effective length of query: 97
effective length of database: 5,649,911
effective search space: 548041367
effective search space used: 548041367
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 55 (25.8 bits)
Medicago: description of AC146721.7


