

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146719.12 + phase: 0 /pseudo
(758 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
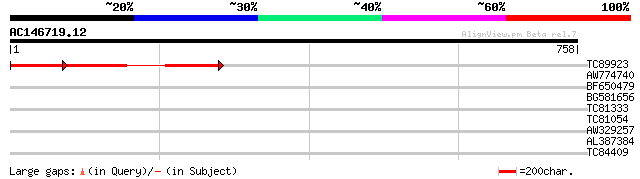
TC89923 weakly similar to GP|12324437|gb|AAG52177.1 hypothetical... 301 e-119
AW774740 34 0.18
BF650479 homologue to PIR|T06643|T066 hypothetical protein T20K1... 34 0.18
BG581656 PIR|A35667|A35 Ty transcription activator TEC1 - yeast ... 34 0.24
TC81333 similar to GP|22776289|dbj|BAC12565. ATP-dependent RNA h... 33 0.31
TC81054 similar to GP|12321513|gb|AAG50816.1 unknown protein {Ar... 31 2.0
AW329257 29 5.9
AL387384 similar to GP|6682235|gb|A unknown protein {Arabidopsis... 29 7.7
TC84409 29 7.7
>TC89923 weakly similar to GP|12324437|gb|AAG52177.1 hypothetical protein;
45165-40041 {Arabidopsis thaliana}, partial (5%)
Length = 783
Score = 301 bits (772), Expect(2) = e-119
Identities = 162/217 (74%), Positives = 163/217 (74%), Gaps = 1/217 (0%)
Frame = +1
Query: 71 ACLQLCIGMINQLFDQYHGLHLAWLPTQGA*LLFAPLMDTLKFIVRPFVIFVLNGLRLWT 130
+CLQLCIGMINQLFDQYHGLHLAWLPTQGA*LLFAPLMDTLKFIVRPFVIFVLNGLRLWT
Sbjct: 280 SCLQLCIGMINQLFDQYHGLHLAWLPTQGA*LLFAPLMDTLKFIVRPFVIFVLNGLRLWT 459
Query: 131 *LKGCMNISDVLSFKAQEGVIH*IFPNRYTCIKATTTKKFPIRWGELHGSNEVIS*TMSV 190
*LKGCMNISDVLSFKAQEGVIH*IFP
Sbjct: 460 *LKGCMNISDVLSFKAQEGVIH*IFPK--------------------------------- 540
Query: 191 YRSMKCR**FGL*LVWFLAIDRVC*KMRQAKWILSHQMMKCQRKCLRANYCH**ALMNML 250
FLAIDRVC*KMRQAKWILSHQMMKCQRKCLRANYCH**ALMNML
Sbjct: 541 ----------------FLAIDRVC*KMRQAKWILSHQMMKCQRKCLRANYCH**ALMNML 672
Query: 251 PAVQCYIRL-LFLGLPCCM*HLNFIPILIEMPLSVFL 286
PAVQCYI L LGLPCCM*HLNFIPILIEMPLSVFL
Sbjct: 673 PAVQCYISLXCXLGLPCCM*HLNFIPILIEMPLSVFL 783
Score = 147 bits (370), Expect(2) = e-119
Identities = 71/77 (92%), Positives = 72/77 (93%)
Frame = +3
Query: 1 MGSQPSPLQPAMLMGSPSFPNALAWSHDNLIAAASGHFVTILRPDLPNGPRGLIKVIPNE 60
MGSQPSPLQPAMLMGSPSFPNALAWSHDNLIAAASGHFVTILRPDLPNGPRGLIKVIPNE
Sbjct: 51 MGSQPSPLQPAMLMGSPSFPNALAWSHDNLIAAASGHFVTILRPDLPNGPRGLIKVIPNE 230
Query: 61 PLILGFIDRKACLQLCI 77
PLILGFIDRK C+
Sbjct: 231 PLILGFIDRKDLHSGCL 281
>AW774740
Length = 739
Score = 34.3 bits (77), Expect = 0.18
Identities = 12/18 (66%), Positives = 15/18 (82%)
Frame = -3
Query: 165 TTTKKFPIRWGELHGSNE 182
TTTK +P +WG LHGSN+
Sbjct: 275 TTTKPYPTKWGRLHGSND 222
>BF650479 homologue to PIR|T06643|T066 hypothetical protein T20K18.200 -
Arabidopsis thaliana, partial (13%)
Length = 580
Score = 34.3 bits (77), Expect = 0.18
Identities = 12/20 (60%), Positives = 15/20 (75%)
Frame = -2
Query: 165 TTTKKFPIRWGELHGSNEVI 184
TTTK +P +WG LHGSN +
Sbjct: 267 TTTKSYPAKWGRLHGSNYAV 208
>BG581656 PIR|A35667|A35 Ty transcription activator TEC1 - yeast
(Saccharomyces cerevisiae), partial (2%)
Length = 795
Score = 33.9 bits (76), Expect = 0.24
Identities = 13/24 (54%), Positives = 16/24 (66%)
Frame = -1
Query: 162 IKATTTKKFPIRWGELHGSNEVIS 185
I TTTK +P +WG LHGS I+
Sbjct: 645 INTTTTKPYPTKWGRLHGSKYAIA 574
>TC81333 similar to GP|22776289|dbj|BAC12565. ATP-dependent RNA helicase
{Oceanobacillus iheyensis}, partial (3%)
Length = 756
Score = 33.5 bits (75), Expect = 0.31
Identities = 18/38 (47%), Positives = 22/38 (57%), Gaps = 7/38 (18%)
Frame = +3
Query: 154 IFPNRY------TCIKATTTKKF-PIRWGELHGSNEVI 184
+ PN Y + + TTT KF P RWG LHGSN +I
Sbjct: 555 LLPNSYEPTRDSSLVYITTTVKFYPTRWGRLHGSNYLI 668
>TC81054 similar to GP|12321513|gb|AAG50816.1 unknown protein {Arabidopsis
thaliana}, partial (32%)
Length = 1050
Score = 30.8 bits (68), Expect = 2.0
Identities = 21/62 (33%), Positives = 31/62 (49%)
Frame = +1
Query: 62 LILGFIDRKACLQLCIGMINQLFDQYHGLHLAWLPTQGA*LLFAPLMDTLKFIVRPFVIF 121
L+L F+ + + + I +IN+L +H L LFA L+ T FI PF IF
Sbjct: 238 LLLLFLFKTLIIIIIIILINKLIHHHHPTITPPLTFSSLIFLFA-LVPTFSFIFHPFSIF 414
Query: 122 VL 123
+L
Sbjct: 415 IL 420
>AW329257
Length = 379
Score = 29.3 bits (64), Expect = 5.9
Identities = 12/20 (60%), Positives = 14/20 (70%)
Frame = -1
Query: 165 TTTKKFPIRWGELHGSNEVI 184
TTT F +WG LHGSN V+
Sbjct: 286 TTT*LFHAKWGRLHGSNYVV 227
>AL387384 similar to GP|6682235|gb|A unknown protein {Arabidopsis thaliana},
partial (3%)
Length = 522
Score = 28.9 bits (63), Expect = 7.7
Identities = 14/29 (48%), Positives = 16/29 (54%), Gaps = 2/29 (6%)
Frame = -1
Query: 165 TTTKKFPIR--WGELHGSNEVIS*TMSVY 191
TTTK +P + WG LHGSN I Y
Sbjct: 378 TTTKPYPTKYKWGRLHGSN*AIGFNHKTY 292
>TC84409
Length = 685
Score = 28.9 bits (63), Expect = 7.7
Identities = 12/20 (60%), Positives = 14/20 (70%)
Frame = -3
Query: 165 TTTKKFPIRWGELHGSNEVI 184
TTTK +P + G LHGSN I
Sbjct: 329 TTTKPYPTKSGRLHGSNYAI 270
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.360 0.162 0.604
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 28,695,476
Number of Sequences: 36976
Number of extensions: 483732
Number of successful extensions: 4579
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 1355
Number of HSP's successfully gapped in prelim test: 203
Number of HSP's that attempted gapping in prelim test: 3174
Number of HSP's gapped (non-prelim): 1667
length of query: 758
length of database: 9,014,727
effective HSP length: 103
effective length of query: 655
effective length of database: 5,206,199
effective search space: 3410060345
effective search space used: 3410060345
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.8 bits)
S2: 62 (28.5 bits)
Medicago: description of AC146719.12


