

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
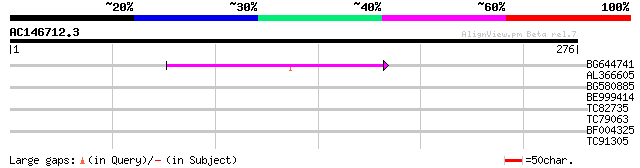
Query= AC146712.3 - phase: 0 /pseudo
(276 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BG644741 45 4e-05
AL366605 40 0.001
BG580885 35 0.024
BE999414 33 0.091
TC82735 weakly similar to GP|6686408|gb|AAF23842.1| F1E22.7 {Ara... 28 2.9
TC79063 similar to GP|15215854|gb|AAK91471.1 At2g43170/F14B2.11 ... 28 2.9
BF004325 weakly similar to GP|11994465|dbj contains similarity t... 27 6.5
TC91305 similar to GP|6175169|gb|AAF04895.1| unknown protein {Ar... 27 8.5
>BG644741
Length = 735
Score = 44.7 bits (104), Expect = 4e-05
Identities = 33/109 (30%), Positives = 46/109 (41%), Gaps = 1/109 (0%)
Frame = -2
Query: 77 LHLFPFSLRDRASAWFHSLEVGSITSWDGMRRAFLARFSPHLKPLNLEIKSHGLIKKMG- 135
L +FP SL A WF L SI +W+ +R FLAR+ P K LN + + + G
Sbjct: 587 LRVFPLSLMGEADIWFTELPYNSIFTWNQLRDVFLARYYPVSKKLNHNDRVNNFVALPGE 408
Query: 136 SLSMKRGRVSRKSLDFVPTMAWKSGL*STPFNGLSYTTKMSVDAAAGGS 184
S+S R + L + G K+ +D AGGS
Sbjct: 407 SVSSSWDRFTSFLRSVPNHRIDDDSLKEYFYRGQDDNNKVVLDTIAGGS 261
>AL366605
Length = 422
Score = 40.0 bits (92), Expect = 0.001
Identities = 23/69 (33%), Positives = 36/69 (51%), Gaps = 2/69 (2%)
Frame = -3
Query: 54 PNLHISSFLRLSGTIKENQEAV*LHLFPFSLRDRASAWFHSLEVGSITSWDGMRRAFLAR 113
P HI ++R G K+N + +H F SL + A+ W+ SL I ++D + AF +
Sbjct: 291 PQNHIIKYVRKMGNYKDNDSLM-IHCFQDSLMEDAAEWYTSLSKNDIHTFDELAAAFKSH 115
Query: 114 --FSPHLKP 120
F+ LKP
Sbjct: 114 YGFNTRLKP 88
>BG580885
Length = 609
Score = 35.4 bits (80), Expect = 0.024
Identities = 19/71 (26%), Positives = 38/71 (52%), Gaps = 2/71 (2%)
Frame = +3
Query: 46 FSGSPTEDPNLHISSFLRLSGTIKENQEAV*LHLFPFSLRDRASAWFH-SLEVGSIT-SW 103
F G+P E P H+S F ++ + + +FP +L + ++ W+ ++E I+ SW
Sbjct: 54 FRGTPNESPITHLSRFNKVCRANNASSVEMQKKIFPVTLEEESALWYDLNIEPYYISLSW 233
Query: 104 DGMRRAFLARF 114
D ++ +FL +
Sbjct: 234 DEIKLSFLQAY 266
>BE999414
Length = 613
Score = 33.5 bits (75), Expect = 0.091
Identities = 23/74 (31%), Positives = 36/74 (48%)
Frame = +3
Query: 42 QQNQFSGSPTEDPNLHISSFLRLSGTIKENQEAV*LHLFPFSLRDRASAWFHSLEVGSIT 101
Q + + G T DP+ H+ + + + T + + AV LF +LR A WF +L SI
Sbjct: 6 QMDSYDG--T*DPDEHMEN-IEVVLTYRSVRGAVKCKLFVTTLRRGAMTWFKNLRRNSIG 176
Query: 102 SWDGMRRAFLARFS 115
SW + F F+
Sbjct: 177 SWGDLCHEFTTHFT 218
>TC82735 weakly similar to GP|6686408|gb|AAF23842.1| F1E22.7 {Arabidopsis
thaliana}, partial (74%)
Length = 965
Score = 28.5 bits (62), Expect = 2.9
Identities = 10/22 (45%), Positives = 17/22 (76%)
Frame = -3
Query: 243 KNSKNLMLMLSHLLLHPLLVKS 264
KNS +L ++ HL+LHP ++K+
Sbjct: 960 KNSMHLKGLIEHLILHPFIIKN 895
>TC79063 similar to GP|15215854|gb|AAK91471.1 At2g43170/F14B2.11 {Arabidopsis
thaliana}, partial (34%)
Length = 1726
Score = 28.5 bits (62), Expect = 2.9
Identities = 15/33 (45%), Positives = 21/33 (63%)
Frame = +2
Query: 229 LNITTLLLRLKL*PKNSKNLMLMLSHLLLHPLL 261
L + LLL L L P+ LM+ L+H++LH LL
Sbjct: 1244 LPVHLLLLVLLLLPRKLSPLMISLTHVVLHQLL 1342
>BF004325 weakly similar to GP|11994465|dbj contains similarity to late
embryogenesis abundant protein~gene_id:MLD14.16
{Arabidopsis thaliana}, partial (8%)
Length = 357
Score = 27.3 bits (59), Expect = 6.5
Identities = 15/49 (30%), Positives = 29/49 (58%)
Frame = +2
Query: 214 PLLLPLRKRQVYTKSLNITTLLLRLKL*PKNSKNLMLMLSHLLLHPLLV 262
PL+ L++ + K I LLR+ *PKN++ ++M+S ++ L++
Sbjct: 170 PLVGKLQQTTLLLKLKEIVR*LLRVSQ*PKNTRGFIIMVSPKMIVVLIL 316
>TC91305 similar to GP|6175169|gb|AAF04895.1| unknown protein {Arabidopsis
thaliana}, partial (34%)
Length = 674
Score = 26.9 bits (58), Expect = 8.5
Identities = 11/21 (52%), Positives = 15/21 (71%)
Frame = -1
Query: 13 SMEEPQAIIVYPTVEGNNFEI 33
++EE +I Y TVEGNNF +
Sbjct: 107 AVEEDGRLIKYGTVEGNNFHV 45
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.325 0.138 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,255,229
Number of Sequences: 36976
Number of extensions: 114564
Number of successful extensions: 753
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 749
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 753
length of query: 276
length of database: 9,014,727
effective HSP length: 95
effective length of query: 181
effective length of database: 5,502,007
effective search space: 995863267
effective search space used: 995863267
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 58 (26.9 bits)
Medicago: description of AC146712.3


