

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146706.4 + phase: 0
(478 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
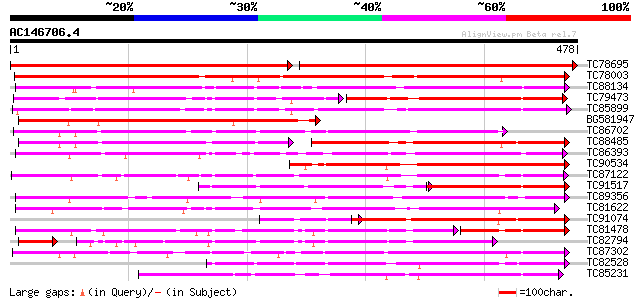
Score E
Sequences producing significant alignments: (bits) Value
TC78695 similar to GP|4115563|dbj|BAA36423.1 UDP-glucose:anthocy... 481 0.0
TC78003 similar to GP|20146093|dbj|BAB88935. glucosyltransferase... 540 e-154
TC88134 weakly similar to SP|Q9MB73|LGT_CITUN Limonoid UDP-gluco... 299 1e-81
TC79473 weakly similar to PIR|T00639|T00639 hypothetical protein... 172 2e-80
TC85899 weakly similar to GP|7385017|gb|AAF61647.1| UDP-glucose:... 295 3e-80
BG581947 similar to GP|20146093|dbj glucosyltransferase NTGT2 {N... 251 4e-67
TC86702 weakly similar to GP|9794913|gb|AAF98390.1| UDP-glucose:... 246 2e-65
TC88485 weakly similar to SP|Q9MB73|LGT_CITUN Limonoid UDP-gluco... 175 4e-63
TC86393 weakly similar to GP|10953887|gb|AAG25643.1 UDP-glucosyl... 197 6e-51
TC90534 weakly similar to GP|19911197|dbj|BAB86925. glucosyltran... 197 7e-51
TC87122 weakly similar to GP|19911203|dbj|BAB86928. glucosyltran... 194 8e-50
TC91517 weakly similar to SP|Q9MB73|LGT_CITUN Limonoid UDP-gluco... 121 3e-46
TC89356 similar to GP|19911207|dbj|BAB86930. glucosyltransferase... 181 5e-46
TC81622 weakly similar to PIR|T00584|T00584 indole-3-acetate bet... 179 2e-45
TC91074 weakly similar to SP|Q9MB73|LGT_CITUN Limonoid UDP-gluco... 169 3e-42
TC81478 weakly similar to GP|21740720|emb|CAD40841. OSJNBa0086B1... 129 4e-41
TC82794 weakly similar to GP|3928543|dbj|BAA34687.1 UDP-glucose ... 148 1e-40
TC87302 similar to GP|19911199|dbj|BAB86926. glucosyltransferase... 161 4e-40
TC82528 weakly similar to PIR|C86356|C86356 hypothetical protein... 159 2e-39
TC85231 similar to GP|13508844|emb|CAC35167. arbutin synthase {R... 156 2e-38
>TC78695 similar to GP|4115563|dbj|BAA36423.1 UDP-glucose:anthocyanin
5-O-glucosyltransferase {Verbena x hybrida}, partial
(49%)
Length = 1741
Score = 481 bits (1237), Expect(2) = 0.0
Identities = 238/238 (100%), Positives = 238/238 (100%)
Frame = +2
Query: 1 MAQNHHFLIITYPLQGHINPALQFTKRLISLGAKVTFATTIHLYSRLINKPTIPGLSFAT 60
MAQNHHFLIITYPLQGHINPALQFTKRLISLGAKVTFATTIHLYSRLINKPTIPGLSFAT
Sbjct: 68 MAQNHHFLIITYPLQGHINPALQFTKRLISLGAKVTFATTIHLYSRLINKPTIPGLSFAT 247
Query: 61 FSDGYDDGQKSFGDEDIVSYMSEFTRRGSEFLTNIILSSKQENHPFTCLIYTLILSWAPK 120
FSDGYDDGQKSFGDEDIVSYMSEFTRRGSEFLTNIILSSKQENHPFTCLIYTLILSWAPK
Sbjct: 248 FSDGYDDGQKSFGDEDIVSYMSEFTRRGSEFLTNIILSSKQENHPFTCLIYTLILSWAPK 427
Query: 121 VAHELHLPSTLLWIQAATVFDIFYYYFHEHGDYITNKSKDETCLISLPGLSFSLKSRDLP 180
VAHELHLPSTLLWIQAATVFDIFYYYFHEHGDYITNKSKDETCLISLPGLSFSLKSRDLP
Sbjct: 428 VAHELHLPSTLLWIQAATVFDIFYYYFHEHGDYITNKSKDETCLISLPGLSFSLKSRDLP 607
Query: 181 SFLLASNTYTFALPSLKEQIQLLNEEINPRVLVNTVEEFELDALNKVDVGKIKMIPIG 238
SFLLASNTYTFALPSLKEQIQLLNEEINPRVLVNTVEEFELDALNKVDVGKIKMIPIG
Sbjct: 608 SFLLASNTYTFALPSLKEQIQLLNEEINPRVLVNTVEEFELDALNKVDVGKIKMIPIG 781
Score = 479 bits (1232), Expect(2) = 0.0
Identities = 234/234 (100%), Positives = 234/234 (100%)
Frame = +1
Query: 245 FLDGKDPTDNSFGGDVVRVDSKDDYIQWLDSKDEKSVVYVSFGTLAVLSKRQMEEIARAL 304
FLDGKDPTDNSFGGDVVRVDSKDDYIQWLDSKDEKSVVYVSFGTLAVLSKRQMEEIARAL
Sbjct: 802 FLDGKDPTDNSFGGDVVRVDSKDDYIQWLDSKDEKSVVYVSFGTLAVLSKRQMEEIARAL 981
Query: 305 LDSGFSFLWVIRDKKLQQQKEEEVDDDELSCREELENNMNGKIVKWCSQVEVLSHRSLGC 364
LDSGFSFLWVIRDKKLQQQKEEEVDDDELSCREELENNMNGKIVKWCSQVEVLSHRSLGC
Sbjct: 982 LDSGFSFLWVIRDKKLQQQKEEEVDDDELSCREELENNMNGKIVKWCSQVEVLSHRSLGC 1161
Query: 365 FMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTGLRMEHDEEGMVKVEEIR 424
FMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTGLRMEHDEEGMVKVEEIR
Sbjct: 1162 FMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTGLRMEHDEEGMVKVEEIR 1341
Query: 425 KCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSSNRNLRSYLNDIACITQI 478
KCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSSNRNLRSYLNDIACITQI
Sbjct: 1342 KCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSSNRNLRSYLNDIACITQI 1503
>TC78003 similar to GP|20146093|dbj|BAB88935. glucosyltransferase NTGT2
{Nicotiana tabacum}, partial (50%)
Length = 1740
Score = 540 bits (1391), Expect = e-154
Identities = 279/475 (58%), Positives = 342/475 (71%), Gaps = 7/475 (1%)
Frame = +3
Query: 5 HHFLIITYPLQGHINPALQFTKRLISLGAKVTFATTIHLYSRLINKPTIPGLSFATFSDG 64
H L++ YP+QGHINPA +F KRLI+LGA VT +TT+H+++R+ NKPT+P LS+ FSDG
Sbjct: 147 HRILLVPYPVQGHINPAFEFAKRLITLGAHVTISTTVHMHNRITNKPTLPNLSYYPFSDG 326
Query: 65 YDDGQKSFGDEDIVSYMSEFTRRGSEFLTNIILSSKQENHPFTCLIYTLILSWAPKVAHE 124
YDDG K G + + Y +EF RRGSEF+++IIL + QE PFTCL+++L+L WA + A E
Sbjct: 327 YDDGFKGTGSDAYLEYHAEFQRRGSEFVSDIILKNSQEGTPFTCLVHSLLLQWAAEAARE 506
Query: 125 LHLPSTLLWIQAATVFDIFYYYFHEHGDYITNKSKDETCLISLPGLSFSLKSRDLPSFLL 184
HLP+ LLW+Q ATVFDI YYYFH D I N S I LPGL SRDLPSFLL
Sbjct: 507 FHLPTALLWVQPATVFDILYYYFHGFSDSIKNPSSS----IELPGLPLLFSSRDLPSFLL 674
Query: 185 AS--NTYTFALPSLKEQIQLLNEEIN--PRVLVNTVEEFELDALNKVDVGKIKMIPIGPL 240
AS + Y+ +EQ L+ E N +LVN+ E E AL V K MI IGPL
Sbjct: 675 ASCPDAYSLMTSFFEEQFNELDVETNLTKTILVNSFESLEPKALRAVK--KFNMISIGPL 848
Query: 241 IPSAFLDGKDPT-DNSFGGDVVRVDSKDDYIQWLDSKDEKSVVYVSFGTLAVLSKRQMEE 299
IPS LD KD T DNS+GG +D ++WLDSK + SVVYVSFG+ VLS+RQ EE
Sbjct: 849 IPSEHLDEKDSTEDNSYGGQTHIFQPSNDCVEWLDSKPKSSVVYVSFGSYFVLSERQREE 1028
Query: 300 IARALLDSGFSFLWVIRDKKLQQQKEEEVDDDELSCREELENNMNGKIVKWCSQVEVLSH 359
IA ALLD GF FLWV+R +KE E +++ REELE GKIVKWCSQ+E+LSH
Sbjct: 1029IAHALLDCGFPFLWVLR------EKEGENNEEGFKYREELEE--KGKIVKWCSQMEILSH 1184
Query: 360 RSLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTGLRMEHD--EEGM 417
SLGCF+THCGWNSTLESL GVPMVAFPQWTDQ TNAKLIEDVWK G+R++ + E+G+
Sbjct: 1185PSLGCFLTHCGWNSTLESLVKGVPMVAFPQWTDQMTNAKLIEDVWKIGVRVDEEVNEDGI 1364
Query: 418 VKVEEIRKCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSSNRNLRSYLNDI 472
V+ +EIR+CLEVVMG GEKGEELRR+ KKWK+LAR AVKEGGSS +NLRS+L+ +
Sbjct: 1365VRGDEIRRCLEVVMGSGEKGEELRRSGKKWKELAREAVKEGGSSEKNLRSFLDGV 1529
>TC88134 weakly similar to SP|Q9MB73|LGT_CITUN Limonoid
UDP-glucosyltransferase (EC 2.4.1.210) (Limonoid
glucosyltransferase) (Limonoid GTase), partial (27%)
Length = 1669
Score = 299 bits (766), Expect = 1e-81
Identities = 174/480 (36%), Positives = 275/480 (57%), Gaps = 13/480 (2%)
Frame = +2
Query: 6 HFLIITYPLQGHINPALQFTKRLISLGAKVTFATTIHLYSRLINKPT-----IP------ 54
H L++ + QGHINP L+ K L++ G VT ATT +Y R+ T +P
Sbjct: 74 HVLLVAFSAQGHINPLLRLGKSLLTKGLNVTLATTELVYHRVFKSTTTDTDTVPTSYSTN 253
Query: 55 GLSFATFSDGYDDGQKSFGDEDIVSYMSEFTRRGSEFLTNIILSSKQEN--HPFTCLIYT 112
G++ FSDG+D Q G + YM + G LTN+I ++ N C+I
Sbjct: 254 GINVLFFSDGFDISQ---GHKSPDEYMELIAKFGPISLTNLINNNFINNPSKKLACIINN 424
Query: 113 LILSWAPKVAHELHLPSTLLWIQAATVFDIFYYYFHEHGDYITNKSKDETCLISLPGLSF 172
+ W VA+EL++P LWIQ T++ I+Y +++E + T ++ + + LPGL
Sbjct: 425 PFVPWVANVAYELNIPCACLWIQPCTLYSIYYRFYNELNQFPTLENPEID--VELPGLPL 598
Query: 173 SLKSRDLPSFLLASNTYTFALPSLKEQIQLLNEEINPRVLVNTVEEFELDALNKVDVGKI 232
LK +DLPSF+L SN L E Q + + VL N+ E E D ++ +
Sbjct: 599 -LKPQDLPSFILPSNPIKTMSDVLAEMFQDMKKL--KWVLANSFYELEKDVIDSM-AEIF 766
Query: 233 KMIPIGPLIPSAFLDGKDPTDNSFGGDVVRVDSKDDYIQWLDSKDEKSVVYVSFGTLAVL 292
+ P+GPL+P + L G+D + + +D ++WL+ K SV+Y+SFG+L L
Sbjct: 767 PITPVGPLVPPSLL-GQDQKQDV---GIEMWTPQDSCMEWLNQKLPSSVIYISFGSLIFL 934
Query: 293 SKRQMEEIARALLDSGFSFLWVIRDKKLQQQKEEEVDDDELSCREELENNMNGKIVKWCS 352
S++QM+ IA AL +S FLWV++ + +K + LS + E G +V WC
Sbjct: 935 SEKQMQSIASALKNSNKYFLWVMKSNDIGAEKVQ------LSEKFLEETKEKGLVVTWCP 1096
Query: 353 QVEVLSHRSLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTGLRMEH 412
Q +VL H ++ CF+THCGWNSTLE++ +GVPM+ +PQWTDQ TNA L+ DV++ G+R++
Sbjct: 1097QTKVLVHPAIACFLTHCGWNSTLEAITAGVPMIGYPQWTDQPTNATLVSDVFRMGIRLKQ 1276
Query: 413 DEEGMVKVEEIRKCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSSNRNLRSYLNDI 472
D +G V+ +E+ + +E ++G GE+ E ++NA + K AR AV +GGSS+RN++ ++++I
Sbjct: 1277DNDGFVESDEVERAIEEIVG-GERSEVFKKNALELKHAAREAVADGGSSDRNIQRFVDEI 1453
>TC79473 weakly similar to PIR|T00639|T00639 hypothetical protein F3I6.2 -
Arabidopsis thaliana, partial (45%)
Length = 1530
Score = 172 bits (435), Expect(2) = 2e-80
Identities = 86/186 (46%), Positives = 126/186 (67%)
Frame = +1
Query: 285 SFGTLAVLSKRQMEEIARALLDSGFSFLWVIRDKKLQQQKEEEVDDDELSCREELENNMN 344
SFG++ L+ Q+EE+A L +S +FLWV+R+ +Q K + D + +
Sbjct: 868 SFGSMVSLTSEQIEELALGLKESEVNFLWVLRES--EQGKLPKGYKDFIKEK-------- 1017
Query: 345 GKIVKWCSQVEVLSHRSLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVW 404
G IV WC+Q+E+L+H ++GCF+THCGWNSTLESL GVP+V PQW DQ +AK +E++W
Sbjct: 1018 GIIVTWCNQLELLAHDAVGCFVTHCGWNSTLESLSLGVPVVCLPQWADQLPDAKFLEEIW 1197
Query: 405 KTGLRMEHDEEGMVKVEEIRKCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSSNRN 464
+ G+R + DE G+VK EE L+VVM + E+ E +RRNA +WK LAR AV EGGS ++N
Sbjct: 1198 EVGVRPKEDENGVVKREEFMLSLKVVM-ESERSEVIRRNASEWKKLARDAVCEGGSFDKN 1374
Query: 465 LRSYLN 470
+ +++
Sbjct: 1375 INQFVD 1392
Score = 145 bits (367), Expect(2) = 2e-80
Identities = 98/281 (34%), Positives = 149/281 (52%), Gaps = 3/281 (1%)
Frame = +2
Query: 4 NHHFLIITYPLQGHINPALQFTKRLISLGAKVTFATTIHLYSRLINKPTIPGLSFATFSD 63
N H L+I YP QGHI+P +QF+KRL+S G K TFATT + + T P +S SD
Sbjct: 68 NVHVLVIPYPAQGHISPLIQFSKRLVSKGIKTTFATTHY----TVKSITAPNISVEPISD 235
Query: 64 GYDDGQKSFGDEDIVSYMSEFTRRGSEFLTNIILSSKQENHPFTCLIYTLILSWAPKVAH 123
G+D+ S +++ +++ F GS+ L+N+I ++ + P TC++Y L WA VA
Sbjct: 236 GFDESGFSQA-KNVELFLNSFKTNGSKTLSNLIQKHQKTSTPITCIVYDSFLPWALDVAK 412
Query: 124 ELHLPSTLLWIQAATVFDIFYYYFHEHGDYITNKSKDETCLISLPGLSFSLKSRDLPSFL 183
+ + + +A V +IF H + DE LI +PGL L SRDLPSF+
Sbjct: 413 QHRIYGAAFFTNSAAVCNIFCRIHHG----LIETPVDELPLI-VPGLP-PLNSRDLPSFI 574
Query: 184 LASNTYTFALPSLKEQIQLLNEEINPRVLVNTVEEFELDALNKVDVGKIKMIP---IGPL 240
+Y + Q LN+ + VNT E E + + G +M P IGP+
Sbjct: 575 RFPESYPAYMAMKLNQFSNLNQA--DWMFVNTFEALEAEVVK----GLTEMFPAKLIGPM 736
Query: 241 IPSAFLDGKDPTDNSFGGDVVRVDSKDDYIQWLDSKDEKSV 281
+PSA+LDG+ D +G ++ + S +D I WL++K +SV
Sbjct: 737 VPSAYLDGRIKGDKGYGANLWKPLS-EDCINWLNAKPSQSV 856
>TC85899 weakly similar to GP|7385017|gb|AAF61647.1| UDP-glucose:salicylic
acid glucosyltransferase {Nicotiana tabacum}, partial
(47%)
Length = 1617
Score = 295 bits (754), Expect = 3e-80
Identities = 178/477 (37%), Positives = 266/477 (55%), Gaps = 6/477 (1%)
Frame = +2
Query: 3 QNH--HFLIITYPLQGHINPALQFTKRLISLGAKVTFATTIHLYSRLINKPTIPGLSFAT 60
+NH H LI+ YP QGH+NP +QF+KRLI G K+T T + + NK + + +
Sbjct: 29 KNHAPHCLILPYPAQGHMNPMIQFSKRLIEKGVKITLITVTSFWKVISNK-NLTSIDVES 205
Query: 61 FSDGYDDGQKSFGDEDIVSYMSEFTRRGSEFLTNIILSSKQENHPFTCLIYTLILSWAPK 120
SDGYD+G E + Y F + GS+ L+ ++ +P C+I+ L W
Sbjct: 206 ISDGYDEGGL-LAAESLEDYKETFWKVGSQTLSELLHKLSSSENPPNCVIFDAFLPWVLD 382
Query: 121 VAHELHLPSTLLWIQAATVFDIFYYYFHEHGDYITNKSKDETCLISLPGLSFSLKSRDLP 180
V L + Q+ +V ++Y H H I L LPGL L DLP
Sbjct: 383 VGKSFGLVGVAFFTQSCSVNSVYY---HTHEKLIELPLTQSEYL--LPGLP-KLAPGDLP 544
Query: 181 SFLLASNTYTFALPSLKEQIQLLNEEINPRVLVNTVEEFELDALNKVDVGKIKMIP---I 237
SFL +Y + Q + + +L N++ E E + ++ + +K+ P I
Sbjct: 545 SFLYKYGSYPGYFDIVVNQFANIGKA--DWILANSIYELEPEVVDWL----VKIWPLKTI 706
Query: 238 GPLIPSAFLDGKDPTDNSFGGDVVRVDSKDDY-IQWLDSKDEKSVVYVSFGTLAVLSKRQ 296
GP +PS LD + D +G V D ++ I+WL+ K + SVVY SFG++A LS+ Q
Sbjct: 707 GPSVPSMLLDKRLKDDKEYG--VSLSDPNTEFCIKWLNDKPKGSVVYASFGSMAGLSEEQ 880
Query: 297 MEEIARALLDSGFSFLWVIRDKKLQQQKEEEVDDDELSCREELENNMNGKIVKWCSQVEV 356
+E+A L DS FLWV+R E D +L + +E++ G IV WC Q+ V
Sbjct: 881 TQELALGLKDSESYFLWVVR----------ECDQSKLP-KGFVESSKKGLIVTWCPQLLV 1027
Query: 357 LSHRSLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTGLRMEHDEEG 416
L+H +LGCF+THCGWNSTLE+L GVP++A P WTDQ TNAKLI DVWK G+R DE+
Sbjct: 1028LTHEALGCFVTHCGWNSTLEALSIGVPLIAMPLWTDQVTNAKLIADVWKMGVRAVADEKE 1207
Query: 417 MVKVEEIRKCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSSNRNLRSYLNDIA 473
+V+ E I+ C++ ++ + EKG E+++NA KWK+LA+++V EGG S++N+ ++ +A
Sbjct: 1208IVRSETIKNCIKEII-ETEKGNEIKKNALKWKNLAKSSVDEGGRSDKNIEEFVAALA 1375
>BG581947 similar to GP|20146093|dbj glucosyltransferase NTGT2 {Nicotiana
tabacum}, partial (8%)
Length = 786
Score = 251 bits (641), Expect = 4e-67
Identities = 136/265 (51%), Positives = 175/265 (65%), Gaps = 10/265 (3%)
Frame = +2
Query: 8 LIITYPLQGHINPALQFTKRLISLGAKVTFATTIHLYSRLI----NKPTIPGLSFATFSD 63
L++TYP QGHINPALQF KRLIS+GA VT T+HLY RLI + TI LS FSD
Sbjct: 5 LLVTYPAQGHINPALQFAKRLISMGAHVTLPITLHLYRRLILLNPSITTISNLSITPFSD 184
Query: 64 GYDDGQKSFG--DEDIVSYMSEFTRRGSEFLTNIILSSKQENHPFTCLIYTLILSWAPKV 121
GY+DG + D D Y S+F RGS+F+TN+ILS+KQE+ PFTCL+YT+I+ WAP+V
Sbjct: 185 GYNDGFIAITNTDADFHQYTSQFNTRGSDFITNLILSAKQESKPFTCLLYTIIIPWAPRV 364
Query: 122 AHELHLPSTLLWIQAATVFDIFYYYFHEHGDYITNKSKDET-CLISLPGLSFSLKSRDLP 180
A +L S LWI+ ATVFDI YYYFH + ++I N+++++ I LPGL F+L RD+P
Sbjct: 365 ARGFNLRSAKLWIEPATVFDILYYYFHGYSNHINNQNQNQNQTTIELPGLPFTLSPRDIP 544
Query: 181 SFLLASN--TYTFALPSLKEQIQLLNEEINPRVLVNTVEEFELDALNKVDV-GKIKMIPI 237
SFL SN +F P ++ L+ E NP +LVNT E E +AL VD +KMIPI
Sbjct: 545 SFLFTSNPSVLSFVFPYFQQDFHELDVETNPIILVNTFEALEPEALRAVDTHHNLKMIPI 724
Query: 238 GPLIPSAFLDGKDPTDNSFGGDVVR 262
GPLIPS D SF GD+++
Sbjct: 725 GPLIPS---------DTSFSGDLLQ 772
>TC86702 weakly similar to GP|9794913|gb|AAF98390.1| UDP-glucose:sinapate
glucosyltransferase {Brassica napus}, partial (34%)
Length = 1366
Score = 246 bits (627), Expect = 2e-65
Identities = 157/428 (36%), Positives = 229/428 (52%), Gaps = 12/428 (2%)
Frame = +3
Query: 4 NHHFLIITYPLQGHINPALQFTKRLISLGAKVTFATT------IHLYSRLINKPTIP--- 54
N H L+I+YP QGHINP L+ K L + GA V F TT I + +I K
Sbjct: 72 NTHILLISYPAQGHINPLLRLAKCLAAKGASVIFITTEKAGKDIRTVNNIIEKSLTSIGD 251
Query: 55 -GLSFATFSDGYDDGQKSFGDEDIVSYMSEFTRRGSEFLTNIILSSKQENHPFTCLIYTL 113
L+F DG +D G+ ++ Y+ G F++ +I + N PF+C+I
Sbjct: 252 GSLTFEFLDDGLEDDHPLRGN--LLGYIEHLKLVGKPFVSQMIKNHADSNKPFSCIINNP 425
Query: 114 ILSWAPKVAHELHLPSTLLWIQAATVFDIFYYYFHEHGDYITNKSKDETCL-ISLPGLSF 172
+ W VA E +PS LLWIQ+ VF +Y YFH+ + S+ E + + LP
Sbjct: 426 FVPWVCDVADEHGIPSALLWIQSTAVFSAYYNYFHK---LVRFPSQTEPYIDVQLPFQV- 593
Query: 173 SLKSRDLPSFLLASNTYTFALPSLKEQIQLLNEEINPRVLVNTVEEFELDALNKVDVGKI 232
LK ++P FL + + F + EQ + L++ VLV T EE E D ++ + I
Sbjct: 594 -LKYNEIPDFLHPFSQFPFLGTLILEQFKNLSKVFC--VLVETYEELEHDFIDYISKKTI 764
Query: 233 KMIPIGPLIPSAFLDGKDPTDNSFGGDVVRVDSKDDYIQWLDSKDEKSVVYVSFGTLAVL 292
+ PIGPL + + G + GD V+ D + I+WL +K + SVVY+SFG++ +
Sbjct: 765 FVRPIGPLFHNPNIKGA----KNIRGDFVKSDDCN-IIEWLSTKPKGSVVYISFGSIVYI 929
Query: 293 SKRQMEEIARALLDSGFSFLWVIRDKKLQQQ-KEEEVDDDELSCREELENNMNGKIVKWC 351
+ Q+ EIA LLDS SFLWV++ + KE + D L E GK+V W
Sbjct: 930 PQEQVNEIAYGLLDSQVSFLWVLKPPSEELGLKEHNLPDGFLEGISE-----RGKVVNWS 1094
Query: 352 SQVEVLSHRSLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTGLRME 411
Q EVL+H S+ CF+THCGWNS++E+L GVPM+ FP W DQ TNAK + DV+ G+R+
Sbjct: 1095PQEEVLAHPSVACFITHCGWNSSMEALSLGVPMLTFPAWGDQVTNAKFLVDVFGVGIRLG 1274
Query: 412 HDEEGMVK 419
+ GM++
Sbjct: 1275Y---GMIE 1289
>TC88485 weakly similar to SP|Q9MB73|LGT_CITUN Limonoid
UDP-glucosyltransferase (EC 2.4.1.210) (Limonoid
glucosyltransferase) (Limonoid GTase), partial (39%)
Length = 1544
Score = 175 bits (444), Expect(2) = 4e-63
Identities = 96/221 (43%), Positives = 135/221 (60%), Gaps = 3/221 (1%)
Frame = +2
Query: 255 SFGGDVVRVDSKDDYIQWLDSKDEKSVVYVSFGTLAVLSKRQMEEIARALLDSGFSFLWV 314
+F D + + D I+WLD+K + SVVYVSFGTL + QM EI LL+S SFLW
Sbjct: 812 TFMVDFAKSNDDDKIIEWLDTKPKDSVVYVSFGTLVNYPQEQMNEIVYGLLNSQVSFLWS 991
Query: 315 IRDKKLQQQKEEEVDDDELSCREELENNMNGKIVKWCSQVEVLSHRSLGCFMTHCGWNST 374
+ + + + DD L E N GK+V+W QV+VL+H S+ CF+THCGWNS+
Sbjct: 992 LSNPGV-------LPDDFLE-----ETNERGKVVEWSPQVDVLAHPSVACFITHCGWNSS 1135
Query: 375 LESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTGLRMEHD---EEGMVKVEEIRKCLEVVM 431
+E+L GVP++ FP DQ TNAK + DV+ G++M +V +E++KCL +
Sbjct: 1136 IEALSLGVPVLTFPSRGDQLTNAKFLVDVFGVGIKMGSGWAVNNKLVTRDEVKKCL-LEA 1312
Query: 432 GKGEKGEELRRNAKKWKDLARAAVKEGGSSNRNLRSYLNDI 472
GEK E+L++NA KWK A A+ GGSS+RNL ++ DI
Sbjct: 1313 TIGEKAEKLKQNANKWKKKAEEALAVGGSSDRNLDEFMEDI 1435
Score = 84.7 bits (208), Expect(2) = 4e-63
Identities = 67/243 (27%), Positives = 114/243 (46%), Gaps = 11/243 (4%)
Frame = +1
Query: 8 LIITYPLQGHINPALQFTKRLISLGAKVTFATT------IHLYSRLINKPTIP----GLS 57
L++++P QGHIN + K L + GA V F TT + + +I+K P +
Sbjct: 73 LLVSFPAQGHINHLVGLGKYLAAKGATVIFTTTETAGKNMRAANNIIDKLATPIGDGTFA 252
Query: 58 FATFSDGYDDGQKSFGDEDIVSYMSEFTRRGSEFLTNIILSSKQENHPFTCLIYTLILSW 117
F F DG DG +S + + +E G ++ +I + N PF+C+I W
Sbjct: 253 FEFFDDGLPDGDRSAFRA--LQHSAEIEVAGRPSISQMIKNHADLNKPFSCIINNYFFPW 426
Query: 118 APKVAHELHLPSTLLWIQAATVFDIFYYYFHEHGDYITNKSK-DETCLISLPGLSFSLKS 176
VA+E ++PS L W +A VF +Y Y H+ + TN+ + LI S LK
Sbjct: 427 VCDVANEHNIPSVLSWTNSAAVFTTYYNYVHKLTPFPTNEEPYIDVQLIP----SRVLKY 594
Query: 177 RDLPSFLLASNTYTFALPSLKEQIQLLNEEINPRVLVNTVEEFELDALNKVDVGKIKMIP 236
++ + ++ F + E+ + L++ VLV+T EE E + ++ + I +
Sbjct: 595 NEISDLVHPFCSFPFLGKLVLEEFKDLSKVF--CVLVDTYEELEHEFIDYISKKSIPIRT 768
Query: 237 IGP 239
+GP
Sbjct: 769 VGP 777
>TC86393 weakly similar to GP|10953887|gb|AAG25643.1 UDP-glucosyltransferase
HRA25 {Phaseolus vulgaris}, partial (47%)
Length = 1332
Score = 197 bits (502), Expect = 6e-51
Identities = 142/474 (29%), Positives = 231/474 (47%), Gaps = 9/474 (1%)
Frame = +1
Query: 6 HFLIITYPLQGHINPALQFTKRLISLGAKVTFATTIHLYSRLIN--KPTIPGLSFATFSD 63
HFL+I YP+ GHINP +Q L G K+TF T + R N + + ++F T D
Sbjct: 16 HFLVIPYPIPGHINPLMQLCHVLAKHGCKITFLNTEFSHKRTNNNNEQSQETINFVTLPD 195
Query: 64 GYDDGQKSFGDEDIVSYMSEFTRRGSEFLTNII--LSSKQENHPFTCLIYTLILSWAPKV 121
G + + + + R L +I +++ + + C+I T + WA +V
Sbjct: 196 GLEPEDDRSDQKKV---LFSIKRNMPPLLPKLIEEVNALDDENKICCIIVTFNMGWALEV 366
Query: 122 AHELHLPSTLLWIQAATVFDIFYYYFHEHGDYITNKS----KDETCLISLPGLSFSLKSR 177
H L + LLW +AT Y D + + + KD+ +S P + + ++
Sbjct: 367 GHNLGIKGVLLWTGSATSLAFCYSIPKLIDDGVIDSAGIYTKDQEIQLS-PNMP-KMDTK 540
Query: 178 DLPSFLLASNTYTFALPSLKEQIQLLNEEINPRVLVNTVEEFELDALNKVDVGKIKMIPI 237
++P + L +Q+Q + ++ L NT + E + K +PI
Sbjct: 541 NVPWRTFDKIIFDH----LAQQMQTM--KLGHWWLCNTTYDLEHATFSISP----KFLPI 690
Query: 238 GPLIPSAFLDGKDPTDNSFGG-DVVRVDSKDDYIQWLDSKDEKSVVYVSFGTLAVLSKRQ 296
GPL+ + D +SF D+ +D WLD + +SVVYVSFG+LAV+ + Q
Sbjct: 691 GPLMEN------DSNKSSFWQEDMTSLD-------WLDKQPSQSVVYVSFGSLAVMDQNQ 831
Query: 297 MEEIARALLDSGFSFLWVIRDKKLQQQKEEEVDDDELSCREELENNMNGKIVKWCSQVEV 356
E+A L FLWV+R K DE GKIV W Q ++
Sbjct: 832 FNELALGLDLLDKPFLWVVRPSN--DNKVNYAYPDEFL-------GTKGKIVSWVPQKKI 984
Query: 357 LSHRSLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTGLRMEHDEEG 416
L+H ++ CF++HCGWNST+E + SG+P + +P TDQ TN I DVWK G ++ DE G
Sbjct: 985 LNHPAIACFISHCGWNSTIEGVYSGIPFLCWPFATDQFTNKSYICDVWKVGFELDKDENG 1164
Query: 417 MVKVEEIRKCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSSNRNLRSYLN 470
+V EEI+K +E ++ + ++++ + K K+L + E G S++NL++++N
Sbjct: 1165IVLKEEIKKKVEQLL----QDQDIKERSLKLKELTLENIVEDGKSSKNLQNFIN 1314
>TC90534 weakly similar to GP|19911197|dbj|BAB86925. glucosyltransferase-7
{Vigna angularis}, partial (71%)
Length = 808
Score = 197 bits (501), Expect = 7e-51
Identities = 100/241 (41%), Positives = 156/241 (64%), Gaps = 5/241 (2%)
Frame = +3
Query: 237 IGPLIPSAFLDG--KDPTDNSFGGDVVRVDSKDDYIQWLDSKDEKSVVYVSFGTLAVLSK 294
+GP +P FLD KD D+S + ++ S D+ I+WL++K ++S VYVSFG++A L++
Sbjct: 39 VGPCLPYTFLDKRVKDDEDHS----IAQLKS-DESIEWLNNKPKRSAVYVSFGSMASLNE 203
Query: 295 RQMEEIARALLDSGFSFLWVIR---DKKLQQQKEEEVDDDELSCREELENNMNGKIVKWC 351
Q+EE+A L D G FLWV++ + KL + E++ + NG IV WC
Sbjct: 204 EQIEEVAHCLKDCGSYFLWVVKTSEETKLPKDFEKKSE--------------NGLIVAWC 341
Query: 352 SQVEVLSHRSLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTGLRME 411
Q+EVL+H ++GCF+THCGWNSTLE+L GVP+VA P ++DQ +AK + D+WK G+R
Sbjct: 342 PQLEVLAHEAIGCFVTHCGWNSTLEALSIGVPIVAIPLYSDQGIDAKFLVDIWKVGIRPL 521
Query: 412 HDEEGMVKVEEIRKCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSSNRNLRSYLND 471
DE+ +V+ + ++ C+ +M EKG+E+ N +WK LA AV + GSS++N+ ++N
Sbjct: 522 VDEKQIVRKDPLKDCICEIMSMSEKGKEIMNNVMQWKTLATRAVGKDGSSHKNMIEFVNS 701
Query: 472 I 472
+
Sbjct: 702 L 704
>TC87122 weakly similar to GP|19911203|dbj|BAB86928. glucosyltransferase-10
{Vigna angularis}, partial (41%)
Length = 1588
Score = 194 bits (492), Expect = 8e-50
Identities = 149/500 (29%), Positives = 235/500 (46%), Gaps = 30/500 (6%)
Frame = +2
Query: 2 AQNHHFLIITYPLQGHINPALQFTKRLISLGAKVTFATTIHLYSRLINK------PTIPG 55
+Q H + + +P QGH+NP +Q K L LG +TF + RLI P
Sbjct: 116 SQKPHVVCVPFPAQGHVNPFMQLAKILNKLGFHITFVNNEFNHKRLIKSLGHDFLKGQPD 295
Query: 56 LSFATFSDGYDDGQKSFGDEDIVSYMSEFTRRG-----SEFLTNIILSSKQENHPFTCLI 110
F T DG ++ + + E R+ E + N L++ E T +I
Sbjct: 296 FRFETIPDGLPPSDENATQS--IGGLVEGCRKHCYGPLKELVEN--LNNSSEVPQVTSII 463
Query: 111 YTLILSWAPKVAHELHLPSTLLWIQAATVFDIFYYYFHE----------HGDYITNKSKD 160
Y ++ +A VA +L + W +A + Y F E YIT+ S D
Sbjct: 464 YDGLMGFAVDVAKDLGIAEQQFWTASACGL-MGYLQFDELIKRNMLPYKDESYITDGSLD 640
Query: 161 ETCLISLPGLSFSLKSRDLPSFLLASNTYTFALPSLKEQIQLLNEEINPRVLVNTVEEFE 220
L PG+ +++ RDLPSF+ + + + Q + + +++NTV+E E
Sbjct: 641 -VHLDWTPGMK-NIRMRDLPSFVRTTTLDEISFVGFGLEAQQCMK--SSAIIINTVKELE 808
Query: 221 LDALNKVDVGKIKMIPIGPLIPSAFLDGKDPT-DNSFGGDVVRVDSKD-DYIQWLDSKDE 278
+ L+ + + IGPL FL P +N F + + D I+WLD +
Sbjct: 809 SEVLDALMAINPNIYNIGPL---QFLANNFPEKENGFKSNGSSLWKNDLTCIKWLDQWEP 979
Query: 279 KSVVYVSFGTLAVLSKRQMEEIARALLDSGFSFLWVIRD-------KKLQQQKEEEVDDD 331
SV+Y+++G++AV+S++ E A L +S FLW+ R + L Q+ +EV D
Sbjct: 980 SSVIYINYGSIAVMSEKHFNEFAWGLANSKLPFLWINRPDLVKGKCRPLSQEFLDEVKD- 1156
Query: 332 ELSCREELENNMNGKIVKWCSQVEVLSHRSLGCFMTHCGWNSTLESLGSGVPMVAFPQWT 391
G I+ WC Q EVL+H S+G F+THCGWNS+LES+ G PM+ +P +
Sbjct: 1157------------RGYIITWCPQSEVLAHPSVGVFLTHCGWNSSLESISEGKPMIGWPFFA 1300
Query: 392 DQTTNAKLIEDVWKTGLRMEHDEEGMVKVEEIRKCLEVVMGKGEKGEELRRNAKKWKDLA 451
+Q TN + I W G+ ++ D VK EE+ + ++ ++ KGEKG++ R +WK
Sbjct: 1301EQQTNCRYISTTWGIGMDIKDD----VKREEVTELVKEMI-KGEKGKKKREKCVEWKKKV 1465
Query: 452 RAAVKEGGSSNRNLRSYLND 471
A K GGSS + L D
Sbjct: 1466VEAAKPGGSSYNDFYRLLKD 1525
>TC91517 weakly similar to SP|Q9MB73|LGT_CITUN Limonoid
UDP-glucosyltransferase (EC 2.4.1.210) (Limonoid
glucosyltransferase) (Limonoid GTase), partial (31%)
Length = 1027
Score = 121 bits (304), Expect(2) = 3e-46
Identities = 56/121 (46%), Positives = 86/121 (70%)
Frame = +1
Query: 352 SQVEVLSHRSLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTGLRME 411
S +VL+H S+ CF+THCGWNS++E++ GVPM+ FP + DQ TNAK + DV+ G+R+
Sbjct: 523 STEQVLAHSSVACFITHCGWNSSMEAITLGVPMLTFPAFGDQLTNAKFLVDVFGAGIRLG 702
Query: 412 HDEEGMVKVEEIRKCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSSNRNLRSYLND 471
+ ++ +V +E++KCL + GEK E L++NA KWK A A+ GGSS+R+L +++ D
Sbjct: 703 YGDKKLVTRDEVKKCLLEAI-TGEKAERLKQNAMKWKTAAEDAMAVGGSSDRHLDAFIQD 879
Query: 472 I 472
I
Sbjct: 880 I 882
Score = 82.0 bits (201), Expect(2) = 3e-46
Identities = 62/198 (31%), Positives = 99/198 (49%)
Frame = +2
Query: 160 DETCLISLPGLSFSLKSRDLPSFLLASNTYTFALPSLKEQIQLLNEEINPRVLVNTVEEF 219
+E L+SL S LK ++P ++ N + QI+ +++ VLV+T EE
Sbjct: 5 EELTLMSLNS-SIVLKYNEIPDYIHPFNPFPILGTLTTAQIKDMSKVFC--VLVDTFEEL 175
Query: 220 ELDALNKVDVGKIKMIPIGPLIPSAFLDGKDPTDNSFGGDVVRVDSKDDYIQWLDSKDEK 279
E D ++ + I + +GPL + +G N+ GD + + I+WL++K +
Sbjct: 176 EHDFIDYISKKSITIRHVGPLFKNPKANG---ASNNTLGDFTKSNDDSTIIEWLNTKPKG 346
Query: 280 SVVYVSFGTLAVLSKRQMEEIARALLDSGFSFLWVIRDKKLQQQKEEEVDDDELSCREEL 339
SVVY+SFGT+ ++ Q+ EIA LL+S SFLWV+ K+ D L
Sbjct: 347 SVVYISFGTIVNHTQEQVNEIAYGLLNSQVSFLWVL--------KQHVFPDGFLE----- 487
Query: 340 ENNMNGKIVKWCSQVEVL 357
E + GK+VKW Q + L
Sbjct: 488 ETSGRGKVVKWSPQNKYL 541
>TC89356 similar to GP|19911207|dbj|BAB86930. glucosyltransferase-12 {Vigna
angularis}, partial (58%)
Length = 1642
Score = 181 bits (459), Expect = 5e-46
Identities = 151/491 (30%), Positives = 231/491 (46%), Gaps = 24/491 (4%)
Frame = +1
Query: 6 HFLIITYPLQGHINPALQFTKRLISLGAKVTFATTIHLYSRLIN--------KPTIPGLS 57
H LI P QGH+N L+ + L +TF T ++++RLI P L
Sbjct: 64 HVLIFPCPAQGHVNSMLKLAELLAIQNIYITFLNTKYIHNRLIQFNDDIQALLECYPKLQ 243
Query: 58 FATFSDGYDDGQKSFGDEDIVSYMSEFTRRGSEFLTNIILSSKQENHPFTCLIYTLILSW 117
F T SD + + + E I ++ + G L +II+S K +C+I I
Sbjct: 244 FKTISDFHSEEKHPGFGERIGDVITSLSLYGKPLLKDIIVSEK-----ISCIILDGIFG- 405
Query: 118 APKVAHELHLPSTLLWIQAATVFDI-FYYYFH-----EHGDYITNKSKD-ETCLISLPGL 170
+A +L + I T+ F+ YF E + +D + + ++PG+
Sbjct: 406 --DLATDLAAEFGIQLIHFRTISSCCFWAYFCVPKLLECNELPIRGDEDMDRIITNIPGM 579
Query: 171 SFSLKSRDLPSFLLASNTYTFALPSLKEQIQLLNEEINPR-------VLVNTVEEFELDA 223
L+ RDLPSF + K+ I+L + + + ++NT E+ E
Sbjct: 580 ENILRCRDLPSFCRENK---------KDHIRLDDVALRTKQSLKANAFILNTFEDLEASV 732
Query: 224 LNKVDVGKIKMIPIGPLIPSAFLDGKDPTDNSF--GGDVVRVDSKDDYIQWLDSKDEKSV 281
L+++ + K+ IGPL K +SF + +VD + WLDS+ KSV
Sbjct: 733 LSQIRIHFPKLYTIGPLHHLLNTTKKSSFPSSFFSKSNFFKVDRT--CMAWLDSQPLKSV 906
Query: 282 VYVSFGTLAVLSKRQMEEIARALLDSGFSFLWVIRDKKLQQQKEEEVDDDELSCREELEN 341
+YVSFG+ + + ++ EI LL+S FLWVIR +Q++ LS EE
Sbjct: 907 IYVSFGSTTPMKREEIIEIWHGLLNSKKQFLWVIRPNMVQEK-------GLLSELEEGTR 1065
Query: 342 NMNGKIVKWCSQVEVLSHRSLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIE 401
G IV W Q EVLSH+++G F+TH GWNSTLES+ GVPM+ +P + DQ N++ +
Sbjct: 1066KEKGLIVGWVPQEEVLSHKAIGAFLTHNGWNSTLESVVCGVPMICWPYFADQQINSRFVS 1245
Query: 402 DVWKTGLRMEHDEEGMVKVEEIRKCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSS 461
DVWK GL M+ + V VE + + V + EE R+A LA +V GGSS
Sbjct: 1246DVWKLGLDMKDVCDRKV-VENMVNDVMV-----NRKEEFVRSAMDIAKLASKSVSPGGSS 1407
Query: 462 NRNLRSYLNDI 472
N + + I
Sbjct: 1408YNNFQDLIQYI 1440
>TC81622 weakly similar to PIR|T00584|T00584 indole-3-acetate
beta-glucosyltransferase homolog T27E13.12 - Arabidopsis
thaliana, partial (25%)
Length = 1735
Score = 179 bits (454), Expect = 2e-45
Identities = 131/471 (27%), Positives = 236/471 (49%), Gaps = 13/471 (2%)
Frame = +2
Query: 6 HFLIITYPLQGHINPALQFTKRLISLGAK---VTFATTIHLYSRLINKPTIPGLSFATFS 62
H + + +P +GHINP L F K L S +TF T + + P + FAT
Sbjct: 167 HVVAMPFPGRGHINPMLSFCKILTSQKPNNLLITFVLTEEWLTFIGADPKPESIRFATIP 346
Query: 63 DGYDDGQKSFGDEDIVSYMSEFTRRGSEFLTNIILSSKQENHPFTCLIYTLILSWAPKVA 122
+ ++ GD Y + T+ + F + Q P ++ + L W V
Sbjct: 347 NVIPPEREKAGDFPGF-YEAVMTKMEAPFEKLL----DQLELPVDVIVGDVELRWPVNVG 511
Query: 123 HELHLPSTLLWIQAATVFDIFYY---YFHEHGDYITNKSKDETCLISLPGLSFSLKSRDL 179
+ ++P W +A+ + + ++ + +H ++T DE ++PG+S S D+
Sbjct: 512 NRRNVPVAAFWTMSASFYSMLHHLDVFSRKH--HLTVDKLDEQAE-NIPGIS-SFHIEDV 679
Query: 180 PSFLLASNTYTFALP-SLKEQIQLLNEEINPRVLVNTVEEFELDALNKV-DVGKIKMIPI 237
+ L ++ L ++ N +L+ TV+E E + ++ + + + PI
Sbjct: 680 QTVLCKNDHQVLQLALGCISKVPKANY-----LLLTTVQELEAETIDSLKSIFPFPIYPI 844
Query: 238 GPLIPSAFLDGKDPTDNSFGGDVVRVDSKDDYIQWLDSKDEKSVVYVSFGTLAVLSKRQM 297
GP IP ++ K+P + D DYI+WLDS+ +SV+Y+S G+ +S QM
Sbjct: 845 GPSIPYLDIEEKNPANT---------DHSQDYIKWLDSQPSESVLYISLGSFLSVSNAQM 997
Query: 298 EEIARALLDSGFSFLWVIRDKKLQQQKEEEVDDDELSCREELENNMNGKIVKWCSQVEVL 357
+EI AL +SG +L+V R + + + + C ++ G ++ WC Q++VL
Sbjct: 998 DEIVEALNNSGIRYLYVARGETSRLKDK---------CGDK------GMVIPWCDQLKVL 1132
Query: 358 SHRSLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTGLRME----HD 413
SH S+G F +HCGWNSTLE++ +GVP++ FP + DQ N+ I D WK G ++E +
Sbjct: 1133SHSSIGGFWSHCGWNSTLETVFAGVPILTFPLFLDQVPNSTQIVDEWKNGWKVEIQSKLE 1312
Query: 414 EEGMVKVEEIRKCLEVVMG-KGEKGEELRRNAKKWKDLARAAVKEGGSSNR 463
+ ++ E+I + ++ M + ++G+++R A++ K + R A+ +GGSS+R
Sbjct: 1313SDVILAKEDIEELVKRFMDLENQEGKKIRDRARELKVMFRKAIGKGGSSDR 1465
>TC91074 weakly similar to SP|Q9MB73|LGT_CITUN Limonoid
UDP-glucosyltransferase (EC 2.4.1.210) (Limonoid
glucosyltransferase) (Limonoid GTase), partial (35%)
Length = 911
Score = 169 bits (427), Expect = 3e-42
Identities = 87/186 (46%), Positives = 118/186 (62%), Gaps = 2/186 (1%)
Frame = +3
Query: 289 LAVLSKRQMEEIARALLDSGFSFLWVIRDKKLQQQKEEEVDDDELSCREELENNMNGKIV 348
L L + Q+ EIA LLDS S LWV++ + ++E V +E E N GK+V
Sbjct: 240 LCYLPQEQVNEIAHGLLDSNVSLLWVLKPPSKESGRKEHVLPNEFL----EETNERGKVV 407
Query: 349 KWCSQVEVLSHRSLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTGL 408
W Q EVL+H S+ CF+THCGWNS++E+L GVPM+ FP W DQ TNAK + DV+ G+
Sbjct: 408 NWSPQEEVLAHPSVACFITHCGWNSSMEALSLGVPMLTFPAWGDQVTNAKFLVDVFGVGI 587
Query: 409 RM--EHDEEGMVKVEEIRKCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSSNRNLR 466
R+ H + +V +E++KCL + GEKGEEL++NA KWK A AV GGSS+RNL
Sbjct: 588 RLGYSHADNKLVTRDEVKKCL-LEATIGEKGEELKQNAIKWKKAAEEAVATGGSSDRNLD 764
Query: 467 SYLNDI 472
++ DI
Sbjct: 765 EFMEDI 782
Score = 52.4 bits (124), Expect = 4e-07
Identities = 34/88 (38%), Positives = 49/88 (55%)
Frame = +2
Query: 211 VLVNTVEEFELDALNKVDVGKIKMIPIGPLIPSAFLDGKDPTDNSFGGDVVRVDSKDDYI 270
VLV++ +E E D ++ + I PIGPL F + K + GD V+ D + I
Sbjct: 20 VLVDSFDELEHDYIDYISKKSILTRPIGPL----FNNPKIKCASDIRGDFVKSDDCN-II 184
Query: 271 QWLDSKDEKSVVYVSFGTLAVLSKRQME 298
+WL+SK SVVY+SFGT+ + S R E
Sbjct: 185 EWLNSKANDSVVYISFGTIVLPSTRTSE 268
>TC81478 weakly similar to GP|21740720|emb|CAD40841. OSJNBa0086B14.13 {Oryza
sativa}, partial (57%)
Length = 1585
Score = 129 bits (324), Expect(2) = 4e-41
Identities = 120/402 (29%), Positives = 190/402 (46%), Gaps = 29/402 (7%)
Frame = +1
Query: 6 HFLIITYPLQGHINPALQFTKRLISLGAKVTFATTIHLYSRLINK------PTIPGLSFA 59
H ++ +P+QGHIN L+ K L G +TF T + + RL+ + SF
Sbjct: 16 HAVLTPFPVQGHINALLKLGKLLHLRGFHITFVNTEYNHKRLLKSRGPNAFDGLTDFSFE 195
Query: 60 TFSDGYDDGQKSFGDEDI--------VSYMSEFTRRGSEFLTNIILSSKQEN-HPFTCLI 110
T DG GD D+ +S M+ F + FL + S+ P TCL+
Sbjct: 196 TIPDGLTPTD---GDGDVSQDLRALCLSIMNNFHQFFGVFLAKLNDSATAGLIPPVTCLV 366
Query: 111 YTLILSWAPKVAHELHLPSTLLWIQAATVFDIFYYYFHEHGDYITN--KSKDETCLIS-- 166
+++ A E LP L +A+ F Y FH + KDE+ L
Sbjct: 367 SDCNMAFTVHAAEEHALPIVLFSPCSASYF---YSTFHITKLFQNGVLPLKDESNLTDGN 537
Query: 167 -------LPGLSFSLKSRDLPSFLLASNTYTFALPSLKEQIQLLNE-EINPRVLVNTVEE 218
+PGL S+ +D P + + +K +I+ ++ + ++ NT E
Sbjct: 538 LDTKVEWIPGLK-SISLKDFPDIIRIKDPDV-----IKYKIEETDKCQRGSTIIFNTSNE 699
Query: 219 FELDALNKVDVGKIKMIPIGPLIPSAFLDGKDPTDN--SFGGDVVRVDSKDDYIQWLDSK 276
E DA+N + + IGP S+FLD + P ++ S ++ + D+K ++WL+SK
Sbjct: 700 LESDAINALSSIFPSVYTIGPF--SSFLD-QIPENHLKSLDSNLWKEDTK--CLEWLESK 864
Query: 277 DEKSVVYVSFGTLAVLSKRQMEEIARALLDSGFSFLWVIRDKKLQQQKEEEVDDDELSCR 336
+ SVVYV+FG++ V+S+ ++ E A L +S FLW+IR L + + D L
Sbjct: 865 EPGSVVYVNFGSITVMSREKLLEFAWGLANSKKPFLWIIRPD-LVIGGSQVLSSDFLK-- 1035
Query: 337 EELENNMNGKIVKWCSQVEVLSHRSLGCFMTHCGWNSTLESL 378
E + G I WC Q +VL+H S+G F+THCGWNS +ES+
Sbjct: 1036---EISDRGLIASWCPQEKVLNHPSIGGFLTHCGWNSIMESI 1152
Score = 57.4 bits (137), Expect(2) = 4e-41
Identities = 31/92 (33%), Positives = 56/92 (60%)
Frame = +2
Query: 381 GVPMVAFPQWTDQTTNAKLIEDVWKTGLRMEHDEEGMVKVEEIRKCLEVVMGKGEKGEEL 440
GVPM+ +P + DQ ++++I + W+ G++++ + VK EE+ K + +M GEKG+++
Sbjct: 1160 GVPMLCWPFFADQPLSSRIICEEWEIGMKIDTN----VKREEVEKLINELM-VGEKGKKM 1324
Query: 441 RRNAKKWKDLARAAVKEGGSSNRNLRSYLNDI 472
R+ A + K A + GGSS NL + D+
Sbjct: 1325 RQKATELKKKAAEDTRLGGSSYMNLDKVIKDV 1420
>TC82794 weakly similar to GP|3928543|dbj|BAA34687.1 UDP-glucose
glucosyltransferase {Arabidopsis thaliana}, partial
(59%)
Length = 1309
Score = 148 bits (374), Expect(2) = 1e-40
Identities = 118/377 (31%), Positives = 199/377 (52%), Gaps = 22/377 (5%)
Frame = +1
Query: 57 SFATFSDGYD----DGQKSFGDEDIVSYMSEFTRRG-----SEFLTNIILSSKQENH--P 105
+F T DG DG S +DI+S +S+ R+ E L + S + H P
Sbjct: 172 TFETIPDGLTPIEGDGDVS---QDIIS-LSDSIRKNFYHPFCELLARL-KDSSNDGHIPP 336
Query: 106 FTCLIYTLILSWAPKVAHELHLPSTLLWIQAATVFDIFYYYFHEHGDYITNKSKDETCLI 165
+CL+ + L++ + A E LPS +L+ A+ + +F D KDE+ L
Sbjct: 337 VSCLVSDIGLTFTIQAAEEHGLPS-VLFSSASACSLLSALHFRTLIDKGVIPLKDESYLT 513
Query: 166 S---------LPGLSFSLKSRDLPSFLLASNTYTFALPSLKEQIQLLNEEINPRVLVNTV 216
+ +PGL +L+ +DLP F+ ++ + + E ++E + ++ NT
Sbjct: 514 NGYLDTKVDWIPGLG-NLRLKDLPDFIRTTDPNDIMIKFIIEAADRVHEANS--IVFNTS 684
Query: 217 EEFELDALNKVDVGKIKMIPIGPLIPSAFLDGKDPTDN--SFGGDVVRVDSKDDYIQWLD 274
+E E D +N + + + IGPL ++FL+ + P +N S G ++ + D K ++WL+
Sbjct: 685 DELENDVINALSIKIPSIYAIGPL--TSFLN-QSPQNNLASIGSNLWKEDMK--CLEWLE 849
Query: 275 SKDEKSVVYVSFGTLAVLSKRQMEEIARALLDSGFSFLWVIRDKKLQQQKEEEVDDDELS 334
SK++ SVVYV+FG++ V++ Q+ E A L +S FLW+IR L + D ++
Sbjct: 850 SKEQGSVVYVNFGSITVMTPDQLLEFAWGLANSKKPFLWIIRPD-LVIGGSVILSSDFVN 1026
Query: 335 CREELENNMNGKIVKWCSQVEVLSHRSLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQT 394
E + G I WC Q +VL+H S+G F+THCGWNST+ES+ +GVPM+ +P + +Q
Sbjct: 1027-----ETSDRGVIASWCPQEKVLNHPSVGGFLTHCGWNSTMESICAGVPMLCWPFFAEQP 1191
Query: 395 TNAKLIEDVWKTGLRME 411
TN + I + W+ G ++
Sbjct: 1192TNCRYICNEWEIGAEID 1242
Score = 36.6 bits (83), Expect(2) = 1e-40
Identities = 17/33 (51%), Positives = 20/33 (60%)
Frame = +3
Query: 8 LIITYPLQGHINPALQFTKRLISLGAKVTFATT 40
+I YPLQGHINP L+ K L G +TF T
Sbjct: 6 VITPYPLQGHINPLLKLAKLLHLRGFHITFVNT 104
>TC87302 similar to GP|19911199|dbj|BAB86926. glucosyltransferase-8 {Vigna
angularis}, partial (53%)
Length = 1787
Score = 161 bits (408), Expect = 4e-40
Identities = 139/498 (27%), Positives = 232/498 (45%), Gaps = 28/498 (5%)
Frame = +1
Query: 3 QNHHFLIITYPLQGHINPALQFTKRLISLGAKVTFATT---IHLYSRLINKPTI------ 53
Q H + + GHI P + + S G KVT TT + L SR I K I
Sbjct: 49 QELHIIFFPFLANGHIIPCVDLARVFSSRGLKVTIVTTHLNVPLISRTIGKAKINIKTIK 228
Query: 54 -PGLSFATFSDGYDDGQKSFGDEDIVSYMSEFTRRGSEFLTNIILSSKQENHPFTCLIYT 112
P +G ++ + + + + +M T E L +++ K + CL+
Sbjct: 229 FPSPEETGLPEGCENSESALAPDKFIKFMKS-TLLLREPLEHVLEQEKPD-----CLVAD 390
Query: 113 LILSWAPKVAHELHLPSTL--------LWIQAATVFDIFYYYFHEHGDYITNKSKDETCL 164
+ W+ A + ++P + L + A T ++ S E +
Sbjct: 391 MFFPWSTDSAAKFNIPRIVFHGLGFFPLCVLACT---------RQYKPQDKVSSYTEPFV 543
Query: 165 I-SLPGLSFSLKSRDLPSFLLASNTYTFALPSLKEQIQLLNEEINPRVLVNTVEEFELDA 223
+ +LPG +L LP +T L E +E + V+ NT E E
Sbjct: 544 VPNLPG-EITLTKMQLPQLPQHDKVFTKLLEESNE-----SEVKSFGVIANTFYELEPVY 705
Query: 224 LN--KVDVGKIKMIPIGPLIPSAFLDGKDPTDNSFGGDVVRVDSKDDYIQWLDSKDEKSV 281
+ + ++G+ K +GP+ L +D + + G +D + + ++WL SK+ SV
Sbjct: 706 ADHYRNELGR-KAWHLGPVS----LCNRDTEEKACRGREASID-EHECLKWLQSKEPNSV 867
Query: 282 VYVSFGTLAVLSKRQMEEIARALLDSGFSFLWVIRDKKLQQQKEEEVDDDELSCREELEN 341
+YV FG++ V S Q++EIA L S F+WV+R K + + E ++ E +E
Sbjct: 868 IYVCFGSMTVFSDAQLKEIAMGLEASEVPFIWVVR--KSAKSEGENLEWLPEGFEERIEG 1041
Query: 342 NMNGKIVK-WCSQVEVLSHRSLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLI 400
+ G I++ W QV +L H S+G F+THCGWNSTLE + +G+PMV +P + +Q NAK +
Sbjct: 1042SGKGLIIRGWAPQVMILDHESVGGFVTHCGWNSTLEGVSAGLPMVTWPMYGEQFYNAKFL 1221
Query: 401 EDVWKTGLRM-EHDEEGM-----VKVEEIRKCLEVVMGKGEKGEELRRNAKKWKDLARAA 454
D+ K G+ + GM VK + I K + +M G++ EE+R AK++ +AR A
Sbjct: 1222SDIVKIGVGVGVQTWIGMGGGEPVKKDVIEKAVRRIM-VGDEAEEMRSRAKEFGKMARRA 1398
Query: 455 VKEGGSSNRNLRSYLNDI 472
V+ GGSS + + + D+
Sbjct: 1399VEVGGSSYNDFSNLIEDL 1452
>TC82528 weakly similar to PIR|C86356|C86356 hypothetical protein AAF87257.1
[imported] - Arabidopsis thaliana, partial (35%)
Length = 1376
Score = 159 bits (403), Expect = 2e-39
Identities = 103/308 (33%), Positives = 161/308 (51%), Gaps = 2/308 (0%)
Frame = +1
Query: 167 LPGLSFSLKSRDLPSFLLASNTYTFALPSLKEQIQLLNEEINPRVLVNTVEEFELDALNK 226
+PG+ + + +DLP F+ + F + E + ++ NT E E D LN
Sbjct: 388 IPGMK-NFRLKDLPDFIRTKDLNDFMVEFFIEAADQFHRA--SAIVFNTYNELESDVLNA 558
Query: 227 VDVGKIKMIPIGPLIPSAFLDGKDPTDNSFGGDVVRVDSKDDYIQWLDSKDEKSVVYVSF 286
+ + IGPL PS S ++ + D+K ++WL+SK+ +SVVYV+F
Sbjct: 559 LHSMFPSLYSIGPL-PSLLSQTPHNHLESLSSNLWKEDTK--CLEWLESKEPESVVYVNF 729
Query: 287 GTLAVLSKRQMEEIARALLDSGFSFLWVIRDKKLQQQKEEEVDDDELSCREELENNMN-- 344
G++ V++ Q+ E A L DS FLW+IR + V E EN ++
Sbjct: 730 GSITVMTPNQLLEFAWGLADSKKPFLWIIRP--------DLVIGGSFILSSEFENEISDR 885
Query: 345 GKIVKWCSQVEVLSHRSLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVW 404
G I WC Q +VL H S+G F+THCGWNST ES+ +GVPM+ +P + DQ TN + I + W
Sbjct: 886 GLITSWCPQEQVLIHPSIGGFLTHCGWNSTTESICAGVPMLCWPFFGDQPTNCRFICNEW 1065
Query: 405 KTGLRMEHDEEGMVKVEEIRKCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSSNRN 464
+ GL ++ D VK +E+ K + + GEKG+++R+ A + K A + GG S N
Sbjct: 1066EIGLEIDMD----VKRDEVEKLVN-ELTVGEKGKKMRQKAVELKKKAEENTRPGGRSYMN 1230
Query: 465 LRSYLNDI 472
L + ++
Sbjct: 1231LDKVIKEV 1254
>TC85231 similar to GP|13508844|emb|CAC35167. arbutin synthase {Rauvolfia
serpentina}, partial (34%)
Length = 1602
Score = 156 bits (394), Expect = 2e-38
Identities = 115/370 (31%), Positives = 194/370 (52%), Gaps = 11/370 (2%)
Frame = +3
Query: 109 LIYTLILSWAPKVAHELHLPSTLLWIQAATVFDIFYYYFHEHGDYITNKSKDETCLISLP 168
L++++ + A VA +L S L + A +F +F + T +++P
Sbjct: 381 LVFSMFSTDAHDVAKHFNLLSYLFFSSGAVLFSLFLTIPNLDEAASTQFLGSSYETVNIP 560
Query: 169 GLSFSLKSRDLPSFLLASNTYTFALPSLKEQIQLLNEEINPRVLVNTVEEFELDALNKV- 227
G S L ++LP + + + A S+ + Q L+ + V++NT + E + + +
Sbjct: 561 GFSIPLHIKELPDPFICERS-SDAYKSILDVCQKLS--LFDGVIMNTFTDLEPEVIRVLQ 731
Query: 228 DVGKIKMIPIGPLIPSAFLDGKDPTDNSFGGDVVRVDSKDDYIQWLDSKDEKSVVYVSFG 287
D K + P+GP+I ++ ++N + ++WL+++ SV++VSFG
Sbjct: 732 DREKPSVYPVGPMI-------RNESNNEANMSMC--------LRWLENQQPSSVLFVSFG 866
Query: 288 TLAVLSKRQMEEIARALLDSGFSFLWVIR--DKKLQQQKEEEVDDDELSCREELENNM-- 343
+ LS+ Q+ EIA L SG FLWV+R K ++D L E L N
Sbjct: 867 SGGTLSQDQLNEIAFGLELSGHKFLWVVRAPSKNSSSAYFSGQNNDPL---EYLPNGFLE 1037
Query: 344 ----NGKIV-KWCSQVEVLSHRSLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAK 398
NG +V W QVE+L H S+G F++HCGW+STLES+ +GVP++A+P + +Q NAK
Sbjct: 1038RTKENGLVVASWAPQVEILGHGSIGGFLSHCGWSSTLESVVNGVPLIAWPLFAEQRMNAK 1217
Query: 399 LIEDVWKTGLRMEHDEE-GMVKVEEIRKCLEVVMGKGEKGEELRRNAKKWKDLARAAVKE 457
L+ DV K +R + D+E G++K EE+ K ++ +M KG++ E+R+ K+ A + E
Sbjct: 1218LLTDVLKVAVRPKVDDETGIIKQEEVAKAIKRIM-KGDESFEIRKKIKELSVGAATVLSE 1394
Query: 458 GGSSNRNLRS 467
GSS + L S
Sbjct: 1395HGSSRKALSS 1424
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.319 0.136 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,277,984
Number of Sequences: 36976
Number of extensions: 222979
Number of successful extensions: 1656
Number of sequences better than 10.0: 172
Number of HSP's better than 10.0 without gapping: 1483
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1511
length of query: 478
length of database: 9,014,727
effective HSP length: 100
effective length of query: 378
effective length of database: 5,317,127
effective search space: 2009874006
effective search space used: 2009874006
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 60 (27.7 bits)
Medicago: description of AC146706.4


